Impact Factor ISSN: 1449-1907
Int J Med Sci 2026; 23(4):1535-1553. doi:10.7150/ijms.128450 This issue Cite
Review
The Multi-dimensional Mechanisms and Transformation Prospects of the Intratumoral Microbiota-Arginine Metabolism Axis in Tumor Progression and Immune Regulation
1. Department of Obstetrics and Gynecology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, 430030, China.
2. Clinical Research Center of Cancer Immunotherapy, Hubei, 430022, Wuhan, China.
* These authors contributed equally to this work.
Received 2025-11-16; Accepted 2026-3-2; Published 2026-3-17
Abstract
The intratumoral microbiota, as an important component of the tumor microenvironment (TME), impact tumor progression by regulating the arginine-ornithine metabolic axis. It has become a new frontier in tumor research. Arginine is a crucial amino acid in TME, and its metabolites, ornithine and polyamines, directly promote tumor proliferation and induce immunosuppression.
Intratumoral microbiota mainly exert their effects through two direct pathways: 1) arginine depletion, such as Streptococcus in gastric cancer. Specific intratumoral microbiota highly express arginine deiminase (ADI) or arginase (Arg) to consume arginine in the TME, leading to T cell dysfunction and enhancing immunosuppressive cells. 2) Ornithine/polyamines supplement, such as fusobacteria in esophageal cancer produce putrescine. The microbiota converts arginine into ornithine, which is then synthesized into polyamines, directly stimulating tumor cell proliferation and reshaping the immunosuppressive TME. Additionally, the metabolic products from the microbiota like short-chain fatty acids (SCFAs) and indole substances, can amplify these effects through signaling pathways including G protein-coupled receptor 43 (GPR43) and aryl hydrocarbon receptors (AHR).
The regulation of intratumoral microbiota-arginine metabolism axis has a “double-edged sword” characteristic, relying on the metabolic dependence of the different tumors, which provides a basis for precise treatment. Furthermore, strategies targeting the axis present great potential, including Arg1 inhibitors (CB-1158) in combination with immunotherapy, engineered probiotics to supply arginine and inhibit polyamine synthesis in situ within the TME. These advancements also indicate there is enormous progress from exploring the intratumoral microbiota-metabolism interaction to developing novel tumor microecological therapies.
Keywords: Intratumoral microbiota, Arginine, Metabolism, Tumor microenvironment, Immunity, Transformation
1. Introduction
Metabolic reprogramming and immune regulation within the tumor microenvironment (TME) are significant driving factors for tumor occurrence and development, and the intratumoral microbiota were discovered as a potential sight to this field in recent years [1]. Traditional biology holds that tumors create a sterile environment, however, recent high-throughput sequencing and spatial imaging reshaped this view [2]. The distribution of the intratumoral microbiota exhibits significant heterogeneity, and its enrichment is positively correlated with the location and exposure to the external environment [2]. Cancers exposed to outside present the more abundant and diverse intratumoral microbiota, such as melanoma, colorectal cancer (CRC), and lung cancer [2-5]. Nevertheless, increasing evidence indicated that certain microbiota also exists in internal tumors including those in the liver, pancreas and breast [6-8]. These microbiota may enter the TME through hematogenous or lymphatic metastasis with the damaged barrier. Also, there are low-biomass microbial DNA and lipopolysaccharide (LPS) signals within brain tumors protected by the blood-brain barrier like glioblastoma (GBM) [9, 10], which suggest that intratumoral microbiota is far more extensive than previously thought. These intratumoral microbiota further profoundly affect the nutrient supply and signal transduction in the TME through metabolism and secreted products, thereby influencing immune surveillance and tumor progression [11, 12].
Arginine serves as a joint among protein metabolism, polyamine production, nitric oxide (NO) synthesis and the urea cycle, determining the cellular proliferation and function [13]. The arginine-ornithine metabolic pathway as a crucial link connects the functions of immune cells and the proliferation of tumor cells [13-15]. Importantly, although T cells have the capacity for arginine synthesis, their metabolic demand exceeds synthesis in the activated state, hence exogenous uptake is necessary for proliferation and effector functions. Also, ornithine and its derivatives polyamines act as important regulatory factors for cell proliferation and differentiation [13, 15, 16]. Due to its characteristics in metabolism and immune, arginine is crucial for microbial regulation compared to other natural amino acids. Tumor cells often upregulate arginine metabolic related enzymes including arginase (Arg) and ornithine decarboxylase (ODC), to serve for their rapid growth with inducing immunosuppression. For example, the immunosuppressive tumor-associated macrophages (TAM) highly express ARG1 and consume arginine [13, 17]. It is worth noting that the intratumoral microbiota also participates in arginine and ornithine metabolic regulation [18]. Numerous studies have shown that certain intratumoral bacteria can efficiently decompose arginine or synthesize ornithine and polyamine metabolites, thereby locally construct a TAM favorable for the tumor, which inhibit anti-tumor immunity and promote tumor growth [18, 19].
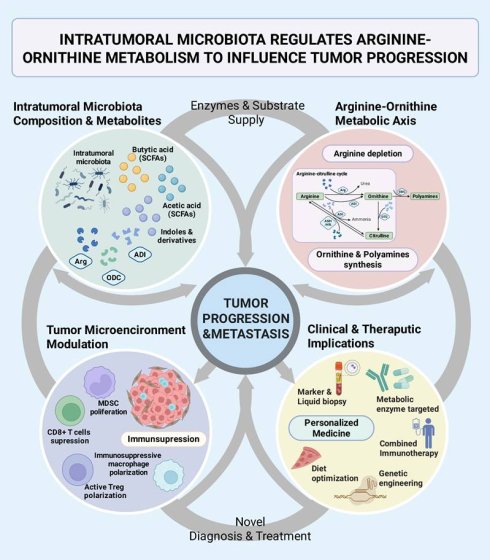
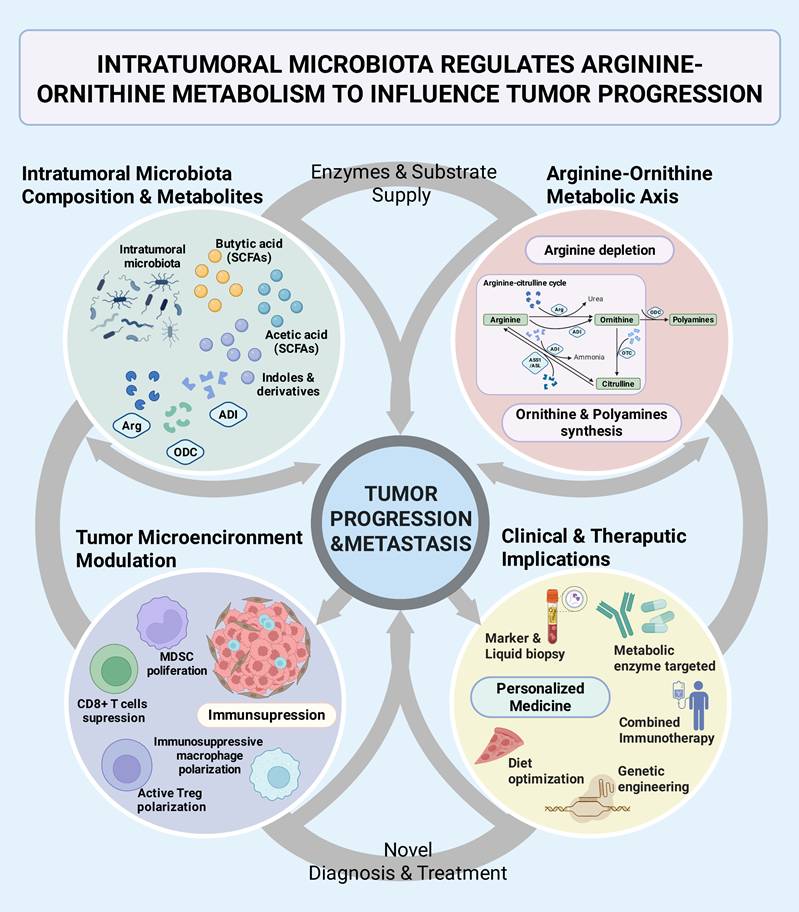
This review will focus on how does the intratumoral microbiota impact tumor progression by regulating the arginine-ornithine metabolism, to systematically summarize the progress of its mechanisms and explore its clinical translations (Figure 1). We will mainly discuss: 1) the effects of microbial arginine metabolic enzyme on immunity and metabolism in the TME; 2) the role of microbiota-derived ornithine metabolism in tumor proliferation and metastasis; 3) the balance of the ornithine-citrulline cycle in tumor metabolism and homeostasis; 4) the indirect regulation by microbial metabolites to related signaling pathways and overall metabolic reprogramming mechanisms; 5) some discoveries in typical cancers (Figure 1). Finally, we will look forward to the new biomarkers and intervention strategies based on the intratumoral microbiota-metabolism axis, including probiotic engineering, metabolic enzyme inhibitors, combined immunotherapy, and other creative progressions (Figure 1).
Intertumoral microbiota regulates arginine-ornithine metabolism to influence tumor progression. The interaction between intratumoral microbiota and the arginine-ornithine metabolic axis was systematically clarified, which jointly impact tumor progression and the tumor microenvironment, as well as provide inspiration for clinical applications. Intratumoral microbiota and their metabolites regulate tumor progression and the immune microenvironment within the tumor through processes such as arginine depletion and polyamine production. The microbiota-metabolism-immune axis in tumor also provides novel perspectives for clinical diagnosis and treatment. SCFAs, short chain fatty acids; Arg, arginase; ADI, arginine deiminase; ODC, ornithine decarboxylase; OTC, ornithine transcarbamylase; ASS1, argininosuccinate synthetase; ASL, argininosuccinatelyase; MDSC, myeloid-derived suppressor cell.
2. Metabolic regulation of arginine and ornithine through the microbiota
2.1. Arginine degradation and metabolic competition mediated by intratumoral microbiota
The metabolic state of arginine in the TME determines the function of immune cells and tumor cells. Some intratumoral microbiota can highly express arginine metabolic enzymes to consume arginine, causing local arginine deficiency and resulting in immunosuppressive effects [18, 20].
On one hand, bacterial arginine deiminase (ADI) can hydrolyze arginine into ornithine, directly depleting the tumor arginine supply [18] (Figure 2). For instance, highly abundant Staphylococcus anginosus (S. anginosus) colonized in gastric cancer was found to have an active arginine metabolism, which can rapidly consume arginine to generate ornithine, leading to a significant decreased arginine in co-cultured tumor models [18]. Further compared with sterile controls, S. anginosus colonization could reduce arginine levels with increased ornithine in tumor tissues, and accelerate tumor growth in vivo [18]. The microbiota-mediated arginine depletion also has a profound impact on immune cells. The proliferation and effector functions of T lymphocytes depend on a sufficient supply of arginine, and T cells will enter cell cycle arrest and functional failure under deficient arginine [18, 21]. The reduction of the competing substrates for inducible nitric oxide synthase (iNOS) in macrophages decreases the NO-mediated anti-tumor killing. At the same time, arginine depletion induces an increased tendency toward regulatory immune cells (Figure 2). Studies found that certain Proteobacteria can enhance the inhibitory function of regulatory T cells (Treg) in the TME by consuming host arginine, then weaken the anti-tumor immune response [22, 23]. Lacking these arginine-consuming bacteria increased the serum arginine levels in mice, which promote the excessive activation of mTOR signaling in Tregs with altered functional state [22]. Conversely, when there were abundant arginine-consuming bacteria, arginine levels decreased and the mTOR pathway was restricted in T cells, that is more conducive to form Tregs mediated immune tolerance [22, 23].
Intratumoral microbiota affects the immune microenvironment and tumor progression through arginine and ornithine metabolism. Arginine depletion: Intratumoral microbiota highly expressed ADI or Arg, consume arginine in the TME to induce the failure of T cell function and enhance immunosuppressive cells (Treg). Ornithine & polyamine supply: Intratumoral microbiota metabolizes arginine into ornithine, then synthesize polyamines from ornithine to stimulate tumor cell proliferation and reshape the immunosuppressive TME. A positive feedback loop forms to promote tumor deterioration through the ornithine-polyamine axis. Ornithine-citrulline cycle: As part of the urea cycle, arginine can be hydrolyzed by Arg to generate ornithine, which is combined with OTC to form citrulline. Citrulline then forms arginine under the action of ASS1/ASL. However, in tumors with arginine synthesis defects (ASS1 deficiency), arginine depletion can starve tumor cells to limit tumor progression. Intratumoral microbiota regulates the direction and degree of arginine conversion to ornithine or citrulline, altering the dynamic of the metabolic cycle and thereby affect cell proliferation and TME homeostasis. Arg, arginase; ADI, arginine deiminase; ODC, ornithine decarboxylase; OTC, ornithine transcarbamylase; ASS1, argininosuccinate synthetase; ASL, argininosuccinatelyase; Tregs, regulatory T cells; IDO1, indoleamine 2,3-dioxygenase 1; TAM, tumor-associated macrophages.
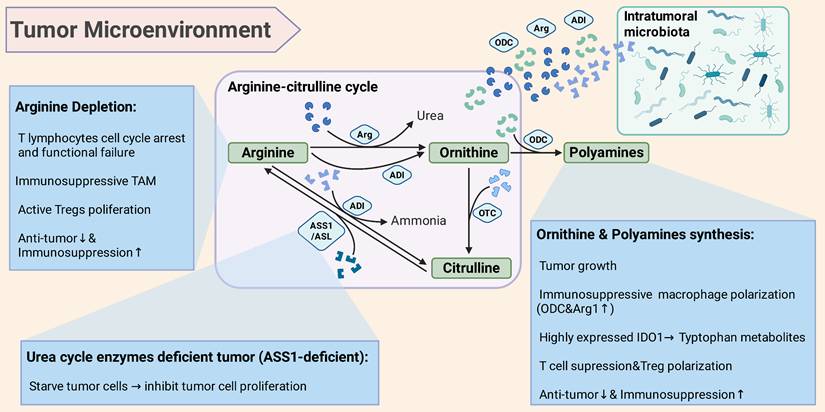
On the other hand, bacterial Arg can also participate in arginine degradation. Some pathogenic bacteria like Helicobacter pylori (H. pylori) expressed Arg can directly compete with host iNOS for substrates, decomposing arginine into ornithine and urea [24]. The arginine metabolism of H. pylori has been proven to significantly inhibit macrophages from synthesizing NO, thus promoting bacteria and tumors to evade immune clearance [24, 25]. Similarly, other Arg expressing microbiota enriched in tumor tissues may also play a similar role in that arginine depletion suppresses the effective function of immune cells, and the produced ornithine may be further utilized by tumor cells [18, 26] (Figure 2).
Moreover, arginine itself is an essential nutrient for tumors that cannot synthesize arginine due to the deficiency of urea cycle enzymes (Figure 2). Therefore, microbiota-mediated arginine depletion may directly starve some tumor cells [16, 27]. One of the therapies for tumors with arginine synthesis defects in clinical settings is to inject PEGylated arginine deiminase (ADI-PEG20) from microbiota to rapidly clear arginine and starve tumor cells, such as argininosuccinate synthetase (ASS1)-deficient liver cancer and melanoma [28, 29] (Figure 2). However, the regulation of arginine metabolism by the microbiota may vary depending on the various TME, leading to complex immune and metabolic regulation. Studies found that after colonizing a strain of bacteria with defective arginine metabolism to reduce the microbial breakdown of arginine, it actually accelerated tumor progression in a mouse model of CRC [26]. The excessive arginine accumulation was utilized by tumors and immunosuppressive cells, further activating the NO synthesis and polyamine synthesis. Thus, it promotes angiogenesis and immunosuppressive macrophage polarization, and activates the proliferative Wnt/β-catenin signaling, ultimately accelerating tumor progression [23, 26].
Overall, in most immunogenic tumors, the local arginine depletion by the intratumoral microbiota often favors tumor escape from immunity, by inhibiting T cells, NK cells and so on [29, 37] (Figure 2). While in some arginine-dependent tumors, the arginine depletion by microbiota may inhibit tumor cell proliferation (Figure 2). Conversely, if the microbiota increases the local arginine supply, it may both supply the tumor and stimulate immunosuppressive pathways, such as the Arg pathway of immunosuppressive macrophages, finally manifesting a tumor-promoting effect [17].
2.2. The synthesis of ornithine and polyamines mediated by intratumoral microbiota
Ornithine is the direct product of arginine degradation, and its accumulation in the TME has various effects on tumor progression. Firstly, ornithine can be converted into polyamines including putrescine, spermidine, and spermine under the action of ODC, which are important promoters of cell proliferation and differentiation [30] (Figure 2). Polyamines can facilitate nucleic acid and protein synthesis, stabilize nucleosome, and affect the cell cycle by regulating the modification of translation initiation factor EIF5A [31]. Furthermore, polyamines not only promote tumor growth and invasion, but can also form an immunosuppressive environment through the reprogramming and feedback of immune cells (Figure 2). Spermine can induce high expressed indoleamine 2,3-dioxygenase (IDO1) in tumor cells and DCs, which lead to the accumulation of tryptophan metabolites, further inhibiting T cell and driving the differentiation of Tregs [32, 33]. For instance, high levels of spermine can induce dysfunction of dendritic cells (DCs) and inhibit T cell proliferation, thereby help tumors evade immune surveillance [31, 34]. The level of polyamines within tumor is related to the malignancy and poor prognosis of tumors, and macrophages with high expressed ODC and Arg1 can support tumor growth and inhibit anti-tumor immunity by generating polyamines [20, 31] (Figure 2).
Recent studies proved that F. nucleatum is a common symbiont in tumors like CRC, which is also highly enriched in esophageal squamous cell carcinoma (ESCC) [19]. Clinical study showed that the F. nucleatum in ESCC tissues was significantly negatively correlated with the prognosis, patients with high abundance had a shorter survival period [19, 35]. F. nucleatum could invade ESCC epithelial cells and generated high levels of polyamine like putrescine, which disrupted the polyamine metabolic balance and promoted the malignant proliferation of tumor cells [35, 36]. This verified the crucial role of the ODC-polyamine metabolism from intratumoral microbiota in tumor progression. Furthermore, F. nucleatum is associated with enhanced tumor progression and metastasis in CRC, and its produced polyamines and toxic metabolites may promote tumor occurrence through activating Wnt/β-catenin signaling [35, 37, 38]. Besides F. nucleatum, others such as S. anginosus and Escherichia coli (E. coli) may also contribute to polyamine metabolic activity. S. anginosus increases ornithine production from arginine, accompanied by the massive accumulation of downstream polyamine derivatives such as N¹-acetylseramine and N⁸-acetylornithine in gastric tumor [18]. Additionally exogenous supplementation of arginine and its downstream acyl spermidine significantly accelerated tumor growth in gastric cancer xenograft model [21, 39]. Therefore, it can be inferred that S. anginosus produces ornithine and polyamines through arginine metabolism, and activates the polyamine pathway to promote tumor proliferation.
In addition to immune and proliferation regulation, the microbiota-arginine-polyamine axis also plays a crucial role in tumor blood and lymphatic vessels. As a downstream of the microbial arginine metabolism, polyamines are necessary regulators in the vascular characteristic. Recent studies showed that, the increased polyamine in the TME from intratumoral microbiota such as F. nucleatum significantly upregulated the expression of vascular endothelial growth factor (VEGF) and basic fibroblast growth factor (bFGF), through the hypusination of EIF5A, thereby accelerating pathological angiogenesis [19, 40]. Moreover, the microbial regulation of arginine directly affects the level of NO in the TME. In CRC models, it has been demonstrated that arginine catabolism mediated by certain microbiota can drive the angiogenesis by activating the Wnt/β-catenin signaling pathway in endothelial cells, promoting vessel branching and damaging vascular integrity, thereby facilitating hematogenous metastasis [26]. It is noteworthy that this metabolic axis also impacts the lymphatic system. Microbiota derived ornithine can be utilized by TAMs to drive polyamine-dependent metabolic reprogramming, which further induce a lymphangiogenic phenotype characterized by high secretion of VEGF-C and VEGF-D [41, 42]. This process not only expands the lymphatic network within the tumor, but also inhibits the recruitment of effector T cells to the lymphatic vessels, thereby protecting metastasis [43]. Therefore, the intratumoral microbiota regulates the TME structure by balancing the conversion of arginine to polyamine, suggesting that targeting this axis may have dual benefits of blocking nutrient supply and its lymph escape pathways.
2.3. Arginine-citrulline cycle: the influence of microbiota on tumor metabolism and homeostasis
Within the arginine metabolism, ornithine and citrulline can be interconverted through the urea cycle and related branches, forming an ornithine-citrulline cycle [44, 45] (Figure 2). This cycle is crucial for maintaining the metabolism and homeostasis of TME, while intratumoral microbiota further increases the dynamics of this cycle. As a part of the urea cycle, arginine can be hydrolyzed to generate ornithine by Arg. And ornithine is combined with carbamoyl phosphate by ornithine transcarbamylase (OTC) to form citrulline, which then integrates with aspartate to generate arginine under the action of ASS1/ argininosuccinatelyase (ASL) [16, 45] (Figure 2).
However, ASS1/ASL lacking tumor cannot effectively utilize citrulline to regenerate arginine, hence exogenous arginine becomes a limiting factor for tumor growth [46, 47]. The arginine metabolism of microbiota can disturb the balance of this cycle (Figure 2). When the intratumoral bacteria highly express ADI, arginine is largely converted into citrulline and ammonia that is not utilized under ASS1 deficiency [47]. Conversely, if the microbiota mainly consumes arginine through Arg pathway, the produced ornithine may be utilized by tumor cells to synthesize polyamines or other molecules, thereby partially supplying for tumor cells [13, 16, 30] (Figure 2). PEG-arginase drugs can convert arginine to ornithine, but the potential risk is that excessive ornithine may synthesize polyamines with tumor promoting effects [48, 49]. Thus, the microbiota mediated transformation of arginine to ornithine or citrulline has a dual impact on the tumor metabolism, depending on the ability of ornithine/citrulline utilization and the activity of downstream pathways of the tumor (Figure 2). If the tumor can effectively utilize ornithine to synthesize polyamines, the microbial Arg will support it. If the tumor cannot utilize citrulline and is arginine-dependent, the ADI pathway will cause arginine starvation to damage it.
Additionally, the by-products of arginine metabolism including ammonia and urea, also impact the metabolic and immune TME. High levels of ammonia generated by microbial ADI can increase the local pH and change the acidic state of the TME, which have been identified as a major driver of T cell dysfunction [32, 50, 51]. A recent study demonstrated that ammonia induced metabolic reprogramming in tumor-infiltrating T cells with lysosomal and mitochondrial stress, ultimately enhancing T cell exhaustion [52]. Tumor cells also recycle ammonia as a fundamental nitrogen source via glutamate dehydrogenase (GDH), to support amino acid synthesis and metabolism, thereby exhibiting a survival advantage under nutrient-deprived conditions [53]. In addition, the accumulation of urea from enhanced Arg further shapes the immunosuppressive TME. Urea cycle dysregulation is recognized not only as a metabolic hallmark of cancer also as a mechanism to evade immune surveillance. It promotes the recruitment and function of myeloid-derived suppressor cells (MDSCs), which further deplete local arginine and secrete inhibitory cytokines, resulting in positive feedback of immune exclusion [54]. Furthermore, these by-products may influence the efficacy of chemotherapy and immunotherapy, through modulate the local pH in TME [55].
Moreover, the correlative pathways also make significant contributions to tumor progression and immune regulation in the presence of arginine regulated microbiota. Ornithine produced through the Arg pathway can be further converted into proline by ornithine aminotransferase, which promotes collagen synthesis and tumor fibrosis [56-58]. In highly fibrotic tumors such as PDAC, studies found that the enriched Pseudomonas present massive Arg and proline metabolic genes. The microbial-driven arginine-proline axis enhance tumor matrix fibrosis and physical barrier for immune cell infiltration by promoting collagen synthesis. It also supports the stress survival of cancer cells as a key bypass for polyamine biosynthesis [16, 59]. Recent spatial multi-omics study further confirmed that this microbial-metabolism interaction is important for the malignant progression and immune evasion of PDAC [60, 61]. Similarly, aspartate is used to rescue arginine synthesis through the ASS1 pathway under the arginine deficiency, further result in the aspartate deficiency. Since aspartate is an important nitrogen source for nucleotide biosynthesis and DNA repair, excessive microbial-induced aspartate depletion limits tumor cell proliferation and increases their sensitivity to oxidative stress [62]. This microbial-driven metabolic redirection provides new logical support for starvation therapy targeting ASS1-deficient tumors. Although there are relatively few studies about it, intratumoral microbial-driven arginine reprogramming indirectly affects other amino acid biosynthesis including proline and aspartate [16, 59]. It also represents an innovative frontier: targeting microbial arginine metabolism within tumor can simultaneously disrupt multiple interdependent amino acid pathways, providing synergistic treatment for tumors such as PDAC.
2.4. The indirect regulation of the arginine pathway by microbial metabolites
In addition to directly participating in arginine and ornithine metabolism, other metabolites and signaling molecules produced by intratumoral microbiota can also affect the arginine metabolism and immune metabolic responses of the host [63]. The most typical include short-chain fatty acids (SCFAs) and derivatives of tryptophan metabolism [64-66].
2.4.1. Short-chain fatty acids (SCFAs)
SCFAs as the fermentation products of the microbiota, are widely present in the intestinal tract and TME. They can regulate the metabolism of immune cells through G protein-coupled receptors (GPCRs) and epigenetic pathways [67-69]. SCFAs produced by intratumoral microbiota, including acetic acid from Bifidobacterium and butyric acid from Lactobacillus, may also be circulated from the intestinal microbiota to TME [70-72].
SCFAs impact the arginine metabolism by altering the immune cell functions, thereby regulating the enzymes activity including iNOS and Arg1[64, 73, 74] (Figure 3). Butyric acid with histone deacetylase inhibitor (HDACi) activity, can promote the fatty acid oxidation in Tregs and enhance their differentiation [75, 76] (Figure 3). The activated Tregs is accompanied by the activated immune-inhibiting metabolic pathway Arg1 and IDO in macrophages and DCs, which force the competitive metabolism of arginine and tryptophan with effector T cells inhibition [22, 77]. And acetic acid acts on its receptor GPR43/41 of macrophages, and its high concentrations can inhibit glycolysis and pro-inflammation in macrophages, driving its immunosuppressive transformation [78, 79] (Figure 3). Immunosuppressive macrophages typically express higher Arg1 and further secrete arginine-derived polyamines with the reinforced immunosuppressive metabolic effect [17, 20] (Figure 3). Bifidobacterium colonized in CRC can secrete metabolites including lactic acid and acetic acid, which facilitate tumor growth and immune escape, leading to poor patient prognosis [80]. Therefore, SCFAs promote pro-inflammatory and immunosuppressive phenotypes, further upregulate the Arg1 activity and arginine consumption with polyamine production in the TME, finally contribute to the tumor immune escape.
The metabolites of intratumoral microbiota impact TME through signaling pathways. Arginine can activate Rag GTPase through CASTOR1, thereby promoting the activation of mTORC1. Under arginine depletion induced by intratumoral microbiota, the mTOR signal in T cells weakens, inducing Treg differentiation rather than effector T cells. Butyric acid can regulate the mTORC1 signal through its HDACi activity, and promote the metabolic bias towards fatty acid oxidation in regulatory T cells with enhanced the differentiation. Acetic acid in SCFAs acts on macrophages through GPR43/41, promoting their immunosuppressive transformation. Tryptophan produces indole and its derivatives (I3A, IPA, IAA) to activate the AHR signaling pathways, which promotes immunosuppressive macrophages and the differentiation of Treg cells, and inhibits the proliferation of effector T cells with an immunotolerant TME. The metabolite-created immunotolerant TME is accompanied by the activation of Arg1 in macrophages and IDO in DC, further accelerate the arginine depletion and tryptophan, finally strengthening immunosuppression and promoting tumor progression. TME, tumor microenvironment; SCFAs, short chain fatty acids; Arg, arginase; HDACi, histone deacetylase inhibitor; FAO, fatty acid oxidation; I3A, indole-3-carboxaldehyde; IPA, indole propionic acid; IAA, indole-3-acetic acid; AHR, aryl hydrocarbon receptor; GPR43/41, G protein-coupled receptor 43/41; Treg, regulatory T cell; IDO, indoleamine 2,3-dioxygenase.
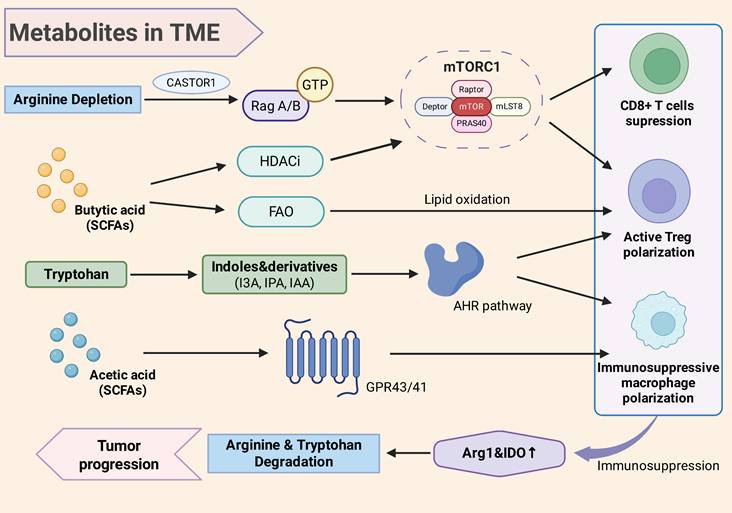
Moreover, SCFAs as metabolic substrates can participate in the tricarboxylic acid cycle of T cells and NK cells, in order to provide energy and regulate their differentiation [81] (Figure 3). And moderate exposure of SCFAs can also perform anti-tumor immunity. Butyric acid can inhibit HDAC and enhance mTOR signaling, to improve the glucose metabolism and cytotoxic activity of CD8+ T cells, which may counteract the adverse effects of arginine deficiency [81]. In summary, SCFAs produced by intratumoral microbiota are important regulators of the immune metabolic reprogramming in the TME, which can both enhance anti-tumor immune and induce immune tolerance, depending on the dose, cell type, and time window [81, 82].
2.4.2. Metabolic derivatives of tryptophan
The tryptophan metabolites and aryl hydrocarbon receptors (AHR) pathways, are also key components of the microbiota-host immune metabolic axis [61, 83, 84]. Many intestinal and intratumoral microbiota can metabolize tryptophan to produce indoles and their derivatives, like indole-3-carboxaldehyde (I3A), indole-3-propionic acid (IPA) and indole-3-acetic acid (IAA). [61, 85]. These microbiota-derived indole compounds as ligands of the AHR, can activate the AHR signaling pathway in immune cells [86] (Figure 3). AHR is a ligand-dependent transcription factor expressed in various immune cells. Its activation drives the transcription of downstream genes, thereby influencing the differentiation and metabolic state of immune cells [86]. In the TME, the continuous activation of AHR signals is often associated with immunosuppression [61, 86, 87]. Studies found that TAMs in PDAC exhibit high AHR activity, which might be concerned with the continuous exposure to indole metabolites from microbiota [61]. Macrophages with high AHR tend to secrete immunosuppressive factors with expressing Arg1, thereby creating an immunotolerant TME [61] (Figure 3). Additionally, the activated AHR can promote the differentiation and function of Treg cells and inhibit the effective T cells [88, 89] (Figure 3). Blocking AHR signals can suppress tumor growth and enhance anti-tumor immune responses in tumor models, supporting the crucial role of AHR in immunosuppressive metabolism [90]. Since AHR is closely related to the IDO-tryptophan metabolism, the AHR activated by intratumoral microbiota often occurs under co-dysregulation of arginine and tryptophan metabolism, together with highly expressed Arg1 and IDO in an immunosuppressive TME [33, 61, 83] (Figure 3). This means that the activated AHR may synergize with bacterial Arg1 pathway to accelerate the depletion of arginine and tryptophan, causing immune tolerance through downstream signaling.
2.4.3. mTOR signaling pathway
Furthermore, the mTOR signaling pathway acts as the cellular central processor to sense amino acid abundance and regulate metabolism [91-93]. Arginine is an important upstream stimulus for mTORC1 complex, and the sense of the arginine concentration mainly relies on CASTOR family proteins, which contains CASTOR1 and CASTOR2. CASTOR1 is the core sensor that directly binds to arginine. When arginine is deficient, CASTOR1 binds to the GATOR2 complex and inhibits its activity, thereby blocking the activation of mTORC1. Therefore, CASTOR1 has a specific arginine binding site, serving as a trigger for concentration sensing. While CASTOR2 as a structural stabilizer form a heterodimer with CASTOR1 to adjusting the sensing sensitivity by influencing the conformation of CASTOR1. Sufficient arginine can activate Rag GTPase through amino acid sensors CASTOR1, then promote mTORC1 localized on the lysosomal membrane and activated by Rheb [94, 95]. The activated mTORC1 further forces anabolic metabolism, inhibits autophagy, and supports cell growth and proliferation [96, 97].
Intratumoral microbiota also affects the activity of the mTOR pathway in immune cells and tumor cells by controlling arginine levels (Figure 3). It has been verified that when the microbiota extensively decomposes arginine, mTOR signal in T cells is decreased, which prefer to inducing Treg differentiation rather than effective T cells [22]. Conversely, if probiotics metabolism increases the arginine level, the enhanced mTOR activity may drive its differentiation towards the effective helper T cell/cytotoxic T lymphocyte (Th1/CTL), improving anti-tumor ability [1, 22]. Therefore, engineered strains were used to provide arginine locally, its results confirmed a significant increase in tumor-infiltrating T cells with enhanced expression of downstream effector molecules like IFN-γ, ultimately enhancing the immunotherapy [98, 99]. Moreover, SCFAs can reprogram T cell metabolism by affecting the AMPK and Akt-mTOR axis [82, 100, 101] (Figure 3).
Moreover, tumor cells also downregulate anabolic metabolism through mTORC1 pathway when faced with arginine starvation from intratumoral microbiota. However, unlike immune cells, tumor cells exhibit a strong metabolic flexibility [94, 102]. They perform a stronger autophagy adaptation ability to supply arginine by degrading intracellular proteins. Tumor cells also compensatorily activate the mTORC2 pathway through lysosomal localization. mTORC2 as a key upstream of the Akt pathway, its enhanced activity can maintain the phosphorylation of Akt in arginine-deficient environments, thereby activating autophagy to recycle endogenous arginine and regulating the anti-apoptotic ability of tumor cells [102]. Therefore, this explains why some tumors can still survive under severe arginine deprivation, by enhancing invasiveness to achieve metabolic escape.
Summary of typical microbiota and their roles in arginine metabolism in various cancers.
| Cancer Type | Typical Microbiota | Roles in Arginine Metabolism | References |
|---|---|---|---|
| Colorectal Cancer (CRC) | Fusobacterium nucleatum | Produces polyamines from arginine, activating Wnt/β-catenin pathway, promoting proliferation and metastasis. | [19, 26, 136] |
| Escherichia coli | Arginine degradation via AST pathway, influencing tumor progression by enhancing angiogenesis and immunosuppressive macrophages. | [26, 104, 136] | |
| Gastric Cancer (GC) | Helicobacter pylori | Arginine consumption via Arg pathway, depletes NO production, and induces immunosuppression. | [25] |
| Streptococcus anginosus | Produces ornithine from arginine, promoting tumor cell proliferation and immune suppression. | [137] | |
| Esophageal Cancer (ESCC) | Fusobacterium nucleatum | Converts arginine to polyamines, disrupting the polyamine metabolic balance and enhancing malignancy. | [19] |
| Pancreatic Cancer (PDAC) | Pseudomonas | Enriches with arginine and proline metabolic genes, promoting immune suppression via macrophages. | [113] |
| Bladder Cancer (BCa) | Fusobacterium nucleatum | Converts arginine to ornithine and polyamines, stimulating tumor proliferation and promoting immunosuppressive TME. | [26] |
| Melanoma | Fusobacterium nucleatum | Increases tumor progression through polyamine metabolism, modulating immune responses. | [138] |
| Porphyromonas gingivalis | LPS production and immune modulation, contributing to tumor progression via NF-κB signaling. | [99] | |
| Head and Neck Cancer (HNSCC) | Streptococcus anginosus | Increases Arg1 expression in macrophages, promoting immunosuppressive phenotypes. | [139] |
3. Microbiota-arginine/ornithine metabolic axis in typical cancers
Due to the various locations, microenvironments and microbiota compositions among different cancers, the mechanisms by which the intratumoral microbiota affect metabolism in TME also differ. Here, we will analyze and summarize the research progress of the microbiota-arginine/ornithine axis combining with representative cancers:
3.1. Colorectal cancer (CRC)
CRC is the tumor with the most extensive research on microbiota that both inside and outside the tumor may be involved in tumor progression. F. nucleatum is a recognized intratumoral bacterium in CRC, and its abundance is significantly correlated with poor prognosis and increased liver metastasis [103-105]. Besides adhering to epithelial cells and activating oncogenic signals, F. nucleatum also impact the TME through metabolism [35, 106]. Its produced polyamines may promote tumor cell proliferation and inhibit immune surveillance in TME, through Wnt/β-catenin pathway activation and IL-10 production from myeloid cells [107-109]. E. coli is also a frequently detected intratumoral strain in CRC [110, 111]. Some of strains carry the pks virulence inducing DNA damage, while others are involved in metabolic regulation. Studies showed that the arginine degradation by intratumoral microbiota is significantly enhanced in CRC, especially some E. coli can consume arginine largely through the arginine succinyltransferase (AST) pathway [26, 111]. In mouse models, inhibiting microbial arginine metabolism could increase arginine levels in tumors, but induced a series of tumor-promoting changes, including increased NO and polyamine synthesis, enhanced angiogenesis, increased infiltration of immunosuppressive macrophages, and upregulation of Wnt signaling, ultimately promoting tumor progression [26]. It suggested that the arginine metabolism of intratumoral microbiota may limit tumor growth, also weaken immunity. Clinically, CRC patients attempted to take high-dose arginine orally to improve immune efficacy, but the effect was not obvious, indicating that the metabolic regulation of host microbiota-arginine is extremely complex [98, 112]. Given the rich and easily modifiable microbiota, CRC is also an important field to exploring microbiota metabolic therapy, and its specific mechanism and intervention still need to be further explored.
3.2. Pancreatic cancer (PDAC)
The TME of PDAC presents highly fibrotic and immunosuppressive, and has been confirmed to contain rich intratumoral microbiota [60, 113]. They mostly originate from the intestine or oral cavity, and enter the pancreas through the bloodstream or bile ducts, such as Clostridiales, Actinomycetales, and Gram-negative bacteria [114-116]. Studies found that intratumoral microbiota imbalance can drive innate and adaptive immune suppression, including increased TAMs and MDSCs, and reduced the infiltration of effective T cells [113, 117]. Pushalkar et al. verified that the tumor growth was slower in germ-free mice, and the microbiota removed by antibiotics could reactivate anti-tumor immunity and improve the efficacy of checkpoint inhibitors (ICB) [113, 118]. In terms of metabolism, the intratumoral microbiota may utilize the abundant free amino acids to compete or cooperate with tumor cells [8, 61, 119]. Pseudomonas in PDAC have enriched Arg enzyme and proline metabolic genes, which are speculated to consume arginine and produce metabolites to affect anti-tumor immunity in the TME [8]. Additionally, there is evidence that the macrophages in PDAC performed high AHR activity and immunosuppressive phenotype, due to the long-term exposure to indole derivatives from microbiota metabolic products [61, 119]. It demonstrated that the intratumoral microbiota strengthens the suppressive immune metabolism of Arg1 and IDO via the indole-AHR axis, synergistically enhancing tumor immune escape. Clinically, H. pylori DNA has also been detected within tumors of PDAC patients. It is supposed that the Arg activity of H. pylori may play a role to inhibit macrophages produced NO in the pancreatic TME [120, 121]. In summary, the intratumoral microbiota exacerbates the "cold tumor" of PDAC through arginine metabolism, the interventions targeting intratumoral microbiota are expected to be explored and improve the immune microenvironment [122, 123].
3.3. Bladder cancer (BCa)
BCa has unique microbial characteristics in the urinary tract, which can regulate arginine metabolism in the TME. Sequencing studies revealed specific intratumoral microbiota different from urinary microbiota, like F. nucleatum [124]. F. nucleatum is known to convert arginine into ornithine and polyamines, which directly promote tumor cell proliferation and shape an immunosuppressive TME. Besides, the microbiota expressing ADI or Arg locally consume arginine, thereby impairing the T cells proliferation and the NO synthesis by macrophages [22]. The metabolic by-products of microbial arginine degradation further affect the immune TME. Excessive polyamines in BCa can suppress the entry of arginine into immune cells and NO-mediated tumor killing activity, then inhibiting anti-tumor immunity. In clinical study, the attenuated Mycobacterium tuberculosis stimulated a strong iNOS activity in the bladder, elevating local NO levels, thereby promoting the killing of tumor cells [125]. Moreover, many invasive BCa also perform loss of ASS1 with arginine dependence. Therefore, ADI-PEG20 significantly inhibits the growth of ASS1-deficient BCa in vitro and in mouse models by depleting extracellular arginine [126]. This highlights that the microbiota-arginine/ornithine metabolic axis is crucial in BCa progression and anti-tumor immune response. Targeting the axis by modulating the intratumoral microbiota and arginine metabolism is an innovative strategy to enhance anti-tumor immunity and drives new clinical research in BCa.
3.4. Melanoma
Melanoma is an immunogenic cancer, the arginine metabolism and microbiota in the TME significantly affect tumor immunity. Most melanomas lack ASS1, which cannot synthesize arginine, hence it can be a therapeutic point targeted by arginine-degrading enzymes [127]. Clinical studies explored that ADI-PEG20 combined with chemotherapy showed persistent control activity in advanced ASS1-negative melanomas [28]. Conversely, arginine supplement can enhance CD8⁺ T cell tumor infiltration and improve the efficacy of anti-PD-1 checkpoint therapy in mouse models [128]. Furthermore, microbiota can systematically regulate arginine levels and immune cell function. In melanoma tissues, translocated bacteria from the oral cavity and skin are frequently detected, such as Porphyromonas gingivalis (P. gingivalis) and F. nucleatum [129]. They carried lipopolysaccharides (LPS) and metabolites stimulate NF-κB through TLR2/4 signaling pathway, promoting the polarization of immunosuppressive macrophages with upregulated Arg1 expression [1,129]. Research revealed that the intestinal probiotic Limosilactobacillus reuteri can migrate to the melanoma and utilize tryptophan to synthesize I3A. I3A then activates the AhR signaling pathway to induce an efficient state of CD8+ T cells, significantly enhancing the ICB efficacy [130]. However, the high abundance bacteria with degrading arginine in TME may weaken this enhancing efficacy. Studies have found that the level of L-arginine in the circulation is an independent predictor of ICB efficacy in melanoma patients, and the abundance of arginine-degrading enzymes in intratumoral microbiota is negatively correlated with this [122, 131].
3.5. Mesothelioma
Malignant mesothelioma as an aggressive pleural tumor, is significantly influenced by arginine metabolism. One of the notable metabolic features is that about 50% tumors present absence of ASS1, resulting in absolutely dependence on exogenous arginine [132]. In the Phase III clinical trial ATOMIC-Meso, researchers conducted a randomized, double-blind controlled study on 249 patients with non-epithelioid mesothelioma [133]. Its results showed that adding ADI-PEG20 (peglotamycin) to the standard chemotherapy regimen (pemetrexed + cisplatin) significantly prolonged the overall survival (OS) and progression-free survival (PFS) of the patients, highlighting a new metabolic efficient therapy [133]. The arginine/ornithine axis also reshapes the immune TME of mesothelioma. Mesothelioma triggers a chronic inflammatory TME dominated by TAM and neutrophils with high ARG1 and iNOS dysregulation, which converts arginine into ornithine and polyamines rather than anti-tumor NO [128]. Excessive polyamines and ornithine enhance fibrosis and immunosuppression, promoting tumor growth. Although mesothelioma typically occurs in sterile sites, the systemic microbiota may affect its course through regulating host arginine and immune microbiota [134]. Eliminating bacteria that consume arginine has been proven to increase arginine levels and enhance CD8⁺ T cell activity in distant tumors, which might be used to enhance anti-tumor immunity in mesothelioma patients [23, 135].
4. The potential for diagnosis and treatment
4.1. Biomarkers and prognosis
The characteristics of intratumoral microbiota and their metabolites have been regarded as potential diagnostic and prognostic markers recently [6, 18, 140-142]. Studies on gastric cancer discovered that patients with high alpha diversity of intratumoral microbiota have better prognosis, while certain bacteria with excessive abundance are present in poor prognosis subtypes such as S. anginosus and F. nucleatum [18, 143-145] (Table 2). The classification according to composition of intratumoral microbiota can independently predict the survival of patients from the clinical stage [18]. Additionally, the microbial signals of metabolic pathways can also serve as markers [11, 146]. Patients with enriched S. anginosus in gastric cancer present an upregulated arginine metabolic pathway, which has been proposed as a risk factor [18]. And the abundance of F. nucleatum in tumors and feces has been explored as an indicator for early screening and prognosis assessment of CRC [104, 147]. High abundance indicates a more aggressive tumor and poorer prognosis, which may be partly attributed to its pro-tumor effects of polyamine metabolism [104] (Table 2).
Summary of transformation targeting intratumoral microbiota-arginine/ornithine axis in TME.
| Transformation | Therapy Directions | Targets | Mechanisms | Reference |
|---|---|---|---|---|
| Diagnosis | Microbiota diversity markers | α-diversity of intratumoral microbiota | Positively correlated with the prognosis of patients | [143-145] |
| Abundance of specific microbiota (Streptococcus, Fusobacterium) | Enrichment indicates an upregulation of the arginine metabolic pathway, leading to immunosuppression and tumor progression | [18] | ||
| Metabolic pathway markers | Genetic abundance of the bacterial Arg/ODC pathway & host expression of Arg1/ODC | By consuming arginine through ADI or Arg, ornithine or polyamines are produced, thereby altering the nutritional status of the TME | [11, 146] | |
| Serum arginine/ornithine/citrulline levels | Decreased arginine and increased citrulline indicate an imbalance in the metabolic pathway | [150] | ||
| Non-invasive liquid biopsy | Detection of F. nucleatum in faeces | Associated with the risk of liver metastasis in CRC | [104, 147] | |
| Blood SCFAs/polyamine metabolite profile | Indirectly reflect the metabolic activity and immune status of the microbiota | [139, 148, 149] | ||
| Treatment | Metabolic enzyme targeted | Arg1/Arg2 inhibitor (CB-1158) | Relieve arginine depletion and restore T cell function | [158, 159] |
| ADI-PEG20 (Arginine Exhaustion) | Selective starvation of ASS1-deficient tumors | [152] | ||
| ODC inhibitor (DFMO) | Block polyamines synthesis and reverse immunosuppression | [153, 154] | ||
| Microbial editing and engineered bacteria | Engineering E. coli Nissle 1917 (arginine synthesis) | Supplementing arginine in the tumor tissue and enhancing T cell infiltration | [98, 160] | |
| Bacteriophage/antimicrobial peptide targeting F. nucleatum | Specific elimination of tumor-promoting bacteria reduces the production of polyamines | [161, 162] | ||
| CRISPR-Cas editing of the microbiome genes | Weaken the specific metabolic functions of the microbiota without disrupting the ecosystem | [165, 166] | ||
| Traditional therapy optimization | Narrow-spectrum antibiotics (ciprofloxacin) | Eliminate specific tumor-promoting microbiota and reinvigorate the response to immunotherapy | [167] | |
| High-fiber/low-red-meat dietary intervention | Adjust the metabolic profile of the microbiota and inhibit the polyamine pathway | [170, 171] | ||
| Combined immunotherapy | Metabolic enzyme inhibitors + ICB (anti-PD-1/CTLA-4) | Metabolic intervention removes the immunosuppression of TME, and ICB activates T cells to kill the target cells | [155, 174, 175] | |
| Delivery of immune factors by engineered bacteria (IL-2, IFN-γ) | Continuously release cytokines, recruit and activate tumor-infiltrating lymphocytes | [178] |
Moreover, the expression of host immune metabolic enzymes also reflects the role of the microbiota. High level of serum Arg1 in head and neck squamous cell carcinoma (HNSCC) patients is associated with poor prognosis, which is regarded to be induced by intratumoral microbiota and inflammatory TME [139, 148, 149] (Table 2). It has also been found that abnormal serum levels of various amino acids in HNSCC patients, including increased citrulline and decreased arginine, are associated with tumor metabolism and can served as auxiliary diagnostic factors [150] (Table 2). These metabolic changes may be partially contributed by arginine metabolism of intratumoral microbiota.
Therefore, it is expected that the composition of intratumoral microbiota and its metabolic indicators will be incorporated into the biomarker spectrum, in order to improve the diagnostic accuracy and prognostic evaluation of tumors. It is possible to consider joint detection involving the gene abundance of Arg and ODC pathway in tumor and fecal microbiota, the systemic levels of arginine and ornithine, and the expression of Arg and ODC in tumor tissues, aiming to assess the activity of the microbiota-metabolism axis from multiple perspectives and infer the biological and immune status of the tumor (Table 2). Large-scale cohort studies and multi-omics analyses in the future will further clarify the clinical value of these indicators.
4.2. Therapeutic targets and intervention strategies
A thorough understanding of the intratumoral microbiota-arginine/ornithine metabolic interaction has provided us with numerous therapeutic targets and innovative ideas. The following are several potential intervention strategies:
4.2.1. Metabolic enzyme targeted therapy
The inhibition and utilization of key enzymes such as Arg1/Arg2, ADI, and ODC is a direct intervention (Table 2). Clinically, PEGylated arginase (Peg-rhArg1, BCT-100) and ADI-PEG20 have been developed for arginine depletion in the body to treat tumors [28, 151, 152]. BCT-100 converts arginine into ornithine, which may potentially promote polyamine synthesis with promoting tumor growth, while ADI-PEG20 converts arginine into citrulline, which cannot be utilized by ASS1-deficient tumors, thereby more effectively starving the tumors [28, 152]. The irreversible inhibitor of ODC, difluoromethyl ornithine (DFMO), has been used clinically for the prevention of colorectal polyps and has shown tumor-suppressing and immune-active effects in neuroblastoma [153, 154] (Table 2). For the microbiota-related arginine metabolism, ADI inhibitors (L-Canavanine) are used to block the arginine metabolism of Streptococcus, and dose-dependently inhibit S. anginosus to convert arginine into ornithine [18]. Therefore, drugs selective targeting microbial ADI/Arg may be developed to precisely regulate the arginine metabolism of intratumoral microbiota, avoiding the impact on host cells.
It is worth noting that combining metabolic enzyme inhibition with immunotherapy has great potential. Arg1 inhibitors (CB-1158) combined with anti-PD-1 antibodies were used in phase I clinical trials for solid tumor patients, in order to alleviate the arginine depletion mediated by intratumoral microbiota and MDSCs, with enhanced efficacy of immunotherapy. Its results primarily indicated good tolerances [155]. In preclinical models, the combination of ODC inhibitors and ICB also showed a synergistic effect on reversing polyamine-mediated immunosuppression and enhancing T cell activity [25, 156, 157]. Therefore, it is possible to develop novel strategies including the simultaneous inhibition of Arg1 and ADI to control the two pathways of arginine metabolism [158, 159]. To sum up, metabolic enzyme targeted therapy provides a direct intervention for the intratumoral microbiota-metabolism axis, whose key lies in selecting appropriate patients, such as screening ASS1-deficient tumors for ADI therapy and controlling potential side effects.
4.2.2. Microbiota editing and probiotic therapy
A strategy that directly regulates the composition and function of intratumoral microbiota can intervene the metabolic interaction from the source. Importing beneficial bacterial strains can reverse the unfavorable metabolic state of the TME. For instance, an engineered E. coli Nissle 1917 has been constructed to reconvert tumor metabolic products into arginine in TME [98, 160] (Table 2). It significantly increases the local arginine concentration, enhances the tumor-infiltrating T cells and synergistically performs a stronger anti-tumor effect with PD-L1 antibody [98, 99]. It suggests that altering microbial function via probiotics or synthetic biology can improve the metabolism and signal environment of TME. On the other hand, eliminating or suppressing tumor-promoting microbiota is also an important direction. The antibacterial peptides or phage preparations specifically targeting F. nucleatum could be used to reduce its products like polyamines and LPS, and improve the prognosis of CRC patients [161, 162] (Table 2). Currently, the phages for F. nucleatum revealed activity in vitro [136, 163, 164].
In addition, CRISPR-Cas delivery vectors are used to deliver gene editing systems that cause the loss of arginine/ADI functions into intratumoral microbiota. It can suppress specific metabolic functions without disrupting the microbiota ecology [165, 166] (Table 2). Although the microbiota editing technologies are still in the early stage, they represent the future perspectives of precise therapy in the tumor microecology. It should be noted that there are challenges including delivery and the survival of the strains, as intratumoral microbiota are usually deeply hid within solid tumors. Hence, it is necessary to develop targeted delivery systems for tumor tissues, while strictly controlling safety to avoid infection or systemic inflammatory responses.
4.2.3. Optimization of traditional therapy for microbiota
The traditional therapy for microbiota involves antibiotics and dietary interventions. Systemic use of antibiotics in tumor treatment should be cautiously performed, as it may disrupt the beneficial microbiota and decrease the immune efficacy [167-169]. However, local or short-term use of antibiotics may be beneficial in cases where certain bacteria promote tumor progress. The systemic antibiotic clearance of the microbiota presents more sensitivity to immunotherapy in pancreatic cancer mice [113, 167]. Clinically, ciprofloxacin was attempted to add into chemotherapy for PDAC patients, and the results showed partly safety and improvement [167] (Table 2). Further developments may involve narrow-spectrum or targeted antibacterial strategies, using specific drugs to inhibit the microbiota with arginine metabolism while preserving the beneficial microbiota. This requires prior detection of intratumoral microbiota in patients and medication based on metabolic characteristics in order to achieve precise microbial treatment. Furthermore, a focus on diet, in particular high-fiber and low-red-meat diets will adjust the microbiota to produce more beneficial metabolites, such as increasing butyrate and reducing harmful amines, which may suppress tumor progression and improve immune function [72, 170, 171] (Table 2). Overall, the optimization of traditional therapy still requires more clinical validation, and it provides simple and feasible adjunctive therapy for patients.
4.2.4. Combination with immunotherapy
Since the intratumoral microbiota-metabolism axis plays a key role in immune regulation, it is expected that it could be combined with immunotherapy to achieve synergistic effects [86, 172, 173]. The metabolic enzyme inhibition and probiotics can alleviate the immune-suppressive state in TME, thereby enhancing the immunotherapy efficacy. For example, Arg1 inhibitors or polyamine blockers can inhibit the function of MDSC and Tregs, enhance T cell proliferation, and then administer PD-1/CTLA-4 antibodies to transform a "cold tumor" into a "hot tumor" [174] (Table 2). The combination of CB-1158 with PD-1 antibodies has been evaluated for safety and efficacy in clinical trials [155, 174, 175]. Additionally, engineered bacteria can directly deliver immune-stimulating molecules. The genes encoding immune-active factors have been introduced into intratumoral microbiota through engineered bacteria, which can maintain continuous expression in the tumor site and enhance the immunotherapy efficacy [176, 177]. Yoon et al. used attenuated Salmonella carrying the IFN-γ gene in mice, which significantly inhibited tumor growth [178] (Table 2). These strategies combined with metabolic regulation may simultaneously regulate metabolism and immunity to achieve more ideal anti-tumor effects. However, broad-spectrum antibiotics combined with ICB were found to weaken efficacy in some researches, suggesting the precise control of the intensity and timing of microbiota intervention should be explored [179]. The metabolic and microbiota regulation should be selected based on different patients, and combined with immunotherapy or chemotherapy, to achieve individualized comprehensive treatment.
5. Discussion and Perspective
More and more pieces of evidence indicate that intratumoral microbiota profoundly affect the TME and disease progression by reprogramming the arginine-ornithine metabolic pathway. The intratumoral bacteria can consume and metabolize arginine, thereby altering arginine supplement and immune mediators in the TME (Figure 2). S. anginosus found in gastric cancer can increase ornithine accumulation through arginine metabolism and significantly inhibit the differentiation and infiltration of CD8+ T cells, then promote tumor proliferation and metastasis [18] (Figure 2). Similarly, the ability of intratumoral microbiota to decompose arginine is closely related to CRC progression. The microbial arginine depletion can increase ornithine content within tumor, thereby activating the polyamine synthesis mediated by NO synthase and ODC, which induce angiogenesis, polarization of immunosuppressive macrophage, and upregulation of Wnt/β-catenin signaling, ultimately accelerating CRC growth [26]. Moreover, polyamines as crucial metabolites of the arginine-ornithine pathway, are termed as important mediators to promote tumor growth and immunosuppression (Figure 2). Tumor cells and myeloid suppressor cells perform excessive polyamine metabolism, and high levels of polyamines can cause immunosuppressive macrophage polarization, MDSC proliferation, and effective T cells suppression [51]. This is typical in "cold tumors" like PDAC, where cancer cells to produce ornithine and synthesize polyamines, shaping a highly immunosuppressive TME [51, 180]. Meanwhile, the by-products of arginine metabolism also synergistically promote tumor progression by reshaping the TME.
At the same time, the metabolites produced by intratumoral microbiota also regulate the host's amino acid metabolism and immune network through signaling pathways. SCFAs including butyric acid can act on the metabolic sensing pathway of T cells and inhibit histone deacetylase, thereby upregulating effector molecules such as IL-2 receptor (CD25) IFN-γ and TNF-α, enhancing the proliferation and anti-tumor activity of CTL and CAR-T cells [100] (Figure 3). Metabolites of tryptophan like indole derivative can activate the AHR signaling pathway and induce the expression of chemokines and cytokines like CXCL9 and IFN-γ, further promoting the proliferation of tumor-infiltrating T cells and inhibiting tumor growth [181, 182] (Figure 3). In digestive tract tumors, changes in microbial arginine metabolic pattern in CRC patients are related to the activation of the arginine and polyamine pathway in TME [26]. Pseudomonas enriched in PDAC were found to be significantly associated with the abnormal enrichment of arginine and proline metabolic pathways [99]. Therefore, the intratumoral microbiota-metabolism-immune interaction constitutes a crucial network that affects the occurrence and development of cancer for clarifying tumor mechanisms.
As the field advancing rapidly, it still faces many challenges and unsolved problems [183]. Firstly, intratumoral microbiota presents a low biomass with the weak microbial signals in the samples that are susceptible to environmental and sequencing contamination, thus making it very difficult for it to be accurately identified and located [184]. Secondly, the heterogeneity and dynamic changes of microbiota among different patients make it complicated to explore the mechanisms and clinical translation [2]. Thirdly, many conclusions about the influence of the microbiota on host metabolism and immunity are mainly based on correlation analyses that need to be strictly proven. For instance, only certain microbiota were found to be associated with amino acid metabolism disorders in PDAC study, but the causal mechanism and specific mediators remain unclear, and require further experimental evidence to support [8]. Additionally, our knowledge of how the intratumoral microbiota can reshape the immune function through metabolism is still limited, and more accurate analytical methods are needed. Nevertheless, deeper exploration for intratumoral microbiota is considered crucial for revealing processes of tumor immune evasion and metabolic re-programming, and contributing to developing new therapeutic strategies and biomarkers [11, 185].
It is expected that the research and applications targeting the intratumoral microbiota-metabolism-immune axis might progress to a breakthrough. The integration of advanced technologies such as spatial multi-omics would provide an entire picture of the interaction between intratumoral microbiota and host cell metabolism in in situ resolution, thereby precisely locating certain microbiota to affect the function of adjacent immune cells through which metabolic pathways [2]. Said high-resolution analysis can uncover key bacterial-host metabolic interaction nodes, providing targets for intervention. Secondly, clarifying the metabolic characteristics of the patient's individual microbiota will promote the development of personalized intervention strategies. The individual intervention strategies could be formulated based on the spectral differences in the metabolites from patient microbiota, improving the treatment accuracy [11].
Therapeutically, combined immunotherapy strategies are to be focused on. Arg1/Arg2 inhibitors and ODC inhibitors combined with ICB can produce a synergistic effect, which can restore the L-arginine level and reduce immunosuppressive polyamines in the TME, thereby increase tumor-infiltrating lymphocytes and enhance ICB efficacy (Table 2). In mouse models, Arg1/Arg2 inhibitors could significantly increase the arginine and reduce polyamine, while increasing immune cell infiltration and activation, further significantly enhancing the efficacy of anti-PD-1 therapy [138]. Moreover, engineered bacteria therapy also shows great potential. They can colonize in tumors and continuously produce beneficial metabolites or enzymes by modifying intestinal or intratumoral microbiota through synthetic biology, which can directly regulate the local immune response (Table 2). For example, Nissle 1917 expressing IAA activated the AHR pathway and induced increased CXCL9 and IFN-γ, together with enhancing CD8+ T cell infiltration, thereby inhibiting tumor growth and improving survival [181, 182]. This demonstrated that engineered bacteria-mediated metabolic intervention can trigger a persistent immune memory response to tumors. Finally, non-invasive biomarker monitoring based on the microbiota-metabolic axis is also an important direction. Changes in intratumoral microbiota and their metabolite spectra are expected to be useful for disease diagnosis and efficacy prediction (Table 2). The concentration of specific microbial metabolites in feces or blood and the composition characteristics of oral or fecal microbiota may become alternative indicators reflecting the immune metabolic state in TME [186, 187]. The development of kits for these biomarkers will provide non-invasive and highly sensitive tools for cancer screening and assessing, which would be expected to be able to achieve early warnings and real-time evaluation of tumor progression and treatment response.
6. Conclusion
In conclusion, the research on intratumoral microbiota-arginine/ornithine metabolism-immune axis is revealing the complex and crucial regulatory levels within the TME. It integrates the tumor evolution in microbiology, metabolism and immunology, which not only deepen the understanding of the immunosuppressive mechanisms in the TME, but also provide novel perspectives for the development of anti-cancer strategies. With the application of multidisciplinary technologies and mechanism being studied, we might more effectively control intratumoral microbiota and its metabolites, further converting them into clinical advantages and establishing a new model for tumor diagnosis and treatment based on microbiota metabolic characteristics. In future, precise intervention of intra-tumoral microbiota-metabolism pathways and integration of immunotherapy approaches may generate a breakthrough for combating the tumor prognosis and improving treatment efficacy.
Abbreviations
TME: Tumor microenvironment; Arg: Arginase; ADI: Arginine deiminase; Treg: Regulatory T cells; GPR43: G protein-coupled receptor 43; AHR: Aryl hydrocarbon receptors; ODC: Ornithine decarboxylase; SCFAs: Short chain fatty acids; OTC: Ornithine transcarbamylase; ASS1: Argininosuccinate synthetase; ASL: Argininosuccinatelyase; MDSCs: Myeloid-derived suppressor cells; iNOS: Inducible nitric oxide synthase; DCs: Dendritic cells; ESCC: Esophageal squamous cell carcinoma; IDO: Indoleamine 2,3-dioxygenase; GPCRs: G protein-coupled receptors; HDACi: Histone deacetylase inhibitor; I3A: Indole-3-carboxaldehyde; IPA: Indole-3-propionic acid; IAA: Indole-3-acetic acid; PDAC: Pancreatic ductal adenocarcinoma; CTL: Cytotoxic T lymphocyte; Th1: Helper T cell; CRC: Colorectal cancer; AST: Arginine succinyltransferase; HNSCC: Head and neck squamous cell carcinoma; DFMO: Difluoromethyl ornithine; LPS: Lipopolysaccharide; ICB: Immune checkpoint blockade; TAM: tumor-associated macrophages.
Acknowledgements
The authors appreciate all members of Wang's Lab for helpful insights and BioRender for providing help on our graphical abstract and figures.
Funding
This research was funded by the National key research and development plan, grant number No. 2023YFC2705400; National Natural Science Foundation of China, grant number 82472965; Wuhan Knowledge Innovation Special project, grant number 202202080101045.
Data availability
The data presented in this study are available in article.
Artificial intelligence (AI) tools
The authors declare that no Generative AI was used in the creation of this manuscript.
Author contributions
Conceptualization, methodology, resources, data curation, writing—original draft preparation, S.Y.; validation, formal analysis, investigation, J.C.; visualization, supervision, W.S., G.C., X.S., S.C., K.D.; writing—review and editing, project administration, funding acquisition, H.W. All authors have read and agreed to the published version of the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Xu J, Cheng M, Liu J, Cui M, Yin B, Liang J. Research progress on the impact of intratumoral microbiota on the immune microenvironment of malignant tumors and its role in immunotherapy. Front Immunol. 2024;15:1389446
2. Galeano Niño JL, Wu H, LaCourse KD, Kempchinsky AG, Baryiames A, Barber B. et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature. 2022;611:810-7
3. Zhang X, Qi J, Dong C, Zhang L, Chen J, Liu J. et al. Intratumoral microbiota in colorectal cancer: roles, therapeutic potential, and challenges. J Adv Res. 2025
4. Sepich-Poore GD, Carter H, Knight R. Intratumoral bacteria generate a new class of therapeutically relevant tumor antigens in melanoma. Cancer Cell. 2021;39:601-3
5. Liu N-N, Yi C-X, Wei L-Q, Zhou J-A, Jiang T, Hu C-C. et al. The intratumor mycobiome promotes lung cancer progression via myeloid-derived suppressor cells. Cancer Cell. 2023 41
6. Fu A, Yao B, Dong T, Chen Y, Yao J, Liu Y. et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell. 2022 185
7. Li C, Cai C, Wang C, Chen X, Zhang B, Huang Z. Gut microbiota-mediated gut-liver axis: a breakthrough point for understanding and treating liver cancer. Clin Mol Hepatol. 2024;31:350-81
8. Luo D, Chen Q, Li Y, Yang J, Tao Y, Ji L. et al. Microbiome-metabolome interplay in pancreatic cancer progression: insights from multi-omics analysis. Mol Cancer. 2025;24:240
9. Nejman D, Livyatan I, Fuks G, Gavert N, Zwang Y, Geller LT. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science. 2020;368:973-80
10. Morad G, Damania AV, Melendez B, Singh BB, Veguilla FJ, Soto RA. et al. Microbial signals in primary and metastatic brain tumors. Nat Med. 2025;31:3675-88
11. Cao Y, Xia H, Tan X, Shi C, Ma Y, Meng D. et al. Intratumoural microbiota: a new frontier in cancer development and therapy. Signal Transduct Target Ther. 2024;9:15
12. Peng F, Hu M, Su Z, Hu L, Guo L, Yang K. Intratumoral Microbiota as a Target for Advanced Cancer Therapeutics. Adv Mater. 2024;36:e2405331
13. Feng T, Xie F, Lyu Y, Yu P, Chen B, Yu J. et al. The arginine metabolism and its deprivation in cancer therapy. Cancer Lett. 2025;620:217680
14. Wu Q, Schapira M, Arrowsmith CH, Barsyte-Lovejoy D. Protein arginine methylation: from enigmatic functions to therapeutic targeting. Nat Rev Drug Discov. 2021;20:509-30
15. Canè S, Geiger R, Bronte V. The roles of arginases and arginine in immunity. Nat Rev Immunol. 2024;25:266-84
16. Lee M-S, Dennis C, Naqvi I, Dailey L, Lorzadeh A, Ye G. et al. Ornithine aminotransferase supports polyamine synthesis in pancreatic cancer. Nature. 2023;616:339-47
17. Zhu Y, Zhou Z, Du X, Lin X, Liang Z-M, Chen S. et al. Cancer cell-derived arginine fuels polyamine biosynthesis in tumor-associated macrophages to promote immune evasion. Cancer Cell. 2025 43
18. Yuan L, Pan L, Wang Y, Zhao J, Fang L, Zhou Y. et al. Characterization of the landscape of the intratumoral microbiota reveals that Streptococcus anginosus increases the risk of gastric cancer initiation and progression. Cell Discov. 2024;10:117
19. Ding N, Cheng Y, Liu H, Wu Y, Weng Y, Cui H. et al. Fusobacterium nucleatum Infection Induces Malignant Proliferation of Esophageal Squamous Cell Carcinoma Cell by Putrescine Production. Microbiol Spectr. 2023;11:e0275922
20. Yurdagul A, Subramanian M, Wang X, Crown SB, Ilkayeva OR, Darville L. et al. Macrophage Metabolism of Apoptotic Cell-Derived Arginine Promotes Continual Efferocytosis and Resolution of Injury. Cell Metab. 2020 31
21. Fu K, Cheung AHK, Wong CC, Liu W, Zhou Y, Wang F. et al. Streptococcus anginosus promotes gastric inflammation, atrophy, and tumorigenesis in mice. Cell. 2024 187
22. Meza-Perez S, Liu M, Silva-Sanchez A, Morrow CD, Eipers PG, Lefkowitz EJ. et al. Proteobacteria impair anti-tumor immunity in the omentum by consuming arginine. Cell Host Microbe. 2024 32
23. Du X, Jie Z, Zou Q. Microbiota alert: Proteobacteria consume arginine to dampen omental antitumor immunity. Cell Host Microbe. 2024;32:1045-7
24. Hu X, Li Y, Cao Y, Shi F, Shang L. The role of nitric oxide synthase/ nitric oxide in infection-related cancers: Beyond antimicrobial activity. Biochim Biophys Acta Rev Cancer. 2024;1879:189156
25. Chaturvedi R, Asim M, Hoge S, Lewis ND, Singh K, Barry DP. et al. Polyamines Impair Immunity to Helicobacter pylori by Inhibiting L-Arginine Uptake Required for Nitric Oxide Production. Gastroenterology. 2010 139
26. Xu S, Zhang Y, Ding X, Yang Y, Gao J, Zou N. et al. Intestinal microbiota affects the progression of colorectal cancer by participating in the host intestinal arginine catabolism. Cell Rep. 2025;44:115370
27. Prasad YR, Anakha J, Pande AH. Treating liver cancer through arginine depletion. Drug Discov Today. 2024;29:103940
28. Yao S, Janku F, Subbiah V, Stewart J, Patel SP, Kaseb A. et al. Phase 1 trial of ADI-PEG20 plus cisplatin in patients with pretreated metastatic melanoma or other advanced solid malignancies. Br J Cancer. 2021;124:1533-9
29. Przystal JM, Hajji N, Khozoie C, Renziehausen A, Zeng Q, Abaitua F. et al. Efficacy of arginine depletion by ADI-PEG20 in an intracranial model of GBM. Cell Death Dis. 2018;9:1192
30. Feng Q, Wang H, Shao Y, Xu X. Antizyme inhibitor family: biological and translational research implications. Cell Commun Signal. 2024;22:11
31. Holbert CE, Cullen MT, Casero RA, Stewart TM. Polyamines in cancer: integrating organismal metabolism and antitumour immunity. Nat Rev Cancer. 2022;22:467-80
32. Proietti E, Rossini S, Grohmann U, Mondanelli G. Polyamines and Kynurenines at the Intersection of Immune Modulation. Trends Immunol. 2020;41:1037-50
33. Mondanelli G, Bianchi R, Pallotta MT, Orabona C, Albini E, Iacono A. et al. A Relay Pathway between Arginine and Tryptophan Metabolism Confers Immunosuppressive Properties on Dendritic Cells. Immunity. 2017;46:233-44
34. Soda K. The mechanisms by which polyamines accelerate tumor spread. J Exp Clin Cancer Res. 2011;30:95
35. Zepeda-Rivera M, Minot SS, Bouzek H, Wu H, Blanco-Míguez A, Manghi P. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature. 2024;628:424-32
36. Nomoto D, Baba Y, Liu Y, Tsutsuki H, Okadome K, Harada K. et al. Fusobacterium nucleatum promotes esophageal squamous cell carcinoma progression via the NOD1/RIPK2/NF-κB pathway. Cancer Lett. 2022;530:59-67
37. Brennan CA, Garrett WS. Fusobacterium nucleatum - symbiont, opportunist and oncobacterium. Nat Rev Microbiol. 2019;17:156-66
38. Mondal T, Chattopadhyay D, Saha Mondal P, Das S, Mondal A, Das A. et al. Fusobacterium nucleatum modulates the Wnt/β-catenin pathway in colorectal cancer development. Int J Biol Macromol. 2025;299:140196
39. Gobert AP, Wilson KT. Polyamine- and NADPH-dependent generation of ROS during Helicobacter pylori infection: A blessing in disguise. Free Radic Biol Med. 2016;105:16-27
40. Chenna SS, Gajula SNR, Nalla LV. Polyamine metabolism in cancer: drivers of immune evasion, ferroptosis and therapy resistance. Expert Rev Mol Med. 2025;27:e39
41. Riabov V, Gudima A, Wang N, Mickley A, Orekhov A, Kzhyshkowska J. Role of tumor associated macrophages in tumor angiogenesis and lymphangiogenesis. Front Physiol. 2014;5:75
42. Schoppmann SF, Birner P, Stöckl J, Kalt R, Ullrich R, Caucig C. et al. Tumor-associated macrophages express lymphatic endothelial growth factors and are related to peritumoral lymphangiogenesis. Am J Pathol. 2002;161:947-56
43. Lund AW, Duraes FV, Hirosue S, Raghavan VR, Nembrini C, Thomas SN. et al. VEGF-C promotes immune tolerance in B16 melanomas and cross-presentation of tumor antigen by lymph node lymphatics. Cell Rep. 2012;1:191-9
44. Morris SM. Regulation of enzymes of the urea cycle and arginine metabolism. Annu Rev Nutr. 2002 22
45. Zhou J, Sun X, Jiang P. The emerging roles of the urea cycle in tumor microenvironment and therapies. Trends Cancer. 2025
46. Yang C, Pataskar A, Feng X, Montenegro Navarro J, Paniagua I, Jacobs JJL. et al. Arginine deprivation enriches lung cancer proteomes with cysteine by inducing arginine-to-cysteine substitutants. Mol Cell. 2024 84
47. Chu Y-D, Lai M-W, Yeh C-T. Unlocking the Potential of Arginine Deprivation Therapy: Recent Breakthroughs and Promising Future for Cancer Treatment. Int J Mol Sci. 2023 24
48. De Santo C, Cheng P, Beggs A, Egan S, Bessudo A, Mussai F. Metabolic therapy with PEG-arginase induces a sustained complete remission in immunotherapy-resistant melanoma. J Hematol Oncol. 2018;11:68
49. Hernandez CP, Morrow K, Lopez-Barcons LA, Zabaleta J, Sierra R, Velasco C. et al. Pegylated arginase I: a potential therapeutic approach in T-ALL. Blood. 2010;115:5214-21
50. Trihemasava K, Bailis W. Dumping polyamines helps regulatory T cells "repair" a cancerous relationship. Immunity. 2025;58:1885-7
51. Jiang S, Ren B, Ding C, Du C, Cao Z, Yang G. et al. Polyamines in pancreatic cancer: reshaping the immunosuppressive tumor microenvironment. Cancer Lett. 2025;633:218016
52. Bell HN, Huber AK, Singhal R, Korimerla N, Rebernick RJ, Kumar R. et al. Microenvironmental ammonia enhances T cell exhaustion in colorectal cancer. Cell Metab. 2022 35
53. Spinelli JB, Yoon H, Ringel AE, Jeanfavre S, Clish CB, Haigis MC. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science. 2017;358:941-6
54. Matos A, Carvalho M, Bicho M, Ribeiro R. Arginine and Arginases Modulate Metabolism, Tumor Microenvironment and Prostate Cancer Progression. Nutrients. 2021 13
55. Wei J, Chen Y, Li Q, Huang F, Zhang C, Hou B. et al. Ammonia in cancer: dual roles and therapeutic strategies. Cancer Cell Int. 2025;25:420
56. Tharp KM, Kersten K, Maller O, Timblin GA, Stashko C, Canale FP. et al. Tumor-associated macrophages restrict CD8+ T cell function through collagen deposition and metabolic reprogramming of the breast cancer microenvironment. Nat Cancer. 2024;5:1045-62
57. Yadav P, Gómez Ortega J, Dabral P, Tamaki W, Chien C, Chang K-C. et al. Myeloid-mesenchymal crosstalk drives ARG1-dependent profibrotic metabolism via ornithine in lung fibrosis. J Clin Invest. 2025 135
58. Sivashanmugam M, J J, V U, K N S. Ornithine and its role in metabolic diseases: An appraisal. Biomed Pharmacother. 2016;86:185-94
59. Yang J-S, Wang C-C, Qiu J-D, Ren B, You L. Arginine metabolism: a potential target in pancreatic cancer therapy. Chin Med J (Engl). 2020;134:28-37
60. Riquelme E, Zhang Y, Zhang L, Montiel M, Zoltan M, Dong W. et al. Tumor Microbiome Diversity and Composition Influence Pancreatic Cancer Outcomes. Cell. 2019 178
61. Hezaveh K, Shinde RS, Klötgen A, Halaby MJ, Lamorte S, Ciudad MT. et al. Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity. 2022 55
62. Xie J, Ni F, Shao J, Zhang D, Guo T, Wang X. Amino acid metabolic reprogramming in clear cell renal cell carcinoma: Pathogenic mechanisms and therapeutic targeting. Cancer Lett. 2026;640:218244
63. Wang D, Cai J, Pei Q, Yan Z, Zhu F, Zhao Z. et al. Gut microbial alterations in arginine metabolism determine bone mechanical adaptation. Cell Metab. 2024 36
64. Tillett BJ, Dwiyanto J, Secombe KR, George T, Zhang V, Anderson D. et al. SCFA biotherapy delays diabetes in humanized gnotobiotic mice by remodeling mucosal homeostasis and metabolome. Nat Commun. 2025;16:2893
65. Depommier C, Everard A, Druart C, Maiter D, Thissen J-P, Loumaye A. et al. Serum metabolite profiling yields insights into health promoting effect of A. muciniphila in human volunteers with a metabolic syndrome. Gut Microbes. 2021;13:1994270
66. Jiang Z-M, Fang Z-Y, Yang X, Ji X-X, Zhao Y-Y, Lin B-Y. et al. Glycyrrhetinic acid ameliorates gastric mucosal injury by modulating gut microbiota and its metabolites via Thbs1/PI3K-Akt/p53 pathway. Phytomedicine. 2025;142:156745
67. Li S, Duan Y, Luo S, Zhou F, Wu Q, Lu Z. Short-chain fatty acids and cancer. Trends Cancer. 2024;11:154-68
68. Chun E, Lavoie S, Fonseca-Pereira D, Bae S, Michaud M, Hoveyda HR. et al. Metabolite-Sensing Receptor Ffar2 Regulates Colonic Group 3 Innate Lymphoid Cells and Gut Immunity. Immunity. 2019 51
69. Xu Z, Wang T, Wang Y, Li Y, Sun Y, Qiu H-J. Short-chain fatty acids: key antiviral mediators of gut microbiota. Front Immunol. 2025;16:1614879
70. Mukhopadhya I, Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol. 2025;23:635-51
71. Fusco W, Lorenzo MB, Cintoni M, Porcari S, Rinninella E, Kaitsas F. et al. Short-Chain Fatty-Acid-Producing Bacteria: Key Components of the Human Gut Microbiota. Nutrients. 2023 15
72. Mann ER, Lam YK, Uhlig HH. Short-chain fatty acids: linking diet, the microbiome and immunity. Nat Rev Immunol. 2024;24:577-95
73. Dong D, Wang H, Bi H, Li Y, Gao T, Feng J. et al. A pectic polysaccharide from Lycium ruthenicum Murray alleviates dextran sulfate sodium-induced colitis in mice. Curr Res Food Sci. 2024;10:100955
74. Laudes M, Geisler C, Rohmann N, Bouwman J, Pischon T, Schlicht K. Microbiota in Health and Disease-Potential Clinical Applications. Nutrients. 2021 13
75. Chen B, Guan L, Wu C, Gong Y, Wu L, Zhang M. et al. Gut Microbiota-Butyrate-PPARγ Axis Modulates Adipose Regulatory T Cell Population. Adv Sci (Weinh). 2025;12:e2411086
76. Masuyama S, Mizui M, Morita M, Shigeki T, Kato H, Yamamoto T. et al. Enhanced fatty acid oxidation by selective activation of PPARα alleviates autoimmunity through metabolic transformation in T-cells. Clin Immunol. 2024;268:110357
77. Laragione T, Harris C, Azizgolshani N, Beeton C, Bongers G, Gulko PS. Magnesium increases numbers of Foxp3+ Treg cells and reduces arthritis severity and joint damage in an IL-10-dependent manner mediated by the intestinal microbiome. EBioMedicine. 2023;92:104603
78. Li Q, Liu H, Bai C, Jiang L, Su C, Qin X. et al. A high-calorie diet exacerbates lipopolysaccharide-induced pneumonia by promoting acetate-mediated macrophage polarization via the HDAC9/10-HIF-1α-glycolysis axis. Front Immunol. 2025;16:1614768
79. Maslowski KM, Vieira AT, Ng A, Kranich J, Sierro F, Yu D. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature. 2009;461:1282-6
80. Li Y, Li Q, Yuan R, Wang Y, Guo C, Wang L. Bifidobacterium breve-derived indole-3-lactic acid ameliorates colitis-associated tumorigenesis by directing the differentiation of immature colonic macrophages. Theranostics. 2024;14:2719-35
81. Bachem A, Makhlouf C, Binger KJ, de Souza DP, Tull D, Hochheiser K. et al. Microbiota-Derived Short-Chain Fatty Acids Promote the Memory Potential of Antigen-Activated CD8+ T Cells. Immunity. 2019 51
82. Yang W, Yu T, Huang X, Bilotta AJ, Xu L, Lu Y. et al. Intestinal microbiota-derived short-chain fatty acids regulation of immune cell IL-22 production and gut immunity. Nat Commun. 2020;11:4457
83. Seo S-K, Kwon B. Immune regulation through tryptophan metabolism. Exp Mol Med. 2023;55:1371-9
84. Li B, Liang X, Li Y, Wang R, Wei Y, Liu Q. et al. Tryptophan catabolites from microbiota ameliorate immune-mediated hepatitis through activating aryl hydrocarbon receptor of T cells. Gut Microbes. 2025;17:2557979
85. Situ Y, Zhang P, Zhang C, Jiang A, Zhang N, Zhu L. et al. The metabolic dialogue between intratumoural microbes and cancer: implications for immunotherapy. EBioMedicine. 2025;115:105708
86. Bender MJ, McPherson AC, Phelps CM, Pandey SP, Laughlin CR, Shapira JH. et al. Dietary tryptophan metabolite released by intratumoral Lactobacillus reuteri facilitates immune checkpoint inhibitor treatment. Cell. 2023 186
87. Sadik A, Somarribas Patterson LF, Öztürk S, Mohapatra SR, Panitz V, Secker PF. et al. IL4I1 Is a Metabolic Immune Checkpoint that Activates the AHR and Promotes Tumor Progression. Cell. 2020 182
88. DeNardo DG, Ruffell B. Macrophages as regulators of tumour immunity and immunotherapy. Nat Rev Immunol. 2019;19:369-82
89. Halbrook CJ, Pontious C, Kovalenko I, Lapienyte L, Dreyer S, Lee H-J. et al. Macrophage-Released Pyrimidines Inhibit Gemcitabine Therapy in Pancreatic Cancer. Cell Metab. 2019 29
90. Yan J, Chen D, Ye Z, Zhu X, Li X, Jiao H. et al. Molecular mechanisms and therapeutic significance of Tryptophan Metabolism and signaling in cancer. Mol Cancer. 2024;23:241
91. Yang L, Chu Z, Liu M, Zou Q, Li J, Liu Q. et al. Amino acid metabolism in immune cells: essential regulators of the effector functions, and promising opportunities to enhance cancer immunotherapy. J Hematol Oncol. 2023;16:59
92. Sancak Y, Peterson TR, Shaul YD, Lindquist RA, Thoreen CC, Bar-Peled L. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science. 2008;320:1496-501
93. Mossmann D, Park S, Hall MN. mTOR signalling and cellular metabolism are mutual determinants in cancer. Nat Rev Cancer. 2018;18:744-57
94. Chantranupong L, Scaria SM, Saxton RA, Gygi MP, Shen K, Wyant GA. et al. The CASTOR Proteins Are Arginine Sensors for the mTORC1 Pathway. Cell. 2016;165:153-64
95. Wyant GA, Abu-Remaileh M, Wolfson RL, Chen WW, Freinkman E, Danai LV. et al. mTORC1 Activator SLC38A9 Is Required to Efflux Essential Amino Acids from Lysosomes and Use Protein as a Nutrient. Cell. 2017 171
96. Liu GY, Sabatini DM. mTOR at the nexus of nutrition, growth, ageing and disease. Nat Rev Mol Cell Biol. 2020;21:183-203
97. Wang G, Dai S, Chen J, Zhang K, Huang C, Zhang J. et al. USP19 potentiates autophagic cell death via inhibiting mTOR pathway through deubiquitinating NEK9 in pancreatic cancer. Cell Death Differ. 2024;32:702-13
98. Canale FP, Basso C, Antonini G, Perotti M, Li N, Sokolovska A. et al. Metabolic modulation of tumours with engineered bacteria for immunotherapy. Nature. 2021;598:662-6
99. Li T, Wang W. New strategy for cancer immunotherapy: using live engineered bacteria for metabolic modulation. Signal Transduct Target Ther. 2021;6:415
100. Luu M, Riester Z, Baldrich A, Reichardt N, Yuille S, Busetti A. et al. Microbial short-chain fatty acids modulate CD8+ T cell responses and improve adoptive immunotherapy for cancer. Nat Commun. 2021;12:4077
101. Sun M, Wu W, Chen L, Yang W, Huang X, Ma C. et al. Microbiota-derived short-chain fatty acids promote Th1 cell IL-10 production to maintain intestinal homeostasis. Nat Commun. 2018;9:3555
102. Saxton RA, Sabatini DM. mTOR Signaling in Growth, Metabolism, and Disease. Cell. 2017;168:960-76
103. Flanagan L, Schmid J, Ebert M, Soucek P, Kunicka T, Liska V. et al. Fusobacterium nucleatum associates with stages of colorectal neoplasia development, colorectal cancer and disease outcome. Eur J Clin Microbiol Infect Dis. 2014;33:1381-90
104. Mima K, Nishihara R, Qian ZR, Cao Y, Sukawa Y, Nowak JA. et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut. 2015;65:1973-80
105. Chen Y, Lu Y, Ke Y, Li Y. Prognostic impact of the Fusobacterium nucleatum status in colorectal cancers. Medicine (Baltimore). 2019;98:e17221
106. Zhang Y, Zhang L, Zheng S, Li M, Xu C, Jia D. et al. Fusobacterium nucleatum promotes colorectal cancer cells adhesion to endothelial cells and facilitates extravasation and metastasis by inducing ALPK1/NF-κB/ICAM1 axis. Gut Microbes. 2022;14:2038852
107. Wang N, Fang J-Y. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 2022;31:159-72
108. Rubinstein MR, Baik JE, Lagana SM, Han RP, Raab WJ, Sahoo D. et al. Fusobacterium nucleatum promotes colorectal cancer by inducing Wnt/β-catenin modulator Annexin A1. EMBO Rep. 2019 20
109. Song Q, Jin Z, Zhang H, Hong K, Zhu B, Yin H. et al. Fusobacterium nucleatum-derived 3-indolepropionic acid promotes colorectal cancer progression via aryl hydrocarbon receptor activation in macrophages. Chem Biol Interact. 2025;414:111495
110. Qu R, Zhang Y, Ma Y, Zhou X, Sun L, Jiang C. et al. Role of the Gut Microbiota and Its Metabolites in Tumorigenesis or Development of Colorectal Cancer. Adv Sci (Weinh). 2023;10:e2205563
111. Jian C, Yinhang W, Jing Z, Zhanbo Q, Zefeng W, Shuwen H. Escherichia coli on colorectal cancer: A two-edged sword. Microb Biotechnol. 2024;17:e70029
112. Peyraud F, Guégan JP, Bodet D, Nafia I, Fontan L, Auzanneau C. et al. Circulating L-arginine predicts the survival of cancer patients treated with immune checkpoint inhibitors. Ann Oncol. 2022;33:1041-51
113. Pushalkar S, Hundeyin M, Daley D, Zambirinis CP, Kurz E, Mishra A. et al. The Pancreatic Cancer Microbiome Promotes Oncogenesis by Induction of Innate and Adaptive Immune Suppression. Cancer Discov. 2018;8:403-16
114. Nalluri H, Jensen E, Staley C. Role of biliary stent and neoadjuvant chemotherapy in the pancreatic tumor microbiome. BMC Microbiol. 2021;21:280
115. Geller LT, Barzily-Rokni M, Danino T, Jonas OH, Shental N, Nejman D. et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357:1156-60
116. Zhu Y, He C, Li X, Cai Y, Hu J, Liao Y. et al. Gut microbiota dysbiosis worsens the severity of acute pancreatitis in patients and mice. J Gastroenterol. 2018;54:347-58
117. Dickson I. Microbiome promotes pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2018;15:328
118. Tabrizi E, Pourteymour Fard Tabrizi F, Mahmoud Khaled G, Sestito MP, Jamie S, Boone BA. Unraveling the gut microbiome's contribution to pancreatic ductal adenocarcinoma: mechanistic insights and therapeutic perspectives. Front Immunol. 2024;15:1434771
119. Tintelnot J, Xu Y, Lesker TR, Schönlein M, Konczalla L, Giannou AD. et al. Microbiota-derived 3-IAA influences chemotherapy efficacy in pancreatic cancer. Nature. 2023;615:168-74
120. Jia K, Chen Y, Xie Y, Wang X, Hu Y, Sun Y. et al. Helicobacter pylori and immunotherapy for gastrointestinal cancer. Innovation (Camb). 2024;5:100561
121. Nilsson H-O, Stenram U, Ihse I, Wadstrom T. Helicobacter species ribosomal DNA in the pancreas, stomach and duodenum of pancreatic cancer patients. World J Gastroenterol. 2006;12:3038-43
122. Ju Y, Xu D, Liao M-M, Sun Y, Bao W-D, Yao F. et al. Barriers and opportunities in pancreatic cancer immunotherapy. NPJ Precis Oncol. 2024;8:199
123. Tan Q, Cao X, Zou F, Wang H, Xiong L, Deng S. Spatial Heterogeneity of Intratumoral Microbiota: A New Frontier in Cancer Immunotherapy Resistance. Biomedicines. 2025 13
124. Bučević Popović V, Šitum M, Chow C-ET, Chan LS, Roje B, Terzić J. The urinary microbiome associated with bladder cancer. Sci Rep. 2018;8:12157
125. Jansson OT, Morcos E, Brundin L, Lundberg JO, Adolfsson J, Söderhäll M. et al. The role of nitric oxide in bacillus Calmette-Guérin mediated anti-tumour effects in human bladder cancer. Br J Cancer. 1998;78:588-92
126. Sahu D, Gupta S, Hau AM, Nakashima K, Leivo MZ, Searles SC. et al. Argininosuccinate Synthetase 1 Loss in Invasive Bladder Cancer Regulates Survival through General Control Nonderepressible 2 Kinase-Mediated Eukaryotic Initiation Factor 2α Activity and Is Targetable by Pegylated Arginine Deiminase. Am J Pathol. 2016;187:200-13
127. Lee H, Park G, Kim S, Son B, Joo J, Park HH. et al. Enhancement of anti-tumor activity in melanoma using arginine deiminase fused with 30Kc19α protein. Appl Microbiol Biotechnol. 2022;106:7531-45
128. Bai D, Zhou Y, Jing L, Guo C, Yang Q. Arginine Metabolism in Cancer Biology and Immunotherapy. Immune Netw. 2025;25:e30
129. Bautista J, Villegas-Chávez JA, Bunces-Larco D, Martín-Aguilera R, López-Cortés A. The microbiome as a therapeutic co-driver in melanoma immuno-oncology. Front Med (Lausanne). 2025;12:1673880
130. Yang X, Yin L, Tian Z, Zhou Q. Intratumoral Microbiota in Lung Cancer: Emerging Roles in TME Modulation and Immunotherapy Response. Int J Mol Sci. 2025 27
131. Chen J, Gao Y, Chen Y, Wang Q, Zhang Y, Huang Y. et al. Identification and validation of intratumoral microbiome associated with sensitization to immune checkpoint inhibitors. Cell Rep Med. 2025;6:102306
132. Szlosarek PW, Steele JP, Nolan L, Gilligan D, Taylor P, Spicer J. et al. Arginine Deprivation With Pegylated Arginine Deiminase in Patients With Argininosuccinate Synthetase 1-Deficient Malignant Pleural Mesothelioma: A Randomized Clinical Trial. JAMA Oncol. 2017;3:58-66
133. Szlosarek PW, Creelan BC, Sarkodie T, Nolan L, Taylor P, Olevsky O. et al. Pegargiminase Plus First-Line Chemotherapy in Patients With Nonepithelioid Pleural Mesothelioma: The ATOMIC-Meso Randomized Clinical Trial. JAMA Oncol. 2024;10:475-83
134. Dzutsev A, Badger JH, Perez-Chanona E, Roy S, Salcedo R, Smith CK. et al. Microbes and Cancer. Annu Rev Immunol. 2017;35:199-228
135. Zhang M, Bzura A, Baitei EY, Zhou Z, Spicer JB, Poile C. et al. A gut microbiota rheostat forecasts responsiveness to PD-L1 and VEGF blockade in mesothelioma. Nat Commun. 2024;15:7187
136. Zheng D-W, Dong X, Pan P, Chen K-W, Fan J-X, Cheng S-X. et al. Phage-guided modulation of the gut microbiota of mouse models of colorectal cancer augments their responses to chemotherapy. Nat Biomed Eng. 2019;3:717-28
137. Fu K, Cheung AHK, Wong CC, Liu W, Zhou Y, Wang F. et al. Streptococcus anginosus promotes gastric inflammation, atrophy, and tumorigenesis in mice. Cell. 2024;187:6783
138. Grzybowski MM, Uçal Y, Muchowicz A, Rejczak T, Kikulska A, Głuchowska KM. et al. Metabolomic reprogramming of the tumor microenvironment by dual arginase inhibitor OATD-02 boosts anticancer immunity. Sci Rep. 2025;15:18741
139. Pokrývková B, Šmahelová J, Dalewská N, Grega M, Vencálek O, Šmahel M. et al. ARG1 mRNA Level Is a Promising Prognostic Marker in Head and Neck Squamous Cell Carcinomas. Diagnostics (Basel). 2021 11
140. Ferreira RM, Pereira-Marques J, Pinto-Ribeiro I, Costa JL, Carneiro F, Machado JC. et al. Gastric microbial community profiling reveals a dysbiotic cancer-associated microbiota. Gut. 2017;67:226-36
141. Dejea CM, Fathi P, Craig JM, Boleij A, Taddese R, Geis AL. et al. Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science. 2018;359:592-7
142. Boleij A, Hechenbleikner EM, Goodwin AC, Badani R, Stein EM, Lazarev MG. et al. The Bacteroides fragilis toxin gene is prevalent in the colon mucosa of colorectal cancer patients. Clin Infect Dis. 2014;60:208-15
143. Hu Y-L, Pang W, Huang Y, Zhang Y, Zhang C-J. The Gastric Microbiome Is Perturbed in Advanced Gastric Adenocarcinoma Identified Through Shotgun Metagenomics. Front Cell Infect Microbiol. 2018;8:433
144. Zhang J-W, Zhang D, Yin H-S, Zhang H, Hong K-Q, Yuan J-P. et al. Fusobacterium nucleatum promotes esophageal squamous cell carcinoma progression and chemoresistance by enhancing the secretion of chemotherapy-induced senescence-associated secretory phenotype via activation of DNA damage response pathway. Gut Microbes. 2023;15:2197836
145. Li Y, Xing S, Chen F, Li Q, Dou S, Huang Y. et al. Intracellular Fusobacterium nucleatum infection attenuates antitumor immunity in esophageal squamous cell carcinoma. Nat Commun. 2023;14:5788
146. Goenka A, Khan F, Verma B, Sinha P, Dmello CC, Jogalekar MP. et al. Tumor microenvironment signaling and therapeutics in cancer progression. Cancer Commun (Lond). 2023;43:525-61
147. Lin Y, Lau HC-H, Liu C, Ding X, Sun Y, Rong J. et al. Multi-cohort analysis reveals colorectal cancer tumor location-associated fecal microbiota and their clinical impact. Cell Host Microbe. 2025 33
148. Moreira D, Sampath S, Won H, White SV, Su Y-L, Alcantara M. et al. Myeloid cell-targeted STAT3 inhibition sensitizes head and neck cancers to radiotherapy and T cell-mediated immunity. J Clin Invest. 2021 131
149. Hofmann L, Harasymczuk M, Huber D, Szczepanski MJ, Dworacki G, Whiteside TL. et al. Arginase-1 in Plasma-Derived Exosomes as Marker of Metastasis in Patients with Head and Neck Squamous Cell Carcinoma. Cancers (Basel). 2023 15
150. Sun Y, Gan M, Wu Y, Gao W, Lu Y. [Study on the difference and clinical value of serum amino acids in patients with laryngeal squamous cell carcinoma]. Lin Chuang Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2024;38:715-21
151. Yau T, Cheng PN, Chan P, Chan W, Chen L, Yuen J. et al. A phase 1 dose-escalating study of pegylated recombinant human arginase 1 (Peg-rhArg1) in patients with advanced hepatocellular carcinoma. Invest New Drugs. 2012 31
152. Mussai F, De Santo C, Cheng P, Thomas IF, Ariti C, Upton L. et al. A randomised evaluation of low-dose Ara-C plus pegylated recombinant arginase BCT-100 versus low dose Ara-C in older unfit patients with acute myeloid leukaemia: Results from the LI-1 trial. Br J Haematol. 2022;200:573-8
153. Thompson PA, Wertheim BC, Zell JA, Chen W-P, McLaren CE, LaFleur BJ. et al. Levels of rectal mucosal polyamines and prostaglandin E2 predict ability of DFMO and sulindac to prevent colorectal adenoma. Gastroenterology. 2010 139
154. Oesterheld J, Ferguson W, Kraveka JM, Bergendahl G, Clinch T, Lorenzi E. et al. Eflornithine as Postimmunotherapy Maintenance in High-Risk Neuroblastoma: Externally Controlled, Propensity Score-Matched Survival Outcome Comparisons. J Clin Oncol. 2023 42
155. Kuboki Y, Koyama T, Matsubara N, Naito Y, Kondo S, Harano K. et al. PD-1 inhibition with retifanlimab and/or arginase inhibition with INCB001158 in Japanese patients with solid tumors: A phase I study. Cancer Med. 2024;13:e6980
156. Lian J, Liang Y, Zhang H, Lan M, Ye Z, Lin B. et al. The role of polyamine metabolism in remodeling immune responses and blocking therapy within the tumor immune microenvironment. Front Immunol. 2022;13:912279
157. Hayes CS, Shicora AC, Keough MP, Snook AE, Burns MR, Gilmour SK. Polyamine-blocking therapy reverses immunosuppression in the tumor microenvironment. Cancer Immunol Res. 2013;2:274-85
158. Ye P-H, Li C-Y, Cheng H-Y, Anuraga G, Wang C-Y, Chen F-W. et al. A novel combination therapy of arginine deiminase and an arginase inhibitor targeting arginine metabolism in the tumor and immune microenvironment. Am J Cancer Res. 2023;13:1952-69
159. Chung S-F, Kim C-F, Tam S-Y, Choi M-C, So P-K, Wong K-Y. et al. A bioengineered arginine-depleting enzyme as a long-lasting therapeutic agent against cancer. Appl Microbiol Biotechnol. 2020;104:3921-34
160. Lynch JP, Goers L, Lesser CF. Emerging strategies for engineering Escherichia coli Nissle 1917-based therapeutics. Trends Pharmacol Sci. 2022;43:772-86
161. Lawrence GW, McCarthy N, Walsh CJ, Kunyoshi TM, Lawton EM, O'Connor PM. et al. Effect of a bacteriocin-producing Streptococcus salivarius on the pathogen Fusobacterium nucleatum in a model of the human distal colon. Gut Microbes. 2022;14:2100203
162. Alzamami A, Alturki NA, Khan K, Basharat Z, Mashraqi MM. Screening inhibitors against the Ef-Tu of Fusobacterium nucleatum: a docking, ADMET and PBPK assessment study. Mol Divers. 2024;28:4259-76
163. Liu H, Yu Y, Dong A, Elsabahy M, Yang Y-W, Gao H. Emerging strategies for combating Fusobacterium nucleatum in colorectal cancer treatment: Systematic review, improvements and future challenges. Exploration (Beijing). 2023;4:20230092
164. Lam HYP, Lai M-J, Wang P-C, Wu W-J, Chen L-K, Fan H-W. et al. A Novel Bacteriophage with the Potential to Inhibit Fusobacterium nucleatum-Induced Proliferation of Colorectal Cancer Cells. Antibiotics (Basel). 2025 14
165. Nguyen MHT, Netter Z, Angermeyer A, Seed KD. A phage weaponizes a satellite recombinase to subvert viral restriction. Nucleic Acids Res. 2022;50:11138-53
166. Wu Y, Zhang J, Wang B, Zhang Y, Li H, Liu Y. et al. Dissecting the Arginine and Lysine Biosynthetic Pathways and Their Relationship in Haloarchaeon Natrinema gari J7-2 via Endogenous CRISPR-Cas System-Based Genome Editing. Microbiol Spectr. 2023;11:e0028823
167. Ma C, Awosika JA, Monge C, Coffman-D'Annibale K, Xie C, Skorupan N. et al. Altering the gut microbiome and tumor microenvironment in advanced liver cancer: A phase II study of nivolumab, tadalafil and oral vancomycin in patients with refractory primary hepatocellular carcinoma or liver dominant metastatic cancer from colorectal or pancreatic cancers. Cancer Lett. 2025;632:217975
168. Prasad R, Rehman A, Rehman L, Darbaniyan F, Blumenberg V, Schubert M-L. et al. Antibiotic-induced loss of gut microbiome metabolic output correlates with clinical responses to CAR T-cell therapy. Blood. 2025;145:823-39
169. Glitza IC, Seo YD, Spencer CN, Wortman JR, Burton EM, Alayli FA. et al. Randomized Placebo-Controlled, Biomarker-Stratified Phase Ib Microbiome Modulation in Melanoma: Impact of Antibiotic Preconditioning on Microbiome and Immunity. Cancer Discov. 2024;14:1161-75
170. Murphy S, Rahmy S, Gan D, Liu G, Zhu Y, Manyak M. et al. Ketogenic Diet Alters the Epigenetic and Immune Landscape of Prostate Cancer to Overcome Resistance to Immune Checkpoint Blockade Therapy. Cancer Res. 2024;84:1597-612
171. Shao X, Liu L, Zhou Y, Zhong K, Gu J, Hu T. et al. High-fat diet promotes colitis-associated tumorigenesis by altering gut microbial butyrate metabolism. Int J Biol Sci. 2023;19:5004-19
172. Zhang T, Li Y, Zhai E, Zhao R, Qian Y, Huang Z. et al. Intratumoral Fusobacterium nucleatum Recruits Tumor-Associated Neutrophils to Promote Gastric Cancer Progression and Immune Evasion. Cancer Res. 2025;85:1819-41
173. Wang X, Fang Y, Liang W, Wong CC, Qin H, Gao Y. et al. Fusobacterium nucleatum facilitates anti-PD-1 therapy in microsatellite stable colorectal cancer. Cancer Cell. 2024 42
174. Steggerda SM, Bennett MK, Chen J, Emberley E, Huang T, Janes JR. et al. Inhibition of arginase by CB-1158 blocks myeloid cell-mediated immune suppression in the tumor microenvironment. J Immunother Cancer. 2017;5:101
175. Lv Y, Li M, Weng L, Huang H, Mao Y, Yang DA. et al. Ginseng-derived nanoparticles reprogram macrophages to regulate arginase-1 release for ameliorating T cell exhaustion in tumor microenvironment. J Exp Clin Cancer Res. 2023;42:322
176. Geng Z, Cao Z, Liu R, Liu K, Liu J, Tan W. Aptamer-assisted tumor localization of bacteria for enhanced biotherapy. Nat Commun. 2021;12:6584
177. Chowdhury S, Castro S, Coker C, Hinchliffe TE, Arpaia N, Danino T. Programmable bacteria induce durable tumor regression and systemic antitumor immunity. Nat Med. 2019;25:1057-63
178. Yoon W, Park YC, Kim J, Chae YS, Byeon JH, Min S-H. et al. Application of genetically engineered Salmonella typhimurium for interferon-gamma-induced therapy against melanoma. Eur J Cancer. 2016;70:48-61
179. Tucker MD, Rini BI. Predicting Response to Immunotherapy in Metastatic Renal Cell Carcinoma. Cancers (Basel). 2020 12
180. Donahue KL, Watkoske HR, Kadiyala P, Du W, Brown K, Scales MK. et al. Oncogenic KRAS-Dependent Stromal Interleukin-33 Directs the Pancreatic Microenvironment to Promote Tumor Growth. Cancer Discov. 2024;14:1964-89
181. Vaaben TH, Lützhøft DO, Koulouktsis A, Dawoodi IM, Stavnsbjerg C, Kvich L. et al. Modulating tumor immunity using advanced microbiome therapeutics producing an indole metabolite. EMBO Rep. 2025;26:1688-708
182. Zhong L, Boopathi S, Purushothaman B, Tu Q, Zhang Y. Gut microbiota - indole-3-acetic acid axis in cancer: dual functions, mechanistic insights, and therapeutic potential. Microbiol Res. 2025;300:128293
183. Liao K, Wen J, Liu Z, Zhang B, Zhang X, Fu Y. et al. The role of intratumoral microbiome in the occurrence, proliferation, metastasis of colorectal cancer and its underlying therapeutic strategies. Ageing Res Rev. 2025;111:102820
184. Franzosa EA, Hsu T, Sirota-Madi A, Shafquat A, Abu-Ali G, Morgan XC. et al. Sequencing and beyond: integrating molecular 'omics' for microbial community profiling. Nat Rev Microbiol. 2015;13:360-72
185. El Tekle G, Garrett WS. Bacteria in cancer initiation, promotion and progression. Nat Rev Cancer. 2023;23:600-18
186. Song P, Peng Z, Guo X. Gut microbial metabolites in cancer therapy. Trends Endocrinol Metab. 2024;36:55-69
187. Lu Y, Yuan X, Wang M, He Z, Li H, Wang J. et al. Gut microbiota influence immunotherapy responses: mechanisms and therapeutic strategies. J Hematol Oncol. 2022;15:47
Author contact
Corresponding author: Hongbo Wang (Email: drwanghb69edu.cn).

 Global reach, higher impact
Global reach, higher impact