Impact Factor ISSN: 1449-1907
Int J Med Sci 2026; 23(6):1966-1981. doi:10.7150/ijms.125197 This issue Cite
Review
The Rare RNA Methylations m²G, Cm, m⁵U and ms²i⁶A: Roles in Disease Pathogenesis and Emerging Therapeutic Implications
1. Department of Anesthesiology, The First Affiliated Hospital of Nanchang University, Nanchang, China.
2. The First Clinical Medical College, Nanchang University, Nanchang, China.
3. Department of Oncology, The First Affiliated Hospital of Nanchang University, Nanchang, China.
† These authors have contributed equally to this work and share first authorship.
Received 2025-9-14; Accepted 2026-3-31; Published 2026-4-23
Abstract
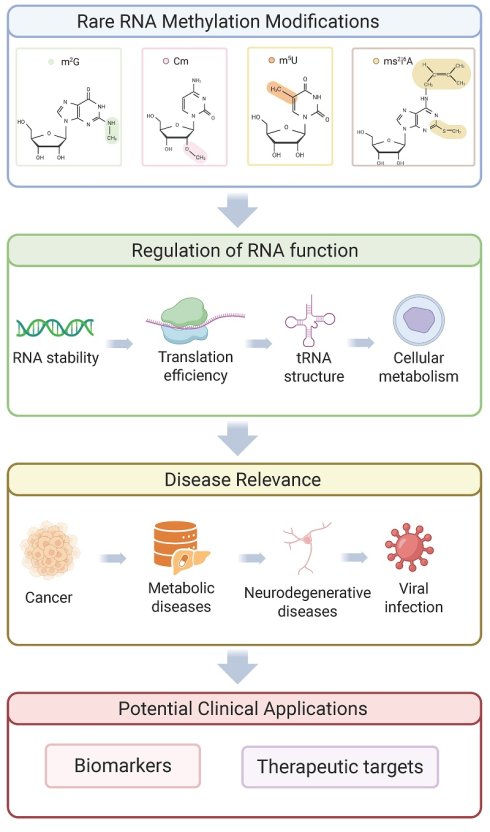
RNA methylation modifications play a central and multifaceted role in various physiological processes by precisely regulating key steps in the RNA life cycle, including nuclear processing, nuclear export, splicing, and cytoplasmic translation. These modifications, which occur on the four nucleotides that constitute RNA strands, are tightly regulated by specific proteins known as “writers,” “readers,” and “erasers.” Advances in high-throughput sequencing and mass spectrometry technologies have progressively unveiled the biological functions of common RNA methylation marks such as N6-methyladenosine (m6A), N1-methyladenosine (m1A), and 5-methylcytosine (m5C). However, our understanding of how RNA modifications influence various cellular processes remains limited, and research focusing on the biological significance of rare RNA methylation modifications is particularly scarce.
This review shifts the research focus toward several relatively understudied and less widely recognized RNA methylation modifications, providing an in-depth analysis of four specific modifications: N2-methylguanosine (m2G), 2′-O-methylcytidine (Cm), 5-methyluridine (m5U), and 2-methylthio-N6-isopentenyladenosine (ms2i6A). It comprehensively elucidates their molecular mechanisms, biological functions, and associations with disease. In addition, this article summarizes the current methodologies available for detecting RNA modifications and discusses the potential applications of these RNA methylation modifications in disease therapy.
Keywords: RNA modification, N2-methylguanosine (m2G), 2′-O-methylcytidine (Cm), 5-methyluridine (m5U), 2-methylthio-N6-isopentenyladenosine (ms2i6A), detection methods, diseases
Introduction
RNA methylation is a common post-transcriptional regulatory mechanism, referring to the transfer of a methyl group from a methyl donor to the nucleotide molecules of RNA under the catalysis of RNA methyltransferases [1-3]. As early as the 1960s, researchers had observed that specific RNA methyltransferases could alter the structure of tRNA [4]. With the continuous advancement of sequencing technologies, the efficiency and accuracy of nucleic acid sequencing have significantly improved. Consequently, in addition to the abundant methylation modifications identified in tRNA and rRNA, diverse types of methylation modifications have also been discovered in many low-abundance RNAs [5-9]. In eukaryotic RNAs, common internal methylation modifications include N6-methyladenosine (m6A), 5-methylcytosine (m5C), 7-methylguanosine (m7G), and N1-methyladenosine (m1A). RNA methylation can occur on any nucleotide residue, including adenosine, uridine, guanosine, or cytidine, and its regulation requires the coordinated actions of methyltransferases (“writers”), demethylases (“erasers”), and RNA-binding proteins (“readers”) [10,11].
Currently, most studies have primarily focused on well-characterized RNA modifications such as m6A, whereas systematic summaries of other less-studied RNA methylation modifications with important biological functions remain limited. Recent studies have demonstrated that RNA modifications such as N2-methylguanosine (m2G), 2′-O-methylcytidine (Cm), 5-methyluridine (m5U), and 2-methylthio-N6-isopentenyladenosine (ms2i6A) play indispensable roles in RNA biosynthesis. N2-methylguanosine is a methylation modification that occurs at the second carbon atom of the ribose moiety in RNA guanosine under the action of methyltransferases. 2′-O-methylcytidine results from the methylation of the 2′-hydroxyl group of the ribose in cytidine, catalyzed by RNA methyltransferases. 5-methyluridine is generated by the methylation of the fifth carbon atom of uridine, catalyzed by pyrimidine methyltransferases. ms2i6A is a complex modification occurring at position 37 of tRNA, involving the sequential addition of an isopentenyl group at the N6 position and a methylthio. These four modifications are predominantly found in tRNAs, and both m5U and m2G share the same type of methyltransferase and function through similar structural folding mechanisms [12,13].
In this review, we focus on the structures, biological functions, and disease relevance of four RNA methylation modifications: m2G, Cm, m5U, and ms2i6A. Based on recent advances, we summarize the regulatory mechanisms, physiological roles, and associations with diseases of these modifications. We also briefly introduce established and emerging techniques for detecting RNA modification sites. Although RNA modifications have garnered increasing attention in recent years, many mechanistic aspects remain to be elucidated. Therefore, a systematic summary of these relatively understudied RNA modifications will help further elucidate the regulatory network of RNA modifications and provide new perspectives for understanding disease mechanisms and developing potential therapeutic strategies. Accordingly, we also provide a preliminary outlook on future research directions in this field.
Regulatory Roles of RNA Methylation Modifications
RNA methylation modifications are likely to exert significant effects on various molecular processes, including RNA metabolism, splicing, stability, and translation [14]. These modifications can alter the biological properties of RNA, affecting its secondary structure, base-pairing capacity, and interactions with proteins. Notably, modifications within ribosomal RNA (rRNA) have been implicated in ribosome biogenesis [15], while those occurring in transfer RNA (tRNA) contribute critically to its biological functions [16]. In messenger RNA (mRNA), modifications often exhibit strong site specificity and ultimately influence gene expression [17]. Moreover, through these mechanisms, RNA methylation modifications may also impact cellular processes such as proliferation, differentiation, and apoptosis, thereby playing important roles in the development of various diseases (Figure 1, Table 1) [18,19].
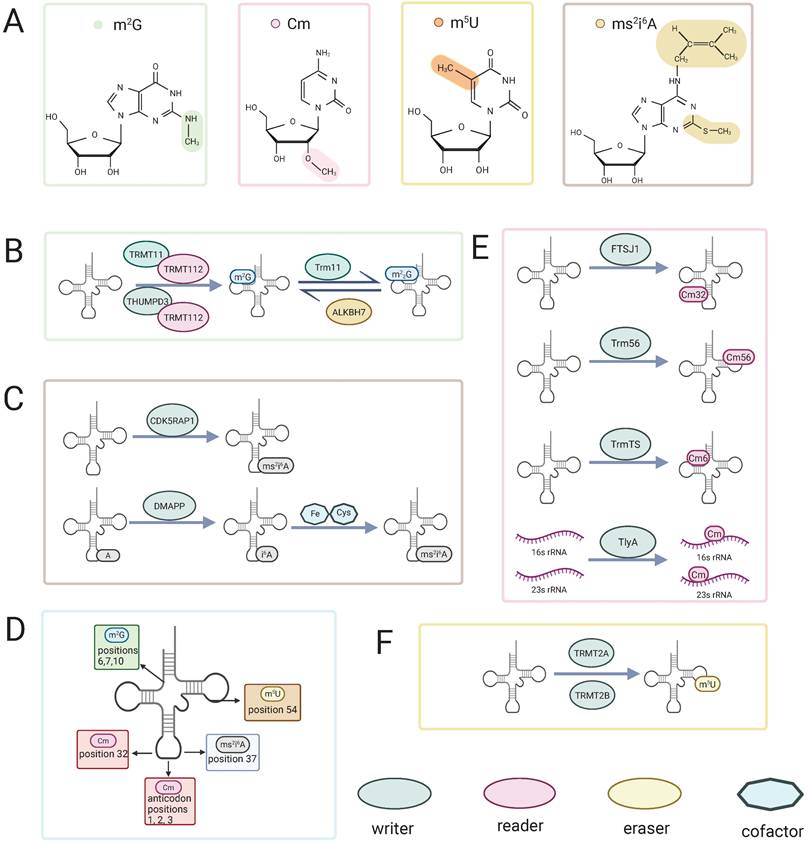
Chemical structures, regulatory enzymes, and tRNA distribution of four rare RNA methylation modifications. (A) Chemical structures of m²G, Cm, m⁵U, and ms²i⁶A.(B, C, E, F) Writers, readers, erasers and cofactors involved in m²G (B), ms²i⁶A (C), Cm (E), and m⁵U (F) methylation. (D) Representative positions of the four modifications on tRNA. Colored shapes denote writers, readers, erasers, and cofactors.
Functions of RNA modifications
| Modification | Process | Enzymes involved | Description | PMID |
|---|---|---|---|---|
| m2G | Protein synthesis | THUMPD3 | Promotes global protein synthesis by reducing rRNA80s monomer accumulation and thereby affecting translation | [18] |
| Cell proliferation | Cell proliferation is suppressed in THUMPD3-deficient cells. | [18] | ||
| mRNA translation | Global protein translation is significantly suppressed in THUMPD3 knockout cell lines. | [23] | ||
| tRNA stability | TRMT11 | Promoting thermal stabilization of tRNAs at the tRNA tenth position | [80] | |
| Formation and functioning of m2G6/7 | N. A | [18] | ||
| Splicing of precursor mRNA | THUMPD2 | The m2G structure formed at position 72 of U6snRNA is functional for precursor mRNA splicing | [18,19] | |
| Mitochondrial activity | ALKBH7 | Conversion of m22G to m2G on mitochondrial tRNA stabilizes mitochondrial activity and function properly | [26] | |
| Processing of nascent tRNAs in mitochondria | Reduces polycis-trans mitochondrial RNA and promotes mitochondria-encoded tRNA processing | [26] | ||
| Cm | Moderated translation | N. A | Maintaining the accuracy of tRNA recognition reading frames | [41-44] |
| tRNA stability | TrmL,TrmJ, Trm7, aTrm56, etc | Helps to correctly identify codons and improve the heat resistance of tRNA molecules. | [37,45] | |
| mRNA stability | N. A | Cm in the cap structure of mRNA distinguishes between autologous and non-autologous RNAs. | [37] | |
| RNA stability | N. A | Altered hydrogen-bonding interactions of nucleosides stabilize RNA double strands and increase the thermal stability of nucleic acid chains | [45,46,48] | |
| Moderated translation | N. A | The anticodon first and second and third positions have opposite roles in translation efficiency | [50] | |
| m5U | tRNA stability | TrmA | Involved in tRNA tertiary structure interactions and enhances tRNA tertiary structure thermal stability | [16] |
| Moderated translation | Reduced translation errors and increased ribosomal A-site binding | [59] | ||
| tmRNA stability | Stabilizing effect on the alanine structure of the stem branch of the Escherichia coli tmRNA receptor | [59] | ||
| Moderated translation | Trm2 | The m5U54 modification of tRNA promoters has an inhibitory effect on translation initiation and elongation m5U54 promoting tRNA modification and regulating the rate of ribosomal translocation | [62,73, 74] | |
| Immune escape | TRMT2A | Synergistic effect of Um54 and m5U54 modifications in tRNAs to avoid recognition and attack of endogenous RNAs by the autoimmune system | [75] | |
| ms2i6A | mRNA translation | tRNA-DMAPP/MiaB | The ms2i6A modification of tRNA stabilizes codon-anticodon interactions on the ribosome | [15] |
| Mitochondrial gene expression | CDK 5 RAP 1 | Cdk5 Rap 1-mediated ms2i6A modification in mt-tRNA regulates mitochondrial translation and energy metabolism | [15,76] |
Abbreviation: m2G, N2-methylguanosine; Cm, 2′-O-methylcytidine; m5U, 5-methyluridine; ms2i6A, 2-methylthio-N6-isopentenyladenosine; tRNA, transfer RNA; rRNA, ribosomal RNA; mRNA, messenger RNA; N. A, not available
m2G RNA modification
Introduction to m2G methylation
N2-methylguanosine (m2G) is a relatively conserved RNA modification, initially discovered in tRNAs from human cells [20]. Subsequent studies have identified m2G modifications in certain small RNAs and mitochondrial RNAs as well [18,21].
The formation of m2G relies on the action of “writer” (methyltransferases). Several methyltransferases responsible for m2G modification have been identified in humans, including TRMT11, THUMPD2, and THUMPD3. These enzymes typically function in coordination with the cofactor protein TRMT112 (known as Trm112 in Saccharomyces cerevisiae) [18,22,23]. These methyltransferases belong to Class I enzymes characterized by a Rossmann fold [18], a structural motif that enables the binding of the methyl donor S-adenosylmethionine (SAM) and stabilizes the methyltransferase structure through interaction with TRMT112 [13]. Moreover, cooperative action of the methyltransferase THUMPD3 and its partner protein TRMT112 enables specific recognition of the G6 position and the 3′-CCA terminus, with an intact tRNA tertiary structure being indispensable for recognition by the modifying enzyme [23]. Similarly, the TRMT11-TRMT112 complex is responsible for the formation of methylguanosine at position 10 in tRNA [20,24]. During this process, TRMT112 serves as a critical scaffold, enhancing the interaction between methyltransferases and tRNA and facilitating the capture of the free methyl donor SAM [13]. Once methylguanosine is formed, it can induce structural alterations in rRNA subunits, subsequently influencing translation and protein synthesis. Interestingly, evidence suggests that methylation at these two specific positions may act synergistically in terms of both formation and function [18]. In addition, recent studies have confirmed the presence of m2G modifications in certain small nuclear RNAs (snRNAs) and mitochondrial RNAs in human cells (Figure 1) [18,21].
Functions of m2G modification
The m2G modification exerts distinct functional effects depending on its presence in different RNA species. Studies have demonstrated that the formation and functional roles of m2G at positions 6/7 and 10 are mutually enhanced. Specifically, when the m2G modification occurs at positions 6/7 of tRNA, it reduces the accumulation of 80S ribosomal monomers, thereby promoting translation efficiency and slightly enhancing protein synthesis. In contrast, the m2G modification at position 10 alone has not been reliably shown to affect protein synthesis. Interestingly, when both the m2G modifications at positions 6/7 and 10 coexist, the translational promotion effect of m2G at position 6/7 is significantly amplified [18]. A previous study reported that, in THUMPD3-knockout cell lines, the levels of tRNA m²G at position 6 in tRNAs bearing guanosine at this site are decreased, accompanied by a marked suppression of global protein translation and inhibition of cell proliferation [23]. This suggests that the m2G modification at position 6 plays a positive role in regulating cell proliferation and protein synthesis. Additionally, in Thermococcus kodakarensis, the 2-amino group of guanosine at position 10 in tRNA is converted, via the intermediate m²G10, to m22G10 by the archaeal Trm11 (tRNA m²G10/ m22G10 methyltransferase), and this modification is thought to promote correct tRNA folding and maintain its structural stability at high temperatures [25].
Beyond tRNA, the THUMPD2-TRMT112 complex catalyzes the formation of m2G at position 72 of U6 small nuclear RNA (snRNA), a modification that is crucial for pre-mRNA splicing, although the precise mechanisms remain incompletely understood [18]. Furthermore, in mitochondria, m2G modifications are generated from N2,2-dimethylguanosine (m22G) via the action of the demethylating enzyme ALKBH7. These modifications play a key role in maintaining mitochondrial function and regulating the processing of nascent mitochondrial tRNAs [26].
In conclusion, m2G modifications on tRNAs are widely distributed in human cells and play vital roles in promoting global protein synthesis and cell proliferation. Moreover, rare m2G modifications at specific RNA sites, such as those found in snRNA and mitochondrial RNA, may participate in pre-mRNA splicing and the stabilization of RNA structures. While substantial progress has been made in understanding m2G modifications, the molecular mechanisms underlying this modification remain incompletely elucidated, warranting further investigation (Figure 1, Table 1).
Cm RNA modification
Introduction to Cm methylation
The 2′ position is the RNA ribose position that undergoes modification in biological systems. In this modification, the hydrogen on the 2′-hydroxyl group of the ribose nucleotide is replaced by another group [27,28]. Its methylation is one of the most widely occurring RNA modifications [29]. This modification primarily appears in tRNA and rRNA molecules. Cm, a type of 2′-O-methylation, is formed by the methylation of the 2′-hydroxyl group of the ribose in cytidine. The Cm modification was first identified in the 1970s in Escherichia coli [30], yeast [31], and mammalian cells [32,33]. Since then, this modification has been widely detected in a variety of prokaryotic and eukaryotic cells. The formation of Cm32 in t RNASer1 and t RNAGln2 in E. coli is catalysed by the enzyme TrmL [34], which is responsible for the addition of Cm34 to E. coli tRNALeu (anticodon CAA) [35]. Cm32 is formed in yeast when Trm7 interacts with Trm732 [36]. In humans, the Cm32 and Gm34 modifications of tRNAphe are catalyzed with the assistance of FTSJ1 [37]. In archaea, Cm is the most common modification in tRNA, and Cm56 is formed at position 56 under the catalysis of the tRNA methyltransferase Trm56 [38]. In prokaryotic 16S and 23S rRNA molecules, 2′-O-methylation modifications, including Cm, account for a significant proportion of the total modifications, ranging from 64% to 86% of the modified residues [28]. The dual substrate rRNA methyltransferase TlyA catalyzes the formation of Cm in both 16S and 23S rRNA [39]. Moreover, recent studies have shown that, similar to the m²G modification, a THUMP domain-containing tRNA methyltransferase, TrmTS, catalyzes the formation of Cm6 in Thermococcus kodakarensis tRNATrp (Figure 1) [40].
Functions of Cm Modification
2′-O-methylation primarily occurs through recognition on pyrimidines and plays a role in regulating gene expression, limiting the ability of nucleosides at the 2′ position to act as proton donors [41-44]. In tRNA, the presence of Cm effectively prevents the hydrolysis of the phosphodiester backbone [45]. In the cap structure of mRNA, Cm helps distinguish self from non-self RNA [37].
Cm modification influences the hydrogen-bonding interactions of nucleosides, thereby affecting their conformation in the RNA chain [46]. Studies have shown that Cm stabilizes the C3′-endo conformation of the ribose ring compared to the C2′-endo conformation [45,47], thus stabilizing the RNA double-stranded structure [48]. The thermal stability of the modified nucleic acid chain also increases [49]. Notably, when Cm is present at the first position of the anticodon, it significantly enhances the efficiency of tRNA recognition of the G-end codon [50]. These modifications, which alter nucleic acid properties, are likely closely related to conformational changes induced by Cm. Interestingly, when Cm is present at the second or third position of the anticodon, it may reduce codon recognition efficiency [50]. Therefore, in organisms, tRNAs typically have Cm modifications at the first nucleotide of the anticodon [51-57], while such modifications are absent at the second or third positions (Figure 1, Table 1).
m5U RNA modification
Introduction to m5U methylation
The m5U modification is widely distributed in various RNA types. It is most abundant in tRNA, typically located at position 54 in the T-loop of tRNA [58]. In addition to tRNA, this modification is also found in a variety of other coding and non-coding RNAs, such as tmRNA [59], rRNA, and mRNA [60,61].
The enzymes catalyzing the formation of the m5U modification, as well as the m2G modification, belong to class I methyltransferases. These enzymes possess a Rossmann fold structure, which facilitates the transfer of a methyl group from S-adenosylmethionine (SAM) to the target nucleoside [12]. Specific enzymes responsible for m5U modification exist in bacteria, yeast, and mammals [62-67]. In human cells, the methyltransferase TRMT2A (tRNA methyltransferase 2 homolog A) primarily provides the m5U modification at position U54 in cytoplasmic tRNA [68]. Meanwhile, TRMT2B (tRNA methyltransferase 2 homolog B) catalyzes the formation of m5U in mitochondrial tRNA and 12S rRNA in humans [69]. TRMT2A and TRMT2B share similar gene sequences and structural composition with the Saccharomyces cerevisiae tRNA methyltransferase Trm2, and they are considered human homologs of Trm2. These enzymes have also been found to be associated with various human diseases (Figure 1) [70].
Functions of m5U modification
The m5U54 modification not only increases the thermal stability of tRNA [16], reduces translation error rates, but also enhances the binding affinity of the ribosomal A-site. Additionally, m5U plays a stabilizing role in the receptor stem-loop of Escherichia coli tmRNA [59]. In summary, in lower organisms such as E. coli, m5U54 acts as a crucial regulator for tRNA maturation and ribosomal translocation during protein synthesis [71-73].
In eukaryotes, the m5U54 modification on initiator tRNA inhibits translation initiation and elongation [62,74]. Recent studies have also revealed that the synergistic effect between Um54 and m5U54 modifications in human tRNALys3 can suppress Toll-like receptor 7 (TLR7)-dependent immune responses, thus preventing endogenous RNA from being recognized and attacked by the immune system (Figure 1, Table 1) [75].
ms2i6A RNA modification
Introduction to ms2i6A methylation
The 2-methylthio-N6-isopentenyladenosine (ms2i6A) modification is a widely conserved modification found in transfer RNA (tRNA) with evolutionary significance [76]. This modification primarily occurs at the 37th position of adenosine in tRNA and is involved in recognizing codons that start with uridine. The biosynthesis of ms2i6A involves multiple steps. Initially, isopentenyl groups are added to the N-6 nitrogen of adenosine, catalyzed by tRNA-dimethylallyl pyrophosphate transferase (tRNA-DMAPP), encoded by the miaA gene in Escherichia coli and the MOD5 gene in yeast, producing the precursor i6A [77]. Subsequently, the MiaB enzyme, encoded by the TM0653 gene, catalyzes the attachment of a methylthio group (-SCH3) at the C(2) position of A37 through a reaction dependent on Fe2+, cysteine, and S-adenosylmethionine (SAM), completing the hypermodification [78]. Notably, in mammalian mitochondria, CDK5 regulatory subunit-associated protein 1 (CDK5 RAP 1), a radical SAM enzyme homologous to the bacterial MiaB protein, specifically converts the A37 i6A modification in DNA-encoded tRNA into ms2i6A [76]. This modification process is critical for mitochondrial translation and energy metabolism in mammals, and its disruption is closely associated with the pathogenesis of mitochondrial diseases (Figure 1).
Functions of ms2i6A modification
Previous studies have shown that the 2-methylthio-N6-isopentenyladenosine (ms2i6A) modification stabilizes the codon-anticodon interaction on the ribosome. When the first codon position A36 on the ribosome correctly pairs with U on tRNA, the ms2i6A modification helps maintain the stability of the codon-anticodon interaction. Conversely, if there is a mismatch at the first codon position, this modification may be weakened or destabilized [15]. Furthermore, research has found that in the absence of the initial isopentenyl modification, there is a significant impact on the ribosome's ability to recognize nonsense codons. For instance, the translation efficiency of the UGA codon decreases by a factor of 60 [79]. Taken together, the ms2i6A modification plays a crucial role in the codon-anticodon interaction on the ribosome and the translation process of tRNA (Figure 1, Table 1).
Techniques for Detecting RNA Modifications
Similar to the detection of other epigenetic modifications, the most established techniques for RNA methylation analysis are primarily based on liquid chromatography and mass spectrometry. Through continuous efforts and improvements, researchers have developed a variety of optimized detection protocols by combining chromatography with mass spectrometry. Advances in sequencing technologies, particularly the advent of high-throughput sequencing, have further accelerated transcriptome-wide studies of RNA modifications, enabling the precise localization of modified sites. Meanwhile, the emergence of deep learning techniques has provided novel approaches for the prediction and detection of RNA methylation sites, opening new avenues for research in this field (Table 2).
Detection Methods for RNA m2G, Cm, m5U, and ms2i6A modifications.
| Methods | Description | Advantages | Limitations | PMID |
|---|---|---|---|---|
| Chromatography | It is a physicochemical separation method that utilizes properties such as solubility and adsorption of substances. | Convenient operation, high sensitivity, accurate quantitative results, can separate complex multi-component mixtures. | Difficulty in completing qualitative analysis. Conventional chromatography takes a long time to detect. | [32] |
| RNA-MS | Mass spectra are obtained by focusing ions with different mass-to-charge ratios at different points in a magnetic field to determine their mass. | Qualitative analysis can be completed with accurate results. | Not suitable for analysis of complex compounds. | [23] |
| LC-MS | Combining chromatography and mass spectrometry. | Combining the advantages of chromatography and mass spectrometry. | The operation process is complex and costly. | [83-85] |
| LC-MS/MS | Combining chromatography and mass spectrometry. | It is more efficient compared to LC-MS, with less sample dosage, faster analyzing speed, and wider detection range. | The operation process is complex and costly. | [83-85] |
| Reversed-phase liquid chromatography | Liquid chromatography in which the polarity of the mobile phase is greater than the polarity of the stationary phase. | High efficiency, high separation capacity, and universal applicability. | Polar and hydrophilic compounds are rarely or not retained. | [83] |
| HILIC | The stationary phase of the HILIC column is hydrophilic, and the more hydrophilic the compound, the longer it is retained. | Suitable for separation of strongly polar and hydrophilic compounds. | Not suitable for separating non-polar compounds, and the separation efficiency of the column deteriorates after a period of use. | [84] |
| ESI-MS | Electrospray is utilized to generate ions that apply a high voltage to a liquid to create an aerosol. | Overcomes the tendency of macromolecules to break up when ionized. | Sensitivity is affected by spray voltage. | [63] |
| UHPLC | Liquid chromatography using small-particle, high-performance particulate stationary phases. | Increases analytical throughput, sensitivity and peak capacity. | Instrument maintenance costs are high. | [85] |
| FICC-Seq | Modification sites were determined using covalent cross-linking principles and high-throughput sequencing. | Efficient and Accurate. | High Costs. | [71] |
| Reaction-Seq | ms²i⁶A was specifically bioorthogonally labeled via chemoselective reactions. | allows detection of ms²i⁶A modification at single-nucleotide resolution | complex and costly. | [87] |
| Machine learning models | Modification sites were predicted by computerized processing of the data. | Simple operation, high efficiency and low cost. | Possible errors and accuracy could be improved. | [88-90] |
Abbreviation: RNA-MS, RNA-mass spectrometry; LC-MS, liquid chromatography-mass spectrometry; LC-MS/MS, liquid chromatography-tandem mass spectrometry; HILIC, hydrophilic interaction liquid chromatography; ESI-MS, electrospray ionization-mass spectrometry; UHPLC, ultrahigh performance liquid chromatography; FICC-Seq, Fluorouracil Induced-Catalytic-Crosslinking-Sequencing.
Liquid chromatography is one of the classical methods for RNA modification analysis. Prior to detection, RNA is hydrolyzed into its constituent components, which are then separated based on the differential interactions between the stationary phase and the mobile phase within the chromatographic column. This technique is simple to operate, offers good reproducibility, and enables precise quantitative analysis of RNA modifications; however, it has relatively weak qualitative capabilities. In the early studies of nucleosides, liquid chromatography was widely employed. As early as the 1970s, chromatography was used to separate methylated ribonucleoside products such as Am, Gm, Um, and Cm, leading to the discovery of methyltransferase activity in rat cells [32].
Mass spectrometry (MS) is a commonly used technique for the qualitative detection of RNA modifications. Based on electromagnetic principles, mass spectrometry separates substances according to their mass-to-charge ratios and identifies their chemical compositions. The combination of RNA mass spectrometry (RNA-MS) with reverse genetics approaches has led to the discovery of the m²G modification [23]. In addition, the use of automated hydrogen-deuterium exchange coupled with mass spectrometry and unbiased proximity labeling methods has elucidated the interaction patterns between TRMT112 and methyltransferases [22,87]. A novel method termed “Single Neuron RNA Modification Analysis by Mass Spectrometry” (SNRMA-MS) enables the detection and quantification of post-transcriptional nucleoside modifications at the single-cell level [88]. Furthermore, nucleic acid isotope labeling coupled with mass spectrometry (NAIL-MS) has revealed a significant increase in Cm levels during the logarithmic phase of cell growth [89].
The combination of mass spectrometry (MS) and liquid chromatography (LC) represents a major approach for RNA modification analysis, leveraging the strengths of both techniques to achieve accurate qualitative and quantitative assessments of RNA modifications. This integrated strategy is particularly important for precise RNA modification sequencing. Liquid chromatography-mass spectrometry (LC-MS)-based methods mainly analyze RNA modifications at the nucleoside level, requiring enzymatic digestion of nucleic acid samples prior to analysis. Identification of modified nucleosides is typically achieved by comparison with authentic standards or RNA modification databases. Using reversed-phase liquid chromatography (LC)-mass spectrometry (MS) and tandem mass spectrometry (LC-MS/MS/MS) combined with stable isotope dilution techniques, significant differences in the abundance of certain modifications, including Cm, have been observed across different tissues [81]. Moreover, hydrophilic interaction liquid chromatography coupled with mass spectrometry/mass spectrometry (HILIC-MS/MS) has been developed to analyze RNA modification-protein interactions and to identify Cm methylation sites [82]. Cheng et al. successfully identified the m⁵U modification in mammalian mRNA and confirmed its methyl donor as S-adenosylmethionine (SAM) by employing N-cyclohexyl-N'-β-(4-methylmorpholinium) ethylcarbodiimide p-toluenesulfonate (CMCT) labeling coupled with liquid chromatography-electrospray ionization tandem mass spectrometry (LC-ESI-MS/MS) [60]. Recently, a novel technique, ultra-high performance liquid chromatography-electrospray ionization-tandem mass spectrometry (UHPLC-ESI-MS/MS), has been developed. This method exhibits high sensitivity for detecting low-abundance modified nucleosides and enables their rapid and accurate quantification [83]. In addition, Danijel Djukovic et al. employed high-performance liquid chromatography coupled with triple quadrupole mass spectrometry (HPLC/TQMS) and demonstrated that serum m²G metabolite levels are significantly elevated in patients with esophageal adenocarcinoma [90].
In addition to the aforementioned approaches, several other methods have been developed to detect RNA methylation modifications. Carter et al. established the 5-fluorouridine-induced catalytic crosslinking sequencing (FICC-Seq) method, which demonstrated that the m⁵U54 modification in cytoplasmic tRNA is catalyzed by TRMT2A in human cells [68]. This technique, based on existing high-throughput sequencing platforms, covalently crosslinks methyltransferases to their target RNA residues. Reverse transcription of these crosslinked RNAs stalls at the crosslinking sites, enabling precise identification of modification sites. Fang et al developed a redox-activated chemical tagging sequencing technique (Reaction-Seq). This method exploits the chemoselective reaction between methylthio groups and oxaziridine moieties to achieve specific bioorthogonal labeling of ms²i⁶A without cross-reactivity toward canonical nucleosides, thereby enabling efficient enrichment of ms²i⁶A-modified RNA prior to sequencing and markedly improving detection accuracy [91]. With the advancement of bioinformatics, several novel deep learning-based frameworks have been developed that can accurately predict RNA m⁵U modification sites, offering tremendous potential for applications in future biotechnological industries [84-86].
In summary, technological advancements have continuously provided more effective tools and methodologies for RNA modification detection. The integration of emerging technologies with classical experimental methods offers researchers a diversified set of strategies. Furthermore, the rapid development of bioinformatics and the emergence of new sequence-based prediction servers have significantly enhanced the efficiency and accuracy of RNA modification site detection, laying a solid foundation for future applications in medical diagnostics and therapeutics.
The Relationship Between RNA Methylation Modifications and Disease
m2G
Research has shown that the level of m2G is elevated in thyroid malignant tumor cells, potentially due to the role of this modification in cell proliferation. However, the underlying mechanisms remain incompletely understood [92]. Studies have reported that the serum m2G levels in colorectal cancer (CRC) patients are significantly lower than those in healthy individuals, suggesting that m2G may serve as a potential serological biomarker for preliminary screening of colorectal tumors in the future [93]. Additionally, it has been found that the simultaneous loss of the m2G-modifying methyltransferases TRMT11 and THUMPD3 affects the proliferation of colorectal cancer cells, indicating that the associated gene loci may become potential targets for controlling tumor proliferation [18]. Further research has revealed that overexpression of ALKBH7 in liver cancer cells, breast cancer (BC) cells, and head and neck squamous cell carcinoma leads to an increase in m2G structures [94]. The possible mechanism involves m22G structures hindering base pairing, while ALKBH7 converts m22G to m2G. The resulting monomethylated guanosine has a minimal impact on the base pairing process, promoting tumor cell proliferation. In lung cancer cells, depletion of THUMPD3 significantly impaired the cell's adaptability, negatively affecting key cellular processes such as proliferation and migration [95]. However, the association between this phenomenon and m2G requires further investigation. Previous studies have shown that serum m²G levels are significantly higher in patients with esophageal adenocarcinoma than in healthy individuals, which may provide a reliable indicator for future screening of esophageal adenocarcinoma [90].
In addition to its role in tumors, m2G modifications have been shown to be effective in preventing the infection of certain retroviruses. The m2G modification may prevent reverse transcriptase and small nuclear capsid proteins from recognizing corresponding sites or may inhibit retrotranscription RNA replication in normal cells by compacting the structure of tRNATrp [96].
A recent study indicated that THUMPD2 catalyzes the methylation of U6 snRNA m2G72. Knockdown of THUMPD2, which eliminates U6 snRNA m2G72, weakens precursor mRNA splicing activity, producing abnormally spliced precursor mRNA, which in turn leads to mRNA degradation. The loss of THUMPD2 can affect retinal function regulation and is associated with age-related macular degeneration (AMD), suggesting that abnormalities in U6 snRNA m2G72 modifications may contribute to the development of this disease (Figure 2, Table 3) [19].
The roles of RNA m2G, Cm, m5U, and ms2i6A methylation modifications in various diseases. m²G: In colorectal cancer, concurrent loss of TRMT11 and THUMPD3 reduces m²G modification on tRNA, thereby regulating tumor cell proliferation. In hepatocellular carcinoma cells, breast cancer (BC) cells, and head and neck squamous cell carcinoma cells, ALKBH7 converts m22G to m²G, which in turn promotes tumor cell proliferation. In the context of retinal function, loss of THUMPD2 leads to a decrease in the m²G72 modification of U6 small nuclear RNA (U6 snRNA), thereby contributing to the development of age-related macular degeneration. ms²i⁶A: CDK5RAP1 converts i⁶A to ms²i⁶A, thereby attenuating the antitumor effect of i⁶A and maintaining the stem-like properties of glioma-initiating cells (GICs). In breast cancer, CDK5RAP1 deficiency activates the ROS/JNK signaling pathway, resulting in upregulation of the p53-mediated pro-apoptotic factors caspase-9 and caspase-3 and ultimately inducing apoptosis. m⁵U: Loss of the tRNA methyltransferase NSun2 reduces tRNA m⁵C modification and diminishes the production of tRF-1s, thereby weakening their interaction with TSR1, suppressing global protein synthesis, and ameliorating liver injury. Metabolomics studies have shown that m⁵U levels are significantly decreased in patients with hypertension, in the serum of breast cancer patients, and in individuals with HIV infection or systemic lupus erythematosus, whereas m⁵U levels are markedly elevated in breast cancer cells, ovarian cancer, NPH, coronary heart disease, and prostate cancer. Cm: In hepatitis C, the triphosphate form of Cm can inhibit the activity of the hepatitis C virus (HCV) RNA polymerase NS5B, thereby suppressing HCV replication and limiting disease progression. Metabolomics analyses indicate that Cm levels are significantly increased in patients with colorectal cancer and clear cell renal cell carcinoma, but markedly decreased in patients with breast cancer, esophageal squamous cell carcinoma, and diabetic retinopathy.
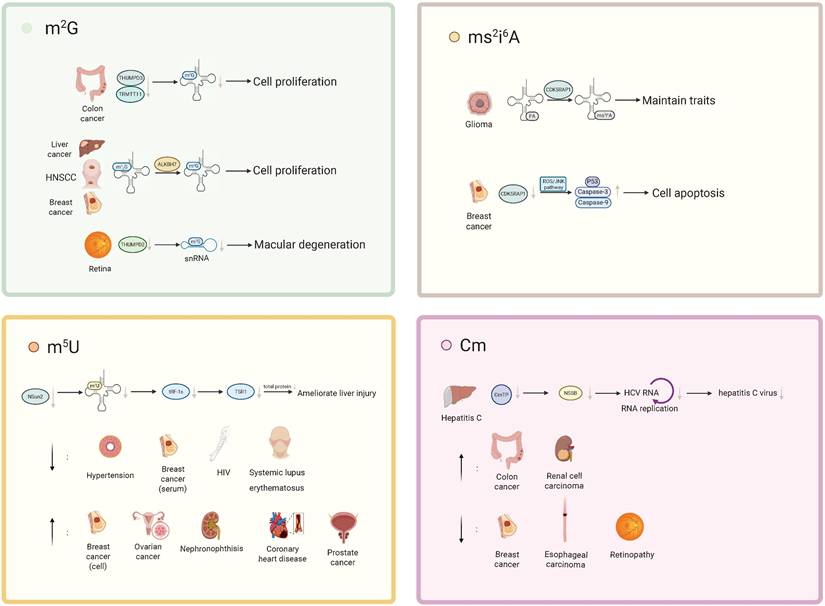
Specific role of modifications in diseases
| Modification | Disease | Enzyme | Target | Description | PMID |
|---|---|---|---|---|---|
| m2G | CRC | THUMPD3 | Position 6 of tRNA | Reduced expression of THUMPD3, THUMPD2, and TRMT11 inhibits global protein synthesis and thus cell proliferation | [18] |
| TRMT11 | Position 10 of tRNA | ||||
| Liver cancer | ALKBH7 | mtRNA | Transformation of mt-RNA m22G to m2G may unblock base pairing by m22G, thereby promoting tumor cell proliferation | [93] | |
| BC | |||||
| HNSCC | |||||
| Lung cancer | THUMPD3 | N. A | Depletion of THUMPD3 significantly impairs cellular adaptability and negatively affects cell proliferation and migration. | [94] | |
| AMD | THUMPD2 | snRNA | Deletion of THUMPD2 eliminates the U6 m2G72 on the 'spliceosome', resulting in aberrant splicing of pre-messenger RNAs, associated with AMD | [19] | |
| EAS | N. A | N. A | The serum level of m²G is significantly higher in patients with esophageal adenocarcinoma than in healthy controls. | [86] | |
| Cm | BC | N. A | N. A | Reduced serum Cm levels in breast cancer patients | [97] |
| CRC | N. A | N. A | Elevated Cm positively correlates with colorectal cancer (CRC) risk | [98-99] | |
| ccRCC | N. A | N. A | Increased Cm levels in clear cell renal carcinoma (ccRCC) | [100] | |
| ESCC | N. A | N. A | Elevated Cm is associated with reduced risk of esophageal squamous cell carcinoma (ESCC) | [96] | |
| Hepatitis C | N. A | N. A | The triphosphate form of Cm can inhibit the activity of the hepatitis C virus (HCV) RNA polymerase NS5B, thereby suppressing HCV replication. | [102] | |
| DR | N. A | N. A | Cm levels are downregulated in DR | [103] | |
| m5U | OC | N. A | N. A | m5U expression level is up-regulated in ovarian cancer tissues, possibly due to hypoxia-associated tumor microenvironment affecting m5U expression | [117] |
| BC | N. A | N. A | m5U was significantly elevated in serum from breast cancer patients and reduced in breast cancer cells cultured in vitro | [97,110] | |
| NPH | N. A | N. A | Deletion of Anks3 affects inner medullary collecting duct cell metabolism in NPH mice, showing accumulation of m5U | [118] | |
| Hypertension | N. A | N. A | m5U urinary metabolites are inversely associated with hypertension | [109] | |
| CAD | N. A | N. A | increased plasma levels of m5U in CAD patients at high risk of major clinical outcomes | [108] | |
| Prostate cancer | N. A | N. A | The concentration of m⁵U is elevated in prostate cancer and may serve as a potential biomarker. | [114] | |
| HIV | N. A | N. A | m⁵U levels are positively correlated with CD4⁺ T cell counts, the CD4/CD8 ratio, and the proportion of CD4⁺ T cells. | [120] | |
| Liver disease | NSun2 | N. A | NSun2 deficiency ameliorates liver injury by reducing the production of tRF-1s, thereby weakening TSR1 interactions. | [126] | |
| ms2i6A | BC | CDK5RAP1 | ROS/JNK pathway | CDK5RAP1 deficiency induces human breast cancer MCF-7 cell cycle arrest and apoptosis through the ROS/JNK signaling pathway | [130] |
| Glioma | CDK5RAP1 | i6A | Conversion of i6A to ms2i6A abolishes the antitumor effect of i6A and protects GIC from excessive autophagy triggered by i6A and maintains GIC traits | [131] |
Abbreviation: m2G, N2-methylguanosine; Cm, 2′-O-methylcytidine; m5U, 5-methyluridine; ms2i6A, 2-methylthio-N6-isopentenyladenosine; CRC, colon cancer; BC, breast cancer; HNSCC, head and neck squamous cell carcinoma; AMD, macular degeneration; EAS, esophageal adenocarcinoma; ccRCC, clear cell renal cell carcinoma; ESCC, esophageal squamous cell carcinoma; DR, diabetic retinopathy; OC, ovarian cancer; NPH, nephronophthisis; Anks3, ankyrin and sterile alpha motif (SAM)-containing 3; CAD, coronary artery disease; tRNA, transfer RNA; rRNA, ribosomal RNA; mRNA, messenger RNA; N.A, not available
Cm
Metabolomic studies have identified Cm as a potential biomarker for several types of cancer, including esophageal squamous cell carcinoma [97], breast cancer [98], colorectal cancer [99,100], clear cell renal carcinoma [101], and liver cancer [102]. Cm is associated with a reduced risk of esophageal squamous cell carcinoma (ESCC) and a positive correlation with colorectal cancer (CRC) risk. Serum levels of Cm are significantly lower in breast cancer patients. In clear cell renal carcinoma (ccRCC), the levels of Cm, along with other modified nucleotides and related enzymes, are elevated. Moreover, the triphosphate form of Cm can inhibit the activity of the hepatitis C virus (HCV) RNA-dependent RNA polymerase NS5B, thereby suppressing HCV replication, and this finding provides a novel therapeutic avenue for the treatment of hepatitis C [103]. Recent studies have shown that Cm levels are downregulated in early diabetic retinopathy (DR), suggesting its potential as a diagnostic biomarker for DR [104]. Furthermore, Cm metabolites have been found to be associated with childhood absence epilepsy (CAE) [105].
Notably, 2'-O-methylated RNA can antagonize various immune-stimulating RNAs, such as by competing with isRNAs to bind to TLR7/8, thereby inhibiting the activation of immune responses. Not only that, the modification also has the ability to activate innate immunity. This mechanism may inspire the design of drugs to modulate the immune response (Figure 2, Table 3) [106-108].
m5U
Metabolomic studies on diseases such as coronary artery disease (CAD), hypertension, breast cancer, multiple sclerosis, and COVID-19 have utilized m5U as a metabolic biomarker for constructing disease risk assessment models [98,109-114]. Notably, the upregulation of m5U in plasma metabolites is associated with a high risk of CAD and can serve as an assessment marker for CAD prognosis and secondary prevention [109]. In contrast, m5U shows a negative correlation with salt-sensitive hypertension [110]. Furthermore, stroke-induced depression is associated with gut metabolic dysregulation, and various metabolites, including m5U, are related to this phenomenon [112]. Recent studies using micellar electrokinetic capillary chromatography with UV detection (MEKC-UV) to compare nucleoside levels in prostate cancer have shown that the concentrations of uridine and 5-methyluridine are elevated in cancer cells, suggesting their potential as biomarkers [115]. In addition, metabolomic analyses in autoimmune diseases [113], dilated cardiomyopathy [116], and high-altitude gastric cancer [117] have also revealed alterations in m⁵U levels.
Research has shown that multiple RNA modifications, including m5U, are upregulated in ovarian cancer tissues. A similar phenomenon is observed in hypoxic environments, suggesting that hypoxia-associated tumor microenvironments may influence the expression of RNA modifications [118]. Interestingly, in vitro studies on breast cancer cells have shown a significant increase in m5U levels in metabolic profiles [111]. At the same time, a study on plasma metabolites in breast cancer patients revealed a significant decrease in m5U metabolites in their plasma [98], indicating that the effect of breast cancer on the generation of m5U metabolites remains to be further investigated.
In Nephronophthisis (NPH), the loss of Anks3 affects the metabolic pathways in the inner medullary collecting duct cells of NPH mice, resulting in m5U accumulation. This suggests that abnormal mutations in Anks3 may affect nucleotide metabolism by causing cellular gene damage [119].
m⁵U has been mentioned in several studies as a potential pharmacological target or therapeutic agent. 5-fluorouracil (5-FU) is a commonly used antimetabolite chemotherapeutic drug, and it has been shown that treatment of HEK293T cells with 5-FU reduces the level of 5-methyluridine in tRNA by approximately 55%, suggesting that 5-FU may exert its antitumor effect, at least in part, by perturbing normal cellular physiology through lowering m⁵U and related tRNA modifications [120]. In addition, m⁵U serves as a key intermediate in the synthesis of the anti-HIV drugs stavudine (d4T) and zidovudine (AZT). Once taken up by cells, m⁵U can be phosphorylated by kinases to its mono- or triphosphate forms, which in turn inhibit viral replication. A metabolomic study in treatment-naïve individuals with HIV infection revealed a significant decrease in m⁵U levels, and demonstrated that m⁵U levels were positively correlated with CD4⁺ T cell counts, the CD4/CD8 ratio, and the proportion of CD4⁺ T cells, but negatively correlated with markers of CD8⁺ T cell activation and inflammation [121]. These findings indicate that m⁵U may play a functional role in HIV therapy and is associated with anti-HIV immune activity, and that further elucidation of the mechanisms underlying m⁵U dynamics in immune cells from patients with HIV could provide additional avenues for the development of anti-HIV agents. Moreover, another study reported reduced m⁵U modification in mRNA from CD4⁺ T cells of patients with systemic lupus erythematosus [122], suggesting that m⁵U may also be implicated in the pathogenic mechanisms of immune-related diseases beyond HIV infection.
In addition, treatment of patients with chronic hepatitis B virus infection with sIFN-α significantly perturbs pyrimidine nucleoside metabolism, leading to decreased levels of 5-methyluridine, which may serve as a potential indicator for monitoring the therapeutic response to IFN-α [123]. IFN-α is generally thought to exert its anti-HBV activity, at least in part, through effects on m⁶A modification [124]; following IFN-α2a treatment, m⁶A modification of pgRNA is increased, thereby regulating viral RNA transcription and suppressing cDNA [125]. IFN-α also induces ISG20-mediated selective degradation of m⁶A-containing HBV RNA [126]. Our findings further suggest that IFN-α may additionally exert antiviral effects by altering m⁵U and other RNA modifications.
Additionally, research has demonstrated that the deletion of tRNA methyltransferase NSun2 reduces both tRNA m5U and m5C modifications while stimulating the production of stress response products, including class I tsRNAs (tRF-1s). The best mutant of tRF-1, tRF-Gln-CTG-026 (tG026), inhibits total protein synthesis by weakening the interaction between TSR1 (pre-rRNA processing protein TSR1 homolog) and tRNA, thereby improving liver injury. This mechanism provides new directions for RNA-based therapies for liver diseases [127]. Interestingly, a similar mechanism is observed in the human m5U methyltransferase TRMT2A, where its deletion leads to low m5U54 modification in tRNA and the formation of tsRNAs, which are associated with cellular stress responses [128]. However, the downstream signaling pathways of this mechanism and its specific relationship with RNA methylation modifications require further exploration. Notably, TRMT2A has been identified as a biomarker for increased recurrence risk in HER2+ breast cancer patients, suggesting that elucidating the physiological mechanisms related to this methyltransferase could provide breakthroughs in understanding the disease (Figure 2, Table 3) [129].
ms2i6A
Previous studies have shown that the ms2i6A modification mediated by CDK5 regulatory subunit-associated protein 1 (Cdk5 rap 1) primarily occurs in mitochondrial DNA-encoded mt-tRNAs, and not in nuclear-encoded RNAs [76]. In Cdk5 rap 1 gene knockout (KO) mice with the transverse aortic constriction (TAC) model, as well as in the stress model induced by KD, the absence of ms2i6A reduced the translation of mitochondrial DNA-encoded respiratory subunits, impairing electron transfer and oxidative respiration. This led to significant damage to heart and skeletal muscle functions due to energy metabolism insufficiency [130]. Therefore, ms2i6A modification is essential for mitochondrial translation.
CDK5RAP1 deficiency induces cell cycle arrest and apoptosis in breast cancer via the ROS/JNK signaling pathway. Studies have shown that CDK5RAP1 deficiency blocks MCF-7 breast cancer cells in the G2/M phase, stimulates ROS generation, and activates p-JNK (a pro-apoptotic factor). This activation of the ROS/JNK pathway results in the overexpression of the tumor suppressor p53 and the upregulation of apoptosis factors caspase-9 and caspase-3, ultimately inducing cell apoptosis [131]. In summary, CDK5RAP1 deficiency provides constructive insights for the development of novel therapies in breast cancer.
CDK5RAP1 eliminates the anti-tumor effects of i6A by converting i6A into ms2i6A, thereby protecting glioma initiating cells (GICs) from excessive autophagy triggered by i6A, maintaining the GIC phenotype [132]. However, CDK5RAP1 is not essential for the normal conversion of i6A to ms2i6A in neural stem cell differentiation [132]. It is understood that i6A inhibits GIC-associated traits and induces autophagic cell death, making targeting i6A a feasible approach for treating GICs (Figure 2, Table 3).
Discussion
Recent research has highlighted the diversity and complexity of RNA modifications in cellular biology. These modifications include well-known methylations such as m6A, as well as other forms such as m2G, Cm, m5U, and ms2i6A methylation [20,32,58,76]. Studies have revealed the physiological functions of RNA methylation modifications and their intricate connections with diseases.
Methylation modifications are crucial for the biological functions of RNA molecules. For instance, modifications like m2G and m5U in tRNA molecules alter their three-dimensional structure, affecting their role in protein synthesis [26,50,80]. These modifications can influence the binding of tRNA to ribosomes, thereby impacting the efficiency and precision of protein synthesis [15,71,72,79]. In mRNA, methylation modifications regulate processes such as nuclear export, splicing, degradation, and translation [18]. The dynamic changes and regulation of these modifications suggest that they may play roles in disease states by influencing gene expression and function. Understanding the specific roles of these modifications in various diseases is essential for revealing the molecular mechanisms of diseases and developing new therapeutic strategies.
RNA methylation modifications have significant potential applications in cancer and other diseases. Particularly in oncology, studies have found that abnormal patterns of RNA modifications are closely associated with tumor initiation, progression, and drug resistance [94,95]. m²G levels are broadly elevated in malignant thyroid tumors, hepatocellular carcinoma, breast cancer and head and neck squamous cell carcinoma, and are linked to enhanced proliferation and migration of tumor cells; loss of the related methyltransferases TRMT11 and THUMPD3 suppresses colon cancer cell proliferation, whereas ALKBH7-mediated conversion of m²2G to m²G promotes cell growth by improving base pairing [91-93]. Serum m²G levels in patients with colorectal cancer and esophageal adenocarcinoma also differ from those in healthy individuals, suggesting that m²G and its modifying enzymes may not only participate in tumorigenic mechanisms, but may also be developed as early screening and prognostic biomarkers in liquid biopsy-based approaches [86]. Cm has mainly been implicated in metabolomics studies, where its altered levels have been associated with the risk and metabolic status of esophageal squamous cell carcinoma, colorectal cancer, clear cell renal cell carcinoma and hepatocellular carcinoma; moreover, the triphosphate form of Cm can inhibit the activity of the HCV RNA polymerase NS5B, providing a potential novel target for antiviral therapy against hepatitis C [99-102].
m⁵U appears to play an even more prominent role in cancer, cardiovascular and immune-related conditions. On the one hand, elevated plasma m⁵U is associated with a higher risk of coronary artery disease and poor prognosis, and has repeatedly emerged as a key metabolite in various disease risk prediction models; on the other hand, imbalances in m⁵U and its related metabolites have been observed in hypertension, post-stroke depression, multiple sclerosis and autoimmune disorders, suggesting that m⁵U participates in the fine-tuning of cardiovascular, neurological and immune regulation [97,108-110]. m⁵U also constitutes an important structural scaffold for several antitumor and anti-HIV nucleoside analogs, and its levels are closely correlated with the degree of immune reconstitution and inflammatory status in individuals with HIV infection [120]. Deficiency of m⁵U methyltransferases such as NSun2 and TRMT2A can induce tsRNA production and alter stress responses, and has been linked to the attenuation of liver injury and to recurrence risk in HER2⁺ breast cancer, further underscoring the potential role of m⁵U in disease pathogenesis [110,122].
ms²i⁶A predominantly occurs in mitochondrial tRNAs and is closely related to energy metabolism. Cdk5RAP1 deficiency leads to reduced ms²i⁶A levels, impairing the translation of mitochondrial respiratory chain subunits and resulting in myocardial and skeletal muscle dysfunction, highlighting its importance in metabolic and cardiovascular diseases [129]. In cancer, Cdk5RAP1 deficiency can, on the one hand, induce cell cycle arrest and apoptosis in breast cancer cells via the ROS/JNK pathway [130], while on the other hand, in gliomas it converts the antitumor modification i⁶A into ms²i⁶A to protect tumor-initiating cells [131], suggesting that ms²i⁶A may exert bidirectional, cell type and context dependent regulatory effects.
In addition, the development of RNA modification detection technologies has provided powerful tools for RNA modification research. Mass spectrometry is now capable of identifying and quantifying single modification sites, while high-throughput sequencing technologies can analyze modification patterns across the transcriptome [68]. Machine learning-based site prediction methods are also being updated continuously [84-86]. These technological advancements not only deepen our understanding of the biological functions of methylation and other modifications but also aid in identifying RNA modification patterns associated with diseases. With the continuous optimization of detection technologies, we expect that future studies will enable RNA modification analysis at the single-cell level, revealing new aspects of cellular heterogeneity in diseases [88].
Overall, current studies indicate that RNA methylation modifications such as m²G, Cm, m⁵U, and ms²i⁶A are not only closely associated with the occurrence, progression, and therapeutic response of various cancers, but also participate in the development and progression of multiple non-neoplastic diseases, including age-related macular degeneration, diabetic retinopathy, epilepsy, viral hepatitis, HIV infection, and autoimmune disorders. These relatively rare RNA methylation modifications and their associated regulatory enzymes therefore hold promise as novel biomarkers for disease diagnosis and prognosis, and may provide potential targets for therapeutic intervention. However, the molecular mechanisms underlying these RNA modifications remain insufficiently understood. The regulatory system of “writers-readers-erasers” has not yet been fully elucidated, and most studies are still limited to metabolite-level observations or correlative analyses. In particular, the precise modification sites, downstream signaling pathways, tissue and cell-type specificity, as well as the synergistic or antagonistic interactions among different RNA modifications remain largely unexplored. Future studies are therefore needed to systematically elucidate the roles of these RNA modifications in gene expression regulation and cell fate determination at the mechanistic level. In addition, integrating high-sensitivity quantitative detection techniques with single-cell sequencing and spatial multi-omics approaches will facilitate a deeper understanding of the dynamic changes of RNA modifications across different cell types and disease microenvironments. Meanwhile, the application of programmable RNA modification editing tools and in vivo disease models will help clarify the functional roles of these modifications in disease development and progression, and evaluate their feasibility and safety as diagnostic biomarkers or potential therapeutic targets in real clinical settings, thereby promoting the translation of RNA modification research toward precision medicine and clinical applications.
Conclusion
This review focuses on four relatively rare RNA methylation marks, m²G, Cm, m⁵U and ms²i⁶A, summarizing their fundamental roles in maintaining RNA structural stability and regulating translation and gene expression, and systematically outlining current evidence linking these modifications to cancer and non-malignant diseases. Available data indicate that these marks are not only involved in the initiation, progression and treatment response of multiple malignancies, but are also closely associated with cardiovascular, metabolic and neurological disorders, as well as viral infections and autoimmune diseases, thereby highlighting their potential as fluid- or tissue-based biomarkers and therapeutic targets. However, compared with classical modifications such as m⁶A, studies on the “writer-reader-eraser” enzyme systems and the precise mechanisms of m²G, Cm, m⁵U and ms²i⁶A remain at an early stage, and most conclusions are still descriptive or correlative in nature. Future work should investigate these modifications in greater mechanistic detail and in larger, well-characterized clinical cohorts to better substantiate their translational potential in precision diagnosis and individualized therapy for cancer and other diseases.
Acknowledgements
The Review Figure created with BioRender.com.
Funding
This study was supported by a grant (grant number: 82360245 to J Min) from the National Natural Science Foundation of China, Beijing, China.
Availability of data and materials
The authors confirm that all data supporting the findings of this study are available in the main manuscript.
Author contributions
KYW and TTN drafted the manuscript and revised the manuscript. QZ compiled and summarized the table and figures. YH and JB were responsible for literature selection and retrieval. JM provided critical revision of the manuscript and supported the project through funding. All authors approved the final draft.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Wang T, Kong S, Tao M. et al. The potential role of RNA N6-methyladenosine in Cancer progression. Mol Cancer. 2020;19(1):88 doi:10.1186/s12943-020-01204-7
2. Chen M, Chen Y, Wang K. et al. Non-m6A RNA modifications in haematological malignancies. Clin Transl Med. 2024;14(6):e1666 doi:10.1002/ctm2.1666
3. Yu L, Xu H, Xiong H. et al. The role of m5C RNA modification in cancer development and therapy. Heliyon. 2024;10(19):e38660 doi:10.1016/j.heliyon.2024.e38660
4. Birnstiel ML, Fleissner E, Borek E. NUCLEOLUS: A CENTER OF RNA METHYLATION. Science. 1963;142(3599):1577-1580 doi:10.1126/science.142.3599.1577
5. Yang Y, Fan X, Mao M. et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 2017;27(5):626-641 doi:10.1038/cr.2017.31
6. He RZ, Jiang J, Luo DX. The functions of N6-methyladenosine modification in lncRNAs. Genes Dis. 2020;7(4):598-605 doi:10.1016/j.gendis.2020.03.005
7. Ranaei-Siadat E, Fabret C, Seijo B. et al. RNA-methyltransferase TrmA is a dual-specific enzyme responsible for C5-methylation of uridine in both tmRNA and tRNA. RNA Biol. 2013;10(4):572-578 doi:10.4161/rna.24327
8. Pendleton KE, Chen B, Liu K. et al. The U6 snRNA m6A Methyltransferase METTL16 Regulates SAM Synthetase Intron Retention. Cell. 2017;169(5):824-835.e14 doi:10.1016/j.cell.2017.05.003
9. Motorin Y, Helm M. RNA nucleotide methylation. Wiley Interdiscip Rev RNA. 2011;2(5):611-631 doi:10.1002/wrna.79
10. Gilbert WV, Bell TA, Schaening C. Messenger RNA modifications: Form, distribution, and function. Science. 2016;352(6292):1408-1412 doi:10.1126/science.aad8711
11. Frye M, Harada BT, Behm M. et al. RNA modifications modulate gene expression during development. Science. 2018;361(6409):1346-1349 doi:10.1126/science.aau1646
12. Kealey JT, Gu X, Santi DV. Enzymatic mechanism of tRNA (m5U54)methyltransferase. Biochimie. 1994;76(12):1133-1142 doi:10.1016/0300-9084(94)90042-6
13. Bourgeois G, Marcoux J, Saliou JM. et al. Activation mode of the eukaryotic m2G10 tRNA methyltransferase Trm11 by its partner protein Trm112. Nucleic Acids Res. 2017;45(4):1971-1982 doi:10.1093/nar/gkw1271
14. Xue C, Zhao Y, Li L. Advances in RNA cytosine-5 methylation: detection, regulatory mechanisms, biological functions and links to cancer. Biomark Res. 2020;8:43 doi:10.1186/s40364-020-00225-0
15. Díaz I, Ehrenberg M. ms2i6A deficiency enhances proofreading in translation. J Mol Biol. 1991;222(4):1161-1171 doi:10.1016/0022-2836(91)90599-2
16. Davanloo P, Sprinzl M, Watanabe K. et al. Role of ribothymidine in the thermal stability of transfer RNA as monitored by proton magnetic resonance. Nucleic Acids Res. 1979;6(4):1571-1581 doi:10.1093/nar/6.4.1571
17. Zhao BS, Roundtree IA, He C. Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Biol. 2017;18(1):31-42 doi:10.1038/nrm.2016.132
18. Wang C, Ulryck N, Herzel L. et al. N 2-methylguanosine modifications on human tRNAs and snRNA U6 are important for cell proliferation, protein translation and pre-mRNA splicing. Nucleic Acids Research. 2023;51(14):7496-7519 doi:10.1093/nar/gkad487
19. Yang WQ, Ge JY, Zhang X. et al. THUMPD2 catalyzes the N2-methylation of U6 snRNA of the spliceosome catalytic center and regulates pre-mRNA splicing and retinal degeneration. Nucleic Acids Res. 2024;52(6):3291-3309 doi:10.1093/nar/gkad1243
20. de Crécy-Lagard V, Boccaletto P, Mangleburg CG. et al. Matching tRNA modifications in humans to their known and predicted enzymes. Nucleic Acids Res. 2019;47(5):2143-2159 doi:10.1093/nar/gkz011
21. Lu L. Mobilizing ER IP3 receptors as a mechanism to enhance calcium signaling. Cell Mol Immunol. 2021;18(9):2284-2285 doi:10.1038/s41423-021-00725-5
22. Purushothaman SK, Bujnicki JM, Grosjean H. et al. Trm11p and Trm112p Are both Required for the Formation of 2-Methylguanosine at Position 10 in Yeast tRNA. Molecular and Cellular Biology. 2005;25(11):4359-4370 doi:10.1128/MCB.25.11.4359-4370.2005
23. Yang WQ, Xiong QP, Ge JY. et al. THUMPD3-TRMT112 is a m2G methyltransferase working on a broad range of tRNA substrates. Nucleic Acids Res. 2021;49(20):11900-11919 doi:10.1093/nar/gkab927
24. Wang SE, Wu FY, Shin I. et al. Transforming growth factor {beta} (TGF-{beta})-Smad target gene protein tyrosine phosphatase receptor type kappa is required for TGF-{beta} function. Mol Cell Biol. 2005;25(11):4703-4715 doi:10.1128/MCB.25.11.4703-4715.2005
25. Hirata A, Suzuki T, Nagano T. et al. Distinct modified nucleosides in tRNATrp from the hyperthermophilic archaeon thermococcus kodakarensis and requirement of tRNA m2G10/m22G10 methyltransferase (archaeal Trm11) for survival at high temperatures. J Bacteriol. 2019;201(21):e00448-19 doi:10.1128/JB.00448-19
26. Zhang LS, Xiong QP, Peña Perez S. et al. ALKBH7-mediated demethylation regulates mitochondrial polycistronic RNA processing. Nat Cell Biol. 2021;23(7):684-691 doi:10.1038/s41556-021-00709-7
27. Cantara WA, Crain PF, Rozenski J. et al. The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Res. 2011;39(Database issue):D195-201 doi:10.1093/nar/gkq1028
28. Limbach PA, Crain PF, McCloskey JA. Summary: the modified nucleosides of RNA. Nucleic Acids Res. 1994;22(12):2183-2196 doi:10.1093/nar/22.12.2183
29. Boccaletto P, Stefaniak F, Ray A. et al. MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res. 2022;50(D1):D231-D235 doi:10.1093/nar/gkab1083
30. Yamada Y, Ishikura H. Identification of a modified nucleoside in Escherichia coli tRNA1Ser as 2'-O-methylcytidine. Biochim Biophys Acta. 1975;402(3):285-287 doi:10.1016/0005-2787(75)90265-8
31. Ladner JE, Jack A, Robertus JD. et al. Structure of yeast phenylalanine transfer RNA at 2.5 A resolution. Proc Natl Acad Sci U S A. 1975;72(11):4414-4418 doi:10.1073/pnas.72.11.4414
32. al-Arif A, Sporn MB. 2'-O-methylation of adenosine, guanosine, uridine, and cytidine in RNA of isolated rat liver nuclei. Proc Natl Acad Sci U S A. 1972;69(7):1716-1719 doi:10.1073/pnas.69.7.1716
33. Heckle WL, Fenton RG, Wood TG. et al. Methylated nucleosides in globin mRNA from mouse nucleated erythroid cells. J Biol Chem. 1977;252(5):1764-1770
34. Purta E, van Vliet F, Tkaczuk KL. et al. The yfhQ gene of Escherichia coli encodes a tRNA:Cm32/Um32 methyltransferase. BMC Mol Biol. 2006;7:23 doi:10.1186/1471-2199-7-23
35. Benítez-Páez A, Villarroya M, Douthwaite S. et al. YibK is the 2'-O-methyltransferase TrmL that modifies the wobble nucleotide in Escherichia coli tRNA(Leu) isoacceptors. RNA. 2010;16(11):2131-2143 doi:10.1261/rna.2245910
36. Guy MP, Podyma BM, Preston MA. et al. Yeast Trm7 interacts with distinct proteins for critical modifications of the tRNAPhe anticodon loop. RNA. 2012;18(10):1921-1933 doi:10.1261/rna.035287.112
37. Ayadi L, Galvanin A, Pichot F. et al. RNA ribose methylation (2'-O-methylation): Occurrence, biosynthesis and biological functions. Biochim Biophys Acta Gene Regul Mech. 2019;1862(3):253-269 doi:10.1016/j.bbagrm.2018.11.009
38. Renalier MH, Joseph N, Gaspin C. et al. The Cm56 tRNA modification in archaea is catalyzed either by a specific 2'-O-methylase, or a C/D sRNP. RNA. 2005;11(7):1051-1063 doi:10.1261/rna.2110805
39. Hibma JL, Munson LM, Jones JD. et al. TlyA is a 23S and 16S 2′-O-methylcytidine methyltransferase important for ribosome assembly in bacillus subtilis. bioRxiv. Published online April 25, 2025:2025.04.21.649808. doi:10.1101/2025.04.21.649808.
40. Matsuda T, Yamagami R, Ihara A. et al. A transfer RNA methyltransferase with an unusual domain composition catalyzes 2′-O-methylation at position 6 in tRNA. Nucleic Acids Res. 2025;53(13):gkaf579 doi:10.1093/nar/gkaf579
41. Agris PF, Vendeix FAP, Graham WD. tRNA's wobble decoding of the genome: 40 years of modification. J Mol Biol. 2007;366(1):1-13 doi:10.1016/j.jmb.2006.11.046
42. Björk GR, Durand JM, Hagervall TG. et al. Transfer RNA modification: influence on translational frameshifting and metabolism. FEBS Lett. 1999;452(1-2):47-51 doi:10.1016/s0014-5793(99)00528-1
43. Urbonavicius J, Qian Q, Durand JM. et al. Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J. 2001;20(17):4863-4873 doi:10.1093/emboj/20.17.4863
44. Yarian C, Townsend H, Czestkowski W. et al. Accurate translation of the genetic code depends on tRNA modified nucleosides. J Biol Chem. 2002;277(19):16391-16395 doi:10.1074/jbc.M200253200
45. Kawai G, Yamamoto Y, Kamimura T. et al. Conformational rigidity of specific pyrimidine residues in tRNA arises from posttranscriptional modifications that enhance steric interaction between the base and the 2'-hydroxyl group. Biochemistry. 1992;31(4):1040-1046 doi:10.1021/bi00119a012
46. He CC, Hamlow LA, Zhu Y. et al. Structural and Energetic Effects of O2'-Ribose Methylation of Protonated Pyrimidine Nucleosides. J Am Soc Mass Spectrom. 2019;30(11):2318-2334 doi:10.1007/s13361-019-02300-9
47. Dominissini D, Moshitch-Moshkovitz S, Schwartz S. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485(7397):201-206 doi:10.1038/nature11112
48. Inoue H, Hayase Y, Imura A. et al. Synthesis and hybridization studies on two complementary nona(2'-O-methyl)ribonucleotides. Nucleic Acids Res. 1987;15(15):6131-6148 doi:10.1093/nar/15.15.6131
49. Noon KR, Bruenger E, McCloskey JA. Posttranscriptional modifications in 16S and 23S rRNAs of the archaeal hyperthermophile Sulfolobus solfataricus. J Bacteriol. 1998;180(11):2883-2888 doi:10.1128/JB.180.11.2883-2888.1998
50. Satoh A, Takai K, Ouchi R. et al. Effects of anticodon 2'-O-methylations on tRNA codon recognition in an Escherichia coli cell-free translation. RNA. 2000;6(5):680-686 doi:10.1017/s1355838200000029
51. Pillay DT, Guillemaut P, Weil JH. Nucleotide sequences of three soybean chloroplast tRNAsLeu and re-examination of bean chloroplast tRNA2Leu sequence. Nucleic Acids Res. 1984;12(6):2997-3001 doi:10.1093/nar/12.6.2997
52. Païs de Barros JP, Keith G, El Adlouni C. et al. 2'-O-methyl-5-formylcytidine (f5Cm), a new modified nucleotide at the “wobble” of two cytoplasmic tRNAs Leu (NAA) from bovine liver. Nucleic Acids Res. 1996;24(8):1489-1496 doi:10.1093/nar/24.8.1489
53. Sprinzl M, Hartmann T, Meissner F. et al. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 1987 15 Suppl(Suppl):r53-188. doi:10.1093/nar/15.suppl.r53
54. Horie N, Yamaizumi Z, Kuchino Y. et al. Modified nucleosides in the first positions of the anticodons of tRNA(Leu)4 and tRNA(Leu)5 from Escherichia coli. Biochemistry. 1999;38(1):207-217 doi:10.1021/bi981865g
55. Pixa G, Dirheimer G, Keith G. Sequence of tRNALeu CmAA from Bacillus stearothermophilus. Biochem Biophys Res Commun. 1983;112(2):578-585 doi:10.1016/0006-291x(83)91503-6
56. Murasugi A, Takemura S. Nucleotide sequence of leucine transfer RNA 1 from Candida (Torulopsis) utilis. J Biochem. 1978;83(4):1029-1038 doi:10.1093/oxfordjournals.jbchem.a131991
57. Glasser AL, el Adlouni C, Keith G. et al. Presence and coding properties of 2'-O-methyl-5-carbamoylmethyluridine (ncm5Um) in the wobble position of the anticodon of tRNA(Leu) (U*AA) from brewer's yeast. FEBS Lett. 1992;314(3):381-385 doi:10.1016/0014-5793(92)81510-s
58. Björk GR, Neidhardt FC. Physiological and biochemical studies on the function of 5-methyluridine in the transfer ribonucleic acid of Escherichia coli. J Bacteriol. 1975;124(1):99-111 doi:10.1128/jb.124.1.99-111.1975
59. Felden B, Hanawa K, Atkins JF. et al. Presence and location of modified nucleotides in Escherichia coli tmRNA: structural mimicry with tRNA acceptor branches. EMBO J. 1998;17(11):3188-3196 doi:10.1093/emboj/17.11.3188
60. Cheng QY, Xiong J, Ma CJ. et al. Chemical tagging for sensitive determination of uridine modifications in RNA. Chem Sci. 2020;11(7):1878-1891 doi:10.1039/c9sc05094a
61. Jones JD, Franco MK, Smith TJ. et al. Methylated guanosine and uridine modifications in S. cerevisiae mRNAs modulate translation elongation. RSC Chem Biol. 2023;4(5):363-378 doi:10.1039/d2cb00229a
62. Nordlund ME, Johansson JO, von Pawel-Rammingen U. et al. Identification of the TRM2 gene encoding the tRNA(m5U54)methyltransferase of Saccharomyces cerevisiae. RNA. 2000;6(6):844-860 doi:10.1017/s1355838200992422
63. Urbonavicius J, Skouloubris S, Myllykallio H. et al. Identification of a novel gene encoding a flavin-dependent tRNA:m5U methyltransferase in bacteria-evolutionary implications. Nucleic Acids Res. 2005;33(13):3955-3964 doi:10.1093/nar/gki703
64. Urbonavicius J, Brochier-Armanet C, Skouloubris S. et al. In vitro detection of the enzymatic activity of folate-dependent tRNA (Uracil-54,-C5)-methyltransferase: evolutionary implications. Methods Enzymol. 2007;425:103-119 doi:10.1016/S0076-6879(07)25004-9
65. Desmolaize B, Fabret C, Brégeon D. et al. A single methyltransferase YefA (RlmCD) catalyses both m5U747 and m5U1939 modifications in Bacillus subtilis 23S rRNA. Nucleic Acids Res. 2011;39(21):9368-9375 doi:10.1093/nar/gkr626
66. Madsen CT, Mengel-Jørgensen J, Kirpekar F. et al. Identifying the methyltransferases for m(5)U747 and m(5)U1939 in 23S rRNA using MALDI mass spectrometry. Nucleic Acids Res. 2003;31(16):4738-4746 doi:10.1093/nar/gkg657
67. Urbonavicius J, Jäger G, Björk GR. Amino acid residues of the Escherichia coli tRNA(m5U54)methyltransferase (TrmA) critical for stability, covalent binding of tRNA and enzymatic activity. Nucleic Acids Res. 2007;35(10):3297-3305 doi:10.1093/nar/gkm205
68. Carter JM, Emmett W, Mozos IR. et al. FICC-Seq: a method for enzyme-specified profiling of methyl-5-uridine in cellular RNA. Nucleic Acids Res. 2019;47(19):e113 doi:10.1093/nar/gkz658
69. Powell CA, Minczuk M. TRMT2B is responsible for both tRNA and rRNA m(5)U-methylation in human mitochondria. RNA Biol. 2020;17(4):451-462 doi:10.1080/15476286.2020.1712544
70. Towns WL, Begley TJ. Transfer RNA methytransferases and their corresponding modifications in budding yeast and humans: activities, predications, and potential roles in human health. DNA Cell Biol. 2012;31(4):434-454 doi:10.1089/dna.2011.1437
71. Jones JD, Franco MK, Tardu M. et al. Conserved 5-methyluridine tRNA modification modulates ribosome translocation. bioRxiv. Published online November 12, 2023:2023.11.12.566704. doi:10.1101/2023.11.12.566704.
72. Schultz SK, Katanski CD, Halucha M. et al. Modifications in the T arm of tRNA globally determine tRNA maturation, function, and cellular fitness. Proc Natl Acad Sci U S A. 2024;121(26):e2401154121 doi:10.1073/pnas.2401154121
73. Jones JD, Franco MK, Giles RN. et al. Conserved 5-methyluridine tRNA modification modulates ribosome translocation. Proc Natl Acad Sci U S A. 2024;121(35):e2401743121 doi:10.1073/pnas.2401743121
74. von Pawel-Rammingen U, Aström S, Byström AS. Mutational analysis of conserved positions potentially important for initiator tRNA function in Saccharomyces cerevisiae. Mol Cell Biol. 1992;12(4):1432-1442 doi:10.1128/mcb.12.4.1432-1442.1992
75. Keller P, Freund I, Marchand V. et al. Double methylation of tRNA-U54 to 2'-O-methylthymidine (Tm) synergistically decreases immune response by Toll-like receptor 7. Nucleic Acids Res. 2018;46(18):9764-9775 doi:10.1093/nar/gky644
76. Fakruddin M, Wei FY, Emura S. et al. Cdk5rap1-mediated 2-methylthio-N6-isopentenyladenosine modification is absent from nuclear-derived RNA species. Nucleic Acids Res. 2017;45(20):11954-11961 doi:10.1093/nar/gkx819
77. Moore JA, Poulter CD. Escherichia coli dimethylallyl diphosphate:tRNA dimethylallyltransferase: a binding mechanism for recombinant enzyme. Biochemistry. 1997;36(3):604-614 doi:10.1021/bi962225l
78. Pierrel F, Hernandez HL, Johnson MK. et al. MiaB protein from Thermotoga maritima. Characterization of an extremely thermophilic tRNA-methylthiotransferase. J Biol Chem. 2003;278(32):29515-29524 doi:10.1074/jbc.M301518200
79. Petrullo LA, Gallagher PJ, Elseviers D. The role of 2-methylthio-N6-isopentenyladenosine in readthrough and suppression of nonsense codons in Escherichia coli. Mol Gen Genet. 1983;190(2):289-294 doi:10.1007/BF00330653
80. Wang C, van Tran N, Jactel V. et al. Structural and functional insights into Archaeoglobus fulgidus m2G10 tRNA methyltransferase Trm11 and its Trm112 activator. Nucleic Acids Res. 2020;48(19):11068-11082 doi:10.1093/nar/gkaa830
81. Fu L, Amato NJ, Wang P. et al. Simultaneous Quantification of Methylated Cytidine and Adenosine in Cellular and Tissue RNA by Nano-Flow Liquid Chromatography-Tandem Mass Spectrometry Coupled with the Stable Isotope-Dilution Method. Anal Chem. 2015;87(15):7653-7659 doi:10.1021/acs.analchem.5b00951
82. Fang Z, Hu Y, Hong X. et al. Simultaneous Determination of Methylated Nucleosides by HILIC-MS/MS Revealed Their Alterations in Urine from Breast Cancer Patients. Metabolites. 2022;12(10):973 doi:10.3390/metabo12100973
83. Kogaki T, Ohshio I, Ura H. et al. Development of a highly sensitive method for the quantitative analysis of modified nucleosides using UHPLC-UniSpray-MS/MS. J Pharm Biomed Anal. 2021;197:113943 doi:10.1016/j.jpba.2021.113943
84. Shaon MSH, Karim T, Ali MM. et al. A robust deep learning approach for identification of RNA 5-methyluridine sites. Sci Rep. 2024;14(1):25688 doi:10.1038/s41598-024-76148-9
85. Noor S, Naseem A, Awan HH. et al. Deep-m5U: a deep learning-based approach for RNA 5-methyluridine modification prediction using optimized feature integration. BMC Bioinformatics. 2024;25(1):360 doi:10.1186/s12859-024-05978-1
86. Ji Y, Sun J, Xie J. et al. m5UMCB: Prediction of RNA 5-methyluridine sites using multi-scale convolutional neural network with BiLSTM. Comput Biol Med. 2024;168:107793 doi:10.1016/j.compbiomed.2023.107793
87. Brūmele B, Mutso M, Telanne L. et al. Human TRMT112-Methyltransferase Network Consists of Seven Partners Interacting with a Common Co-Factor. Int J Mol Sci. 2021;22(24):13593 doi:10.3390/ijms222413593
88. Clark KD, Rubakhin SS, Sweedler JV. Single-Neuron RNA Modification Analysis by Mass Spectrometry: Characterizing RNA Modification Patterns and Dynamics with Single-Cell Resolution. Anal Chem. 2021;93(43):14537-14544 doi:10.1021/acs.analchem.1c03507
89. Heiss M, Reichle VF, Kellner S. Observing the fate of tRNA and its modifications by nucleic acid isotope labeling mass spectrometry: NAIL-MS. RNA Biol. 2017;14(9):1260-1268 doi:10.1080/15476286.2017.1325063
90. Djukovic D, Baniasadi HR, Kc R. et al. Targeted serum metabolite profiling of nucleosides in esophageal adenocarcinoma. Rapid Commun Mass Spectrom. 2010;24(20):3057-3062 doi:10.1002/rcm.4739
91. Fang Z, Lu Z, Han S. et al. The transcriptome-wide mapping of 2-methylthio-N6-isopentenyladenosine at single-base resolution. J Am Chem Soc. 2023;145(9):5467-5473 doi:10.1021/jacs.2c13894
92. You XJ, Liu T, Ma CJ. et al. Determination of RNA hydroxylmethylation in mammals by mass spectrometry analysis. Anal Chem. 2019;91(16):10477-10483 doi:10.1021/acs.analchem.9b01318
93. Zheng W, Wang M, Chai X. et al. Targeted metabolomics analysis of nucleosides and the identification of biomarkers for colorectal adenomas and colorectal cancer. Front Mol Biosci. 2023;10:1163089 doi:10.3389/fmolb.2023.1163089
94. Chen K, Shen D, Tan L. et al. A Pan-Cancer Analysis Reveals the Prognostic and Immunotherapeutic Value of ALKBH7. Front Genet. 2022;13:822261 doi:10.3389/fgene.2022.822261
95. Klimontova M, Zhang H, Campos-Laborie F. et al. THUMPD3 regulates alternative splicing of ECM transcripts in human lung cancer cells and promotes proliferation and migration. PLoS One. 2024;19(12):e0314655 doi:10.1371/journal.pone.0314655
96. Keith G, Heyman T. Heterogeneities in vertebrate tRNAs(Trp) avian retroviruses package only as a primer the tRNA(Trp) lacking modified m2G in position 7. Nucleic Acids Res. 1990;18(4):703-710 doi:10.1093/nar/18.4.703
97. Li X, Zhao L, Wei M. et al. Serum metabolomics analysis for the progression of esophageal squamous cell carcinoma. J Cancer. 2021;12(11):3190-3197 doi:10.7150/jca.54429
98. Fang Z, Hu Y, Chen J. et al. Mass Spectrometry-Based Targeted Serum Monomethylated Ribonucleosides Profiling for Early Detection of Breast Cancer. Front Mol Biosci. 2021;8:741603 doi:10.3389/fmolb.2021.741603
99. Neef SK, Janssen N, Winter S. et al. Metabolic Drug Response Phenotyping in Colorectal Cancer Organoids by LC-QTOF-MS. Metabolites. 2020;10(12):494 doi:10.3390/metabo10120494
100. McCullough ML, Hodge RA, Campbell PT. et al. Pre-Diagnostic Circulating Metabolites and Colorectal Cancer Risk in the Cancer Prevention Study-II Nutrition Cohort. Metabolites. 2021;11(3):156 doi:10.3390/metabo11030156
101. Mohl DA, Lagies S, Zodel K. et al. Integrated Metabolomic and Transcriptomic Analysis of Modified Nucleosides for Biomarker Discovery in Clear Cell Renal Cell Carcinoma. Cells. 2023;12(8):1102 doi:10.3390/cells12081102
102. Wu H, Lin H, Long F. et al. Urinary metabolomics-driven discovery of metabolic markers and molecular subtyping in liver cancer. Front Biosci (Landmark Ed). 2025;30(10):45438 doi:10.31083/FBL45438
103. Carroll SS, Tomassini JE, Bosserman M. et al. Inhibition of hepatitis C virus RNA replication by 2'-modified nucleoside analogs. J Biol Chem. 2003;278(14):11979-11984 doi:10.1074/jbc.M210914200
104. Yao C, Lv D, Zhou X. et al. Exploring urinary modified nucleosides as biomarkers for diabetic retinopathy: Development and validation of a ultra performance liquid chromatography-tandem mass spectrometry method. J Chromatogr B Analyt Technol Biomed Life Sci. 2024;1232:123968 doi:10.1016/j.jchromb.2023.123968
105. Liu J, Li R, Yu H. et al. Metabolite Associations with Childhood and Juvenile Absence Epilepsy: A Bidirectional Mendelian Randomization Study. Psychiatry Clin Psychopharmacol. 2024;35(1):14-21 doi:10.5152/pcp.2024.24951
106. Furset G, Fløisand Y, Sioud M. Impaired expression of indoleamine 2, 3-dioxygenase in monocyte-derived dendritic cells in response to Toll-like receptor-7/8 ligands. Immunology. 2008;123(2):263-271 doi:10.1111/j.1365-2567.2007.02695.x
107. Sioud M. Deciphering the code of innate immunity recognition of siRNAs. Methods Mol Biol. 2009;487:41-59 doi:10.1007/978-1-60327-547-7_2
108. Sioud M. Does the understanding of immune activation by RNA predict the design of safe siRNAs? Front Biosci. 2008;13:4379-4392 doi:10.2741/3011
109. Zhu Q, Qin M, Wang Z. et al. Plasma metabolomics provides new insights into the relationship between metabolites and outcomes and left ventricular remodeling of coronary artery disease. Cell Biosci. 2022;12(1):173 doi:10.1186/s13578-022-00863-x
110. Shi M, He J, Li C. et al. Metabolomics study of blood pressure salt-sensitivity and hypertension. Nutr Metab Cardiovasc Dis. 2022;32(7):1681-1692 doi:10.1016/j.numecd.2022.04.002
111. Bullinger D, Neubauer H, Fehm T. et al. Metabolic signature of breast cancer cell line MCF-7: profiling of modified nucleosides via LC-IT MS coupling. BMC Biochem. 2007;8(1):25 doi:10.1186/1471-2091-8-25
112. Jiang W, Gong L, Liu F. et al. Alteration of Gut Microbiome and Correlated Lipid Metabolism in Post-Stroke Depression. Front Cell Infect Microbiol. 2021;11:663967 doi:10.3389/fcimb.2021.663967
113. de Fátima Cobre A, Surek M, Stremel DP. et al. Diagnosis and prognosis of COVID-19 employing analysis of patients' plasma and serum via LC-MS and machine learning. Comput Biol Med. 2022;146:105659 doi:10.1016/j.compbiomed.2022.105659
114. Song S, Zhang Q, Yu J. A mendelian randomization study investigating the causal relationships between 1400 serum metabolites and autoimmune diseases. Heliyon. 2024;10(14):e34560 doi:10.1016/j.heliyon.2024.e34560
115. Rocha I, de Brito IR, Carvalho HF. et al. Modified nucleosides as potential biomarkers of prostate cancer: Targeted metabolomics of in vitro cell samples by MEKC-UV. Electrophoresis. 2025;46(13-14):1006-1013 doi:10.1002/elps.8120
116. Zeng X, Zeng Q, Wang X. et al. Causal association between 1400 metabolites and dilated cardiomyopathy: A bidirectional two-sample mendelian randomization analysis. Front Endocrinol (Lausanne). 2024;15:1423142 doi:10.3389/fendo.2024.1423142
117. Zhu LH, Jin ZX, Ma YQ. et al. Untargeted metabolomics analysis of metabolite changes in gastric cancer patients from plateau regions. World J Gastrointest Oncol. 2025;17(9):109777 doi:10.4251/wjgo.v17.i9.109777
118. Monoe Y, Miyamoto S, Jingushi K. et al. Hypoxia regulates tumour characteristic RNA modifications in ovarian cancers. FEBS J. 2023;290(8):2085-2096 doi:10.1111/febs.16688
119. Schlimpert M, Lagies S, Budnyk V. et al. Metabolic Phenotyping of Anks3 Depletion in mIMCD-3 cells - a Putative Nephronophthisis Candidate. Sci Rep. 2018;8(1):9022 doi:10.1038/s41598-018-27389-y
120. Berg M, Li C, Kaiser S. NAIL-MS reveals tRNA and rRNA hypomodification as a consequence of 5-fluorouracil treatment. Nucleic Acids Res. 2025;53(4):gkaf090 doi:10.1093/nar/gkaf090
121. Wan LY, Lam SM, Huang HH. et al. Multi-omics dissection of metabolic dysregulation associated with immune recovery in people living with HIV-1. J Transl Med. 2025;23(1):143 doi:10.1186/s12967-025-06168-0
122. Guo G, Wang H, Shi X. et al. Disease activity-associated alteration of mRNA m5 C methylation in CD4+ T cells of systemic lupus erythematosus. Front Cell Dev Biol. 2020;8:430 doi:10.3389/fcell.2020.00430
123. Ye X, Qiu R, He X. et al. Interferon therapy for chronic hepatitis B virus infection affects nucleoside metabolism: A metabolomics study. BMC Gastroenterol. 2025;25(1):523 doi:10.1186/s12876-025-04126-0
124. Yang Z, Sun B, Xiang J. et al. Role of epigenetic modification in interferon treatment of hepatitis B virus infection. Front Immunol. 2022;13:1018053 doi:10.3389/fimmu.2022.1018053
125. Ding WB, Wang MC, Yu J. et al. HBV/pregenomic RNA increases the stemness and promotes the development of HBV-related HCC through reciprocal regulation with insulin-like growth factor 2 mRNA-binding protein 3. Hepatology. 2021;74(3):1480-1495 doi:10.1002/hep.31850
126. Imam H, Kim GW, Mir SA. et al. Interferon-stimulated gene 20 (ISG20) selectively degrades N6-methyladenosine modified hepatitis B virus transcripts. PLoS Pathog. 2020;16(2):e1008338 doi:10.1371/journal.ppat.1008338
127. Ying S, Li P, Wang J. et al. tRF-Gln-CTG-026 ameliorates liver injury by alleviating global protein synthesis. Signal Transduct Target Ther. 2023;8(1):144 doi:10.1038/s41392-023-01351-5
128. Pereira M, Ribeiro DR, Pinheiro MM. et al. m(5)U54 tRNA Hypomodification by Lack of TRMT2A Drives the Generation of tRNA-Derived Small RNAs. Int J Mol Sci. 2021 22(6). doi:10.3390/ijms22062941
129. Hicks DG, Janarthanan BR, Vardarajan R. et al. The expression of TRMT2A, a novel cell cycle regulated protein, identifies a subset of breast cancer patients with HER2 over-expression that are at an increased risk of recurrence. BMC Cancer. 2010;10:108 doi:10.1186/1471-2407-10-108
130. Wei FY, Zhou B, Suzuki T. et al. Cdk5rap1-mediated 2-methylthio modification of mitochondrial tRNAs governs protein translation and contributes to myopathy in mice and humans. Cell Metab. 2015;21(3):428-442 doi:10.1016/j.cmet.2015.01.019
131. Wang H, Wei L, Li C. et al. CDK5RAP1 deficiency induces cell cycle arrest and apoptosis in human breast cancer cell line by the ROS/JNK signaling pathway. Oncol Rep. 2015;33(3):1089-1096 doi:10.3892/or.2015.3736
132. Yamamoto T, Fujimura A, Wei FY. et al. 2-Methylthio Conversion of N6-Isopentenyladenosine in Mitochondrial tRNAs by CDK5RAP1 Promotes the Maintenance of Glioma-Initiating Cells. iScience. 2019;21:42-56 doi:10.1016/j.isci.2019.10.012
Author contact
Corresponding author: Jia Min, Department of Anesthesiology, The First Affiliated Hospital of Nanchang University, No. 17, Yongwaizheng Street, Nanchang, 330006, Jiangxi Province, PR China. (Email: ndyfy02136edu.cn).

 Global reach, higher impact
Global reach, higher impact