Impact Factor ISSN: 1449-1907
Int J Med Sci 2026; 23(4):1278-1284. doi:10.7150/ijms.129445 This issue Cite
Research Paper
Association of MIAT genetic variants and expression with tumor grade in oral tongue cancer
1. Department of Dentistry, Changhua Christian Hospital, Changhua, Taiwan.
2. Department of Post-Baccalaureate Medicine, College of Medicine, National Chung Hsing University, Taichung, Taiwan.
3. Institute of Medicine, Chung Shan Medical University, Taichung, Taiwan.
4. Institute of Oral Sciences, Chung Shan Medical University, Taichung, Taiwan.
5. Department of Dentistry, Chung Shan Medical University Hospital, Taichung, Taiwan.
6. School of Medicine, Chung Shan Medical University, Taichung, Taiwan.
7. Department of Otolaryngology, Chung Shan Medical University Hospital, Taichung, Taiwan.
8. Department of Medical Research, Chung Shan Medical University Hospital, Taichung, Taiwan.
Received 2025-12-4; Accepted 2026-2-13; Published 2026-2-26
Abstract
Oral tongue cancer is an aggressive malignancy and represents the most common subsite of head and neck cancer. The long non-coding RNA myocardial infarction associated transcript (MIAT) has been implicated in the development and progression of several cancers. This study aimed to investigate the association between MIAT single-nucleotide polymorphisms (SNPs) and oral cancer susceptibility as well as related clinicopathological characteristics. In this study, MIAT SNPs (rs4274, rs1061540, and rs1894720) were genotyped using real-time polymerase chain reaction in 1,194 controls and 397 male patients with tongue cancer. A significant association was observed between the MIAT rs4274 AA genotype and the occurrence of tongue cancer compared with controls (p = 0.018). Among the patients, carriers of the rs4274 A allele exhibited a higher likelihood of developing moderately or poorly differentiated tumors [OR (95% CI) = 2.246 (1.217-4.147); p = 0.001]. Betel-quid chewers and smokers carrying the A allele showed similarly elevated risks for poor tumor differentiation. Bioinformatic analyses further indicated that the rs4274 A allele is associated with increased MIAT expression, and higher MIAT levels correlated with higher tumor grade. In conclusion, the MIAT rs4274 A allele is linked to poorer tumor differentiation, particularly among betel-quid chewers and smokers, and elevated MIAT expression supports its potential as a biomarker of tumor aggressiveness.
Keywords: oral cancer, MIAT, polymorphism, cell differentiation, betel-quid chewing
Introduction
Oral cancer is a major subtype of head and neck malignancies, encompassing cancers of the tongue, lips, oral cavity, and oropharynx [1, 2]. Among these, oral tongue squamous cell carcinoma (OTSCC) is the most prevalent, accounting for 25-40% of oral cancer cases [3, 4]. Exposure to environmental risk factors, including tobacco smoking, alcohol consumption, and betel quid chewing, has been identified as a significant contributor to oral carcinogenesis [5-8]. Despite extensive research, it remains unclear whether the functional consequences of genetic alterations are consistent across OTSCC. Consequently, further studies are warranted to evaluate the reproducibility and clinical significance of these genetic phenotypes in OTSCC.
Myocardial infarction associated transcript (MIAT) was first identified as a novel long non-coding RNA significantly associated with myocardial infarction (MI) on chromosome 22q12.1 [9]. In OTSCC, upregulated MIAT was shown to promote epithelial-mesenchymal transition (EMT) in OTSCC cells via activation of the Wnt/β-catenin signaling pathway [10]. Furthermore, several studies have investigated MIAT expression in oral squamous cell carcinoma (OSCC) and suggested that it may serve as a potential prognostic biomarker, reflecting its possible role in disease progression and patient outcomes [11, 12]. In addition, MIAT polymorphisms have been reported to be associated with cancer development and progression across multiple malignancies [9, 13-21]. For example, in ovarian cancer, the C/C genotype of MIAT rs1061540 was significantly more frequent in malignant epithelial tumors compared to low-grade tumors, whereas the T/T genotype was linked to a lower risk of malignancy, suggesting that specific MIAT genetic variants may influence tumor behavior and patient prognosis [14]. However, the associations between MIAT polymorphisms and oral tongue cancer progression, as well as clinicopathologic characteristics, remain unexplored. In this study, we examined three SNPs of MIAT rs4274, rs1061540, and rs1894720, and try to elucidate their associations to oral cancer susceptibility and clinicopathologic characteristics with environmental risk factors.
Materials and Methods
Study subjects
A total of 1194 cancer-free controls and 397 male patients with tongue cancer were participated in our study. All participants in this study were enrolled at Chung Shan Medical University Hospital, Taichung, Taiwan. Tumor differentiation in male patients with tongue cancer was evaluated and graded by pathologists according to the American Joint Committee on Cancer (AJCC) classification system. In addition, TNM staging of the male tongue cancer patients included in this study was clinically determined at the time of diagnosis in accordance with the AJCC guidelines. This project was approved by the institutional review board of Chung Shan Medical University Hospital (CSMUH No: CS1-21151). The informed written consent was provided to each patient who enrolled in this study. The control group who participated in our study was those individuals who without self-reported diseases such as asthma, autoimmune and neurological diseases, cardiovascular diseases, diabetes, and history of cancer of any sites.
Sample preparation and DNA extraction
Peripheral blood samples from normal controls and oral tongue cancer patients participating in this study were collected for genomic DNA extraction [22]. Each sample was preserved in EDTA-containing tubes. The blood samples were centrifuged at 3,000 rpm for 10 minutes, and the resulting buffy coats were collected for DNA extraction. Genomic DNA was isolated using the QIAamp DNA Blood Mini Kit according to the manufacturer's instructions. The final elution was performed with Tris-EDTA (TE) buffer. The extracted DNA was subsequently used as a template for real-time polymerase chain reaction (PCR) analyses.
Selection of MIAT SNPs
In this study, three MIAT SNPs rs4274, rs1061540, and rs1894720 were selected based on data from the International HapMap Project database [23]. The MIAT rs4274 polymorphism was chosen because carriers of the AA genotype have been reported to have an increased risk of paranoid schizophrenia in a Chinese Han population [21]. The MIAT rs1061540 polymorphism was selected due to its reported association with the severity of coronary artery disease (CAD) and the Gensini score in an Egyptian population [24]. In addition, a study on ovarian cancer demonstrated that malignant epithelial tumors exhibited a higher frequency of the MIAT rs1061540 C/C genotype compared with low-grade epithelial tumor cohorts [14]. Regarding MIAT rs1894720, a study investigating ischemic stroke (IS) susceptibility in a Chinese population reported that individuals carrying the GG genotype had a significantly increased susceptibility to the large-artery atherosclerosis (LAA) subtype compared with healthy controls. Moreover, carriers of the rs1894720 GG genotype exhibited higher serum MIAT levels than those with the GT or TT genotypes [25].
MIAT SNPs genotyping
Assessment of allelic discrimination for the MIAT rs4274 (assay IDs: C__11476158_20), rs1061540 (assay IDs: C___2467719_1_), and rs1894720 (assay IDs: C__11476152_10) SNP was performed with an ABI StepOne Software v2.3 Real-Time PCR System. The TaqMan assay was applied for the analysis of genotyping [26]. The analysis and calculation of the collected data of genotyping was processed with the SDS 7000 series software (Applied Biosystems, Foster City, CA, USA).
Published databases for validation
Multiple publicly available databases were employed to validate our findings. The Genotype-Tissue Expression (GTEx) portal provides comprehensive data on human gene expression, regulation, and genetic variation, which we used to assess MIAT expression and the SNP rs4274 [27]. The University of Alabama at Birmingham CANcer (UALCAN) database offers an interactive platform for analyzing cancer OMICS data from TCGA, allowing us to examine MIAT expression in relation to clinicopathological features of head and neck squamous cell carcinoma (HNSC) [28].
Statistical analysis
To compare the age, betel quid chewing, cigarette smoking, alcohol drinking, tumor stage, tumor T status, lymph node status, metastasis, and cell differentiation between the oral tongue cancer patients and the controls, the student's t test or Chi-squared test was applied to evaluated the data in these two groups. A p < 0.05 was considered to represent statistical significant. To analyze the odds ratio (OR) with their 95% confidence intervals (CIs) of the association between the genotypic frequencies of MIAT and the clinical pathological statuses in oral tongue cancer patients, the multiple logistic regression models was introduced for the analysis of the data. The SAS statistical software (Version 9.1, 2005; SAS Institute, Cary, NC) was used to evaluate all the data analysis in our study.
Results
The distribution of demographic characteristics among 1,194 controls and 397 male patients with tongue cancer is presented in Table 1. In this study, the proportion of participants aged <55 years was 564 (47.2%) in controls and 192 (48.4%) in male tongue cancer patients, whereas the proportion aged ≥55 years was 630 (52.8%) in controls and 205 (51.6%) in patients. The distributions of environmental risk factor exposures differed significantly between controls and male tongue cancer patients: 198 (16.6%) vs. 264 (66.5%) for betel quid chewing (p < 0.001), 634 (53.1%) vs. 307 (77.3%) for cigarette smoking (p < 0.001), and 237 (19.9%) vs. 149 (37.5%) for alcohol consumption (p < 0.001).
The distributions of demographical characteristics in 1194 controls and 397 male patients with tongue cancer.
| Variable | Controls (N=1194) | Patients (N=397) | p value |
|---|---|---|---|
| Age (yrs) | |||
| < 55 | 564 (47.2%) | 192 (48.4%) | p = 0.743 |
| ≥ 55 | 630 (52.8%) | 205 (51.6%) | |
| Betel quid chewing | |||
| No | 996 (83.4%) | 133 (33.5%) | |
| Yes | 198 (16.6%) | 264 (66.5%) | p < 0.001* |
| Cigarette smoking | |||
| No | 560 (46.9%) | 90 (22.7%) | |
| Yes | 634 (53.1%) | 307 (77.3%) | p < 0.001* |
| Alcohol drinking | |||
| No | 957 (80.1%) | 248 (62.5%) | |
| Yes | 237 (19.9%) | 149 (37.5%) | p < 0.001* |
| Stage | |||
| I+II | 179 (45.1%) | ||
| III+IV | 218 (54.9%) | ||
| Tumor T status | |||
| T1+T2 | 201 (50.6%) | ||
| T3+T4 | 196 (49.4%) | ||
| Lymph node status | |||
| N0 | 244 (61.5%) | ||
| N1+N2+N3 | 153 (38.5%) | ||
| Metastasis | |||
| M0 | 395 (99.5%) | ||
| M1 | 2 (0.5%) | ||
| Cell differentiation | |||
| Well differentiated | 48 (12.1%) | ||
| Moderately or poorly differentiated | 349 (87.9%) |
* p value < 0.05 as statistically significant.
The adjusted odds ratios (AORs) and 95% confidence intervals (CIs) for tongue cancer associated with MIAT genotype frequencies are shown in Table 2. Among patients with tongue cancer, the most frequent MIAT genetic polymorphisms were rs4274 G variant, rs1061540 T variant, and rs1894720 G variant. After adjusted for betel quid chewing, cigarette smoking, and alcohol consumption. A significant association was observed between tongue cancer and the MIAT rs4274 “AA” genotype (p = 0.018, Table 2).
The adjusted odds ratio and 95% confidence interval (CI) of tongue cancer associated with MIAT genotypic frequencies.
| Variable | Controls (N=1194) (%) | Patients (N=397) (%) | AOR (95% CI) | p value |
|---|---|---|---|---|
| rs4274 | ||||
| GG | 446 (37.3%) | 162 (40.8%) | 1.000 (reference) | |
| GA | 555 (46.5%) | 187 (47.1%) | 0.997 (0.754-1.319) | p=0.985 |
| AA | 193 (16.2%) | 48 (12.1%) | 0.609 (0.404-0.917) | p=0.018* |
| GA+AA | 748 (62.7%) | 235 (59.2%) | 0.887 (0.681-1.155) | p=0.373 |
| rs1061540 | ||||
| TT | 397 (33.2%) | 119 (30.0%) | 1.000 (reference) | |
| TC | 574 (48.1%) | 200 (50.4%) | 1.215 (0.904-1.633) | p=0.197 |
| CC | 223 (18.7%) | 78 (19.6%) | 1.179 (0.811-1.715) | p=0.388 |
| TC+CC | 797 (66.8%) | 278 (70.0%) | 1.205 (0.911-1.593) | p=0.191 |
| rs1894720 | ||||
| GG | 514 (43.0%) | 155 (39.0%) | 1.000 (reference) | |
| GT | 525 (44.0%) | 191 (48.1%) | 1.093 (0.828-1.422) | p=0.532 |
| TT | 155 (13.0%) | 51 (12.9%) | 0.984 (0.651-1.487) | p=0.937 |
| GT+TT | 680 (57.0%) | 242 (61.0%) | 1.068 (0.820-1.389) | p=0.626 |
The adjusted odds ratio (AOR) with their 95% confidence intervals were estimated by multiple logistic regression models after controlling for betel quid chewing, cigarette smoking, and alcohol drinking.
We further analyzed the ORs and 95% CIs of clinical characteristics according to MIAT rs4274 genotypes in male tongue cancer patients. A significant association was identified between MIAT rs4274 polymorphisms and tumor differentiation grade [OR (95% CI) = 2.246 (1.217-4.147); p = 0.001] (Table 3). Additionally, when stratifying male tongue cancer patients by betel quid chewing status, betel quid chewers carrying the MIAT rs4274 “GA+AA” variants had a higher risk of moderate or poor tumor differentiation [(OR) 2.968; 95% CI: 1.374-6.410; p = 0.006] (Table 4). Among smokers, those with the “GA+AA” genotype similarly exhibited an increased risk of moderate or poor differentiation [(OR) 2.867; 95% CI: 1.405-5.849; p = 0.004], whereas non-smokers carrying the same genotype showed a significantly higher risk of developing advanced clinical stage (stage III+IV) [(OR) 2.704; 95% CI: 1.087-6.726; p = 0.033] (Table 5).
Odds ratio (OR) and 95% confidence intervals (CI) of clinical statuses associated with genotypic frequencies of MIAT rs4274 in 397 male tongue cancer patients.
| Variable | GG (N=162) | GA+AA (N=235) | OR (95% CI) | p value |
|---|---|---|---|---|
| Clinical Stage | ||||
| Stage I+II | 78 (48.1%) | 101 (43.0%) | 1.000 (reference) | p=0.309 |
| Stage III+IV | 84 (51.9%) | 134 (57.0%) | 1.232 (0.824-1.842) | |
| Tumor size | ||||
| ≤ T2 | 78 (48.1%) | 123 (52.3%) | 1.000 (reference) | p=0.418 |
| > T2 | 84 (51.9%) | 112 (47.7%) | 0.846 (0.566-1.262) | |
| Lymph node metastasis | ||||
| No | 104 (64.2%) | 140 (59.6%) | 1.000 (reference) | p=0.353 |
| Yes | 58 (35.8%) | 95 (40.4%) | 1.217 (0.805-1.840) | |
| Metastasis | ||||
| M0 | 161 (99.4%) | 234 (99.6%) | 1.000 (reference) | p=0.792 |
| M1 | 1 (0.6%) | 1 (0.4%) | 0.688 (0.043-11.080) | |
| Cell differentiated grade | ||||
| Well | 28 (17.3%) | 20 (8.5%) | 1.000 (reference) | p=0.001* |
| Moderate or poor | 134 (82.7%) | 215 (91.5%) | 2.246 (1.217-4.147) |
The odds ratio (OR) with their 95% confidence intervals were estimated by logistic regression models.
* p value < 0.05 as statistically significant.
Clinical statuses and genotypic frequencies of MIAT rs4274 in 397 male tongue cancer patients who are betel quid chewers or not betel quid chewers.
| Non-Betel Quid Chewers (N=133) | Betel Quid Chewers (N=264) | |||||||
|---|---|---|---|---|---|---|---|---|
| Variable | GG (N=47) | GA+AA (N=86) | OR (95% CI) | p value | GG (N=115) | GA+AA (N=149) | OR (95% CI) | p value |
| Clinical Stage | ||||||||
| Stage I+II | 23 (48.9%) | 36 (41.9%) | 1.000 (reference) | 0.433 | 55 (47.8%) | 65 (43.6%) | 1.000 (reference) | 0.497 |
| Stage III+IV | 24 (51.1%) | 50 (58.1%) | 1.331 (0.651-2.720) | 60 (52.2%) | 84 (56.4%) | 1.185 (0.727-1.931) | ||
| Tumor size | ||||||||
| ≦ T2 | 23 (48.9%) | 46 (53.5%) | 1.000 (reference) | 0.616 | 55 (47.8%) | 77 (51.7%) | 1.000 (reference) | 0.535 |
| > T2 | 24 (51.1%) | 40 (46.5%) | 0.833 (0.409-1.698) | 60 (52.2%) | 72 (48.3%) | 0.857 (0.527-1.395) | ||
| Lymph node metastasis | ||||||||
| No | 27 (57.4%) | 49 (57.0%) | 1.000 (reference) | 0.958 | 77 (67.0%) | 91 (61.1%) | 1.000 (reference) | 0.325 |
| Yes | 20 (42.6%) | 37 (43.0%) | 1.019 (0.497-2.092) | 38 (33.0%) | 58 (38.9%) | 1.291 (0.776-2.149) | ||
| Cell differentiation | ||||||||
| Well | 6 (12.8%) | 9 (10.5%) | 1.000 (reference) | 0.689 | 22 (19.1%) | 11 (7.4%) | 1.000 (reference) | 0.006* |
| Moderate or poor | 41 (87.2%) | 77 (89.5%) | 1.252 (0.417-3.762) | 93 (80.9%) | 138 (92.6%) | 2.968 (1.374-6.410) | ||
The odds ratio (OR) with their 95% confidence intervals were estimated by logistic regression models.
* p value < 0.05 as statistically significant.
Clinical statuses and genotypic frequencies of MIAT rs4274 in 397 male tongue cancer patients who are smoker and non-smokers.
| Non-smoker (N=90) | Smoker (N=307) | |||||||
|---|---|---|---|---|---|---|---|---|
| Variable | GG (N=29) | GA+AA (N=61) | OR (95% CI) | p value | GG (N=133) | GA+AA (N=174) | OR (95% CI) | p value |
| Clinical Stage | ||||||||
| Stage I+II | 18 (62.1%) | 23 (37.7%) | 1.000 (reference) | 0.033* | 60 (45.1%) | 78 (44.8%) | 1.000 (reference) | 0.960 |
| Stage III+IV | 11 (37.9%) | 38 (62.3%) | 2.704 (1.087-6.726) | 73 (54.9%) | 96 (55.2%) | 1.012 (0.643-1.592) | ||
| Tumor size | ||||||||
| ≦ T2 | 14 (48.3%) | 31 (50.8%) | 1.000 (reference) | 0.822 | 64 (48.1%) | 92 (52.9%) | 1.000 (reference) | 0.409 |
| > T2 | 15 (51.7%) | 30 (49.2%) | 0.903 (0.373-2.188) | 69 (51.9%) | 82 (47.1%) | 0.827 (0.526-1.299) | ||
| Lymph node metastasis | ||||||||
| No | 20 (69.0%) | 31 (50.8%) | 1.000 (reference) | 0.108 | 84 (63.2%) | 109 (62.6%) | 1.000 (reference) | 0.926 |
| Yes | 9 (31.0%) | 30 (49.2%) | 2.151 (0.846-5.468) | 49 (36.8%) | 65 (37.4%) | 1.022 (0.641-1.631) | ||
| Cell differentiation | ||||||||
| Well | 3 (10.3%) | 7 (11.5%) | 1.000 (reference) | 0.873 | 25 (18.8%) | 13 (7.5%) | 1.000 (reference) | 0.004* |
| Moderate or poor | 26 (89.7%) | 54 (88.5%) | 0.890 (0.213-3.724) | 108 (81.2%) | 161 (92.5%) | 2.867 (1.405-5.849) | ||
The odds ratio (OR) with their 95% confidence intervals were estimated by logistic regression models.
* p value < 0.05 as statistically significant.
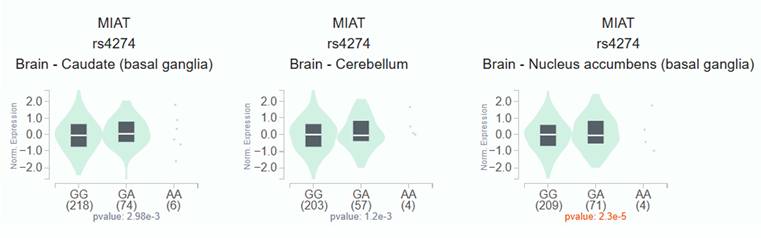
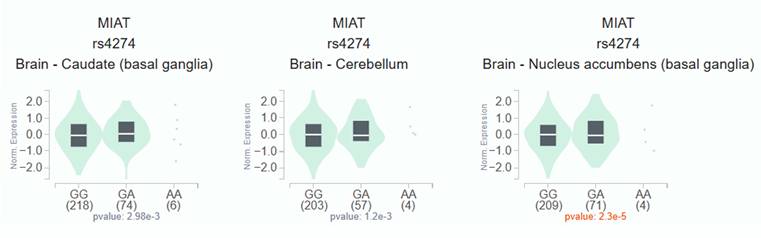
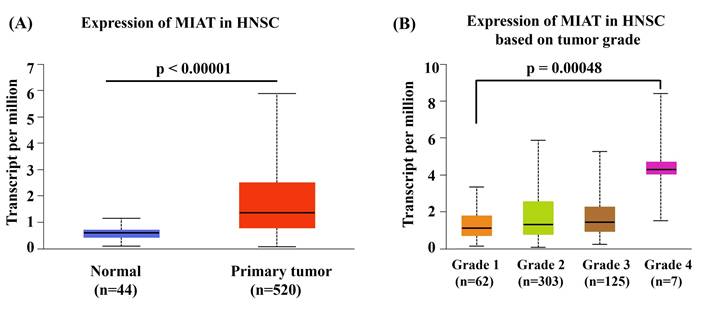
To investigate the functional relevance of MIAT rs4274, we analyzed publicly available bioinformatic databases. GTEx analysis demonstrated that carriers of the rs4274 A allele exhibited significantly higher MIAT expression across various tissues compared to GG homozygotes (Figure 1). Analysis of TCGA data via the UALCAN platform revealed that MIAT expression was significantly elevated in HNSC tumor tissues relative to normal tissues (Figure 2A). Furthermore, higher MIAT mRNA levels were associated with increased tumor grade, with grade 4 tumors showing significantly greater expression compared to grade 1 tumors (p = 0.00048) (Figure 2B).
Association of MIAT expression with MIAT rs4274 polymorphisms in GTEx dataset. MIAT mRNA expression levels in brain tissue according to rs4274 genotypes, showing higher expression in carriers of the A allele.
MIAT expression patterns in head and neck squamous cell carcinoma. (A) UALCAN analysis showing elevated MIAT expression in head and neck squamous cell carcinoma (HNSC) tissues compared with normal tissues. (B) Boxplot from the UALCAN database showing MIAT mRNA expression levels in HNSC tissues across different tumor grades.
Discussion
In this study, we investigated the associations between MIAT SNPs and male tongue cancer. Alcohol consumption, betel quid chewing, and tobacco smoking are well-established environmental risk factors contributing to tongue cancer initiation, development, and progression [29, 30]. Oral tongue squamous cell carcinoma is considered the most common malignancy of the oral cavity [3, 31, 32]. In our current study, statistically significant associations of these environmental risk factors including alcohol drinking, betel quid chewing, and cigarette smoking were found in 397 male patients with tongue cancer compared with 1194 controls, respectively. Although evidence linking alcohol consumption and betel quid chewing to MIAT expression is limited, a study on chronic obstructive pulmonary disease (COPD) reported that MIAT is upregulated in the lung tissues of cigarette smoke (CS)-exposed mice [33]. Furthermore, MIAT knockdown attenuated CS- or CS-extract-induced inflammatory responses, collagen deposition, and EMT [33], suggesting a potential role of MIAT in mediating the effects of cigarette smoking and its contribution to disease progression.
We further examined the association for tongue cancer in relation to MIAT genotypic frequencies. Although previous studies have suggested a potential association between the lncRNA MIAT rs1061540 and ovarian cancer [14], and reported that MIAT rs1894720 variants are associated with various malignancies [16, 17, 19, 34]. However, no significant associations were observed between these two polymorphisms and male tongue cancer in our study. This finding suggests a limited contribution of MIAT rs1061540 and rs1894720 polymorphisms to disease susceptibility and carcinogenesis in male tongue cancer. Additionally, previous research indicated that the rs4274 “AA” genotype may confer an increased risk of paranoid schizophrenia in the Chinese Han population [21]. Consistent with this, our study found a statistically significant association between the MIAT rs4274 “AA” genotype and male tongue cancer, suggesting a potentially more prominent role for MIAT rs4274 polymorphisms in the pathogenesis and susceptibility of male tongue cancer.
We further analyzed the associations of MIAT rs4274 genotypic frequencies in 397 male tongue cancer patients. The MIAT rs4274 “GA+AA” genotype was significantly associated with moderate or poor tumor cell differentiation. Interestingly, when clinical status was analyzed in combination with environmental risk factors, a significant association was observed between moderate or poor differentiation and betel quid chewing. Betel quid chewing is recognized as a major carcinogenic risk factor for oral tongue cancer [30, 35], and MIAT has been suggested as an independent predictor of poor prognosis in OTSCC [10].
However, to date, no study has investigated the interaction between MIAT SNP and betel quid chewing in tongue cancer. In the present study, we observed that male tongue cancer patients who were betel quid chewers and carried the MIAT rs4274 “GA+AA” polymorphic variants had a significantly higher risk of moderate or poor tumor cell differentiation. Given that MIAT has been reported to promote oral cancer cell invasion by regulating EMT marker expression through Wnt/β-catenin pathway activation [10, 36], our findings suggest a potential synergistic effect of MIAT rs4274 “GA+AA” variants and betel quid chewing on tongue cancer susceptibility and progression. Nevertheless, the precise mechanisms underlying this interaction remain unclear.
Considering that the MIAT rs4274 “GA+AA” carriers exhibit moderate or poor cell differentiation, it is plausible that the “A” allele may upregulate MIAT expression independently of environmental risk factors. Previous studies have reported elevated MIAT levels in both oral cancer tissue and serum, implicating MIAT in tumor growth, migration, invasion, and EMT [10]. Moreover, bioinformatic analyses also indicated that the MIAT rs4274 A allele is associated with increased MIAT expression, and higher MIAT levels correlated with higher tumor grade. Therefore, MIAT rs4274 “A” allele may contribute directly to oral tongue cancer progression.
For smokers carrying the MIAT rs4274 “A” allele, we observed a higher risk of moderate or poor differentiation. Supporting this, prior research has shown that MIAT mediates cigarette smoke-induced EMT and airway remodeling via the miR-29c-3p-HIF3A axis in COPD [33]. Although these patients did not show a significantly higher risk of advanced clinical stage in our cohort, it is reasonable to propose that elevated MIAT expression may promote more aggressive tumor progression, potentially contributing to poor prognosis and increased cancer mortality. A limitation of our study is the absence of serum MIAT level data from the enrolled patients, which precludes a more comprehensive analysis. Future well-designed studies are warranted to elucidate the relationship between MIAT rs4274 variants and MIAT expression in oral tongue cancer, which may help identify MIAT as a potential biomarker for tongue cancer susceptibility and disease progression.
In conclusion, our study demonstrates associations between MIAT rs4274 polymorphisms and tongue cancer susceptibility, as well as clinical outcomes in relation to environmental risk factors. To our knowledge, this is the first study to report an interaction between betel quid chewing and the MIAT rs4274 “A” allele in promoting male tongue cancer progression. Specifically, betel quid chewers carrying the rs4274 “A” allele tended to develop moderate or poor cell differentiation, and smokers with the same genotype were at higher risk of poor differentiation. Collectively, MIAT rs4274 polymorphisms may serve as a pivotal biomarker for predicting MIAT-mediated tongue cancer susceptibility and progression in males.
Acknowledgements
We thank the Human Biobank of Chung Shan Medical University Hospital, Taichung, Taiwan for specimen preparation. This study was supported by Chung Shan Medical University and Changhua Christian Hospital (CSMU-CCH-114-07). This study was also supported by research grants from the Chung Shan Medical University, Taiwan (CSMU-INT-111-05).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Feller G, Mmereki D, Mahomed F, Khammissa RA, Ramiah D. Demographic, clinicopathological, and treatment of oral squamous cell carcinoma patients at a Johannesburg Academic Hospital, South Africa: a 5-year retrospective observational study. BMC Cancer. 2025;25:1917
2. Lin CW, Yang WE, Su CW, Lu HJ, Su SC, Yang SF. IGF2BP2 promotes cell invasion and epithelial-mesenchymal transition through Src-mediated upregulation of EREG in oral cancer. Int J Biol Sci. 2024;20:818-30
3. Marini A, Zakopoulou R, Ntritsos G, Iliou K, Giannousis C, Varvarousis D. et al. The prognostic value of lymphatic vessel density in squamous cell carcinoma of the tongue: a systematic review and meta-analysis. Eur Arch Otorhinolaryngol. 2025
4. Lu HJ, Chiu YW, Peng CY, Tseng HC, Hsin CH, Chuang CY. et al. Parameters to assess the necessity of adjuvant therapy for early-stage oral squamous cell carcinoma. Oral Dis. 2025;31:756-68
5. Su SC, Lin CW, Chen MK, Lee YC, Su CW, Bai S. et al. Multimodal profiling of oral squamous cell carcinoma identifies genomic alterations and expression programs associated with betel quid chewing. Neoplasia. 2025;68:101218
6. Su SC, Chang LC, Huang HD, Peng CY, Chuang CY, Chen YT. et al. Oral microbial dysbiosis and its performance in predicting oral cancer. Carcinogenesis. 2021;42:127-35
7. Hsieh MJ, Lin CW, Su SC, Reiter RJ, Chen AW, Chen MK. et al. Effects of miR-34b/miR-892a Upregulation and Inhibition of ABCB1/ABCB4 on Melatonin-Induced Apoptosis in VCR-Resistant Oral Cancer Cells. Mol Ther Nucleic Acids. 2020;19:877-89
8. Yeh JC, Chen YT, Chou YE, Su SC, Chang LC, Chen YL. et al. Interactive effects of CDKN2B-AS1 gene polymorphism and habitual risk factors on oral cancer. J Cell Mol Med. 2023;27:3395-403
9. Ishii N, Ozaki K, Sato H, Mizuno H, Susumu S, Takahashi A. et al. Identification of a novel non-coding RNA, MIAT, that confers risk of myocardial infarction. J Hum Genet. 2006;51:1087-99
10. Zhong W, Xu Z, Wen S, Xie T, Wang F, Wang Q. et al. Long non-coding RNA myocardial infarction associated transcript promotes epithelial-mesenchymal transition and is an independent risk factor for poor prognosis of tongue squamous cell carcinoma. J Oral Pathol Med. 2019;48:720-7
11. Qiu L, Tao A, Liu F, Ge X, Li C. Potential prognostic value of a eight ferroptosis-related lncRNAs model and the correlative immune activity in oral squamous cell carcinoma. BMC Genom Data. 2022;23:80
12. Khayamzadeh M, Ghaderian SMH, Garajei A, Jolani MS, Tavassoli A, Khodaee P. Evaluation of Salivary, Plasma, and Tissue ITGB8 and MIAT-lncRNA Expression as a Biomarker in Oral Squamous Cell Carcinoma: A Cross-Sectional Study. Biochem Genet. 2025
13. Weng YH, Chen J, Yu WT, Luo YP, Liu C, Yang J. et al. lncRNA-MIAT rs9625066 polymorphism could be a potential biomarker for ischemic stroke. BMC Med Genomics. 2024;17:58
14. Fawzy MS, Ibrahiem AT, Osman DM, Almars AI, Alshammari MS, Almazyad LT. et al. Angio-Long Noncoding RNA MALAT1 (rs3200401) and MIAT (rs1061540) Gene Variants in Ovarian Cancer. Epigenomes. 2024 8
15. Motawi TK, Sadik NAH, Shaker OG, Ghaleb MMH, Elbaz EM. Expression, Functional Polymorphism, and Diagnostic Values of MIAT rs2331291 and H19 rs217727 Long Non-Coding RNAs in Cerebral Ischemic Stroke Egyptian Patients. Int J Mol Sci. 2024 25
16. Asadabadi T, Mokhtari MJ, Bayat M, Safari A, Borhani-Haghighi A. Association between Genetic Polymorphism of The lncRNA MIAT rs1894720 with Ischemic Stroke Risk and lncRNA MIAT Expression Levels in The Blood after An Ischemic Stroke: A Case-Control Study. Cell J. 2023;25:863-73
17. Li Y, Zhang W, Ke H, Wang Y, Duan C, Zhu Q. et al. Rs1894720 polymorphism is associated with the risk of age-related cataract by regulating the proliferation of epithelial cells in the lens via the signalling pathway of MIAT/miR-26b/BCL2L2. Arch Med Sci. 2022;18:223-36
18. Ma R, He X, Zhu X, Pang S, Yan B. Promoter polymorphisms in the lncRNA-MIAT gene associated with acute myocardial infarction in Chinese Han population: a case-control study. Biosci Rep. 2020 40
19. Hao S, Wang L, Zhao K, Zhu X, Ye F. Rs1894720 polymorphism in MIAT increased susceptibility to age-related hearing loss by modulating the activation of miR-29b/SIRT1/PGC-1alpha signaling. J Cell Biochem. 2019;120:4975-86
20. Zheng C, Li X, Qian B, Feng N, Gao S, Zhao Y. et al. The lncRNA myocardial infarction associated transcript-centric competing endogenous RNA network in non-small-cell lung cancer. Cancer Manag Res. 2018;10:1155-62
21. Rao SQ, Hu HL, Ye N, Shen Y, Xu Q. Genetic variants in long non-coding RNA MIAT contribute to risk of paranoid schizophrenia in a Chinese Han population. Schizophr Res. 2015;166:125-30
22. Hsu HT, Lu YT, Chen YT, Hsing MT, Su CW, Su SC. et al. Impact of LINC00312 gene polymorphism coupled with habitual risks on buccal mucosa cancer. J Cancer. 2024;15:2354-60
23. International HapMap C. The International HapMap Project. Nature. 2003;426:789-96
24. Elwazir MY, Hussein MH, Toraih EA, Al Ageeli E, Esmaeel SE, Fawzy MS. et al. Association of Angio-LncRNAs MIAT rs1061540/MALAT1 rs3200401 Molecular Variants with Gensini Score in Coronary Artery Disease Patients Undergoing Angiography. Biomolecules. 2022 12
25. Guo F, Wang N, Yu C, Fan Y, Chen D. Genetic association between long non-coding RNA MIAT polymorphism and ischemic stroke susceptibility in the Chinese population. J Stroke Cerebrovasc Dis. 2024;33:107813
26. Hung LC, Huang CC, Lu YT, Su CW, Lin CW, Chu HJ. et al. NSUN2 rs13181449 variant decreases the risk of oral cancer development. Int J Med Sci. 2025;22:3022-31
27. Carithers LJ, Moore HM. The Genotype-Tissue Expression (GTEx) Project. Biopreserv Biobank. 2015;13:307-8
28. Chandrashekar DS, Karthikeyan SK, Korla PK, Patel H, Shovon AR, Athar M. et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18-27
29. Ribeiro V, Teillaud JL, Dieu-Nosjean MC, Lescaille G, Rochefort J. The prognostic significance of tertiary lymphoid structures in oral squamous cell carcinomas: a systematic review. Front Oral Health. 2024;5:1524313
30. Lenoci D, Moresco E, Cavalieri S, Bergamini C, Torchia E, Botta L. et al. Oral cancer in young adults: incidence, risk factors, prognosis, and molecular biomarkers. Front Oncol. 2024;14:1452909
31. Iaria R, Giovannacci I, Meleti M, Leao JC, Vescovi P. Management of a carcinoma in situ of the tongue margin, using a Nd:YAG laser, cross-linked hyaluronic acid (xHyA) gel and a porcine pericardium resorbable membrane: a case report. BMC Oral Health. 2025;25:378
32. Chang S, Li L, Liu Y, Huang G, Pan ST. Advances in tumor micro-environment and immunotherapy of squamous cell carcinoma of the tongue. Biochim Biophys Acta Rev Cancer. 2025;1880:189483
33. Gu W, Wang L, Deng G, Gu X, Tang Z, Li S. et al. Knockdown of long noncoding RNA MIAT attenuates cigarette smoke-induced airway remodeling by downregulating miR-29c-3p-HIF3A axis. Toxicol Lett. 2022;357:11-9
34. Guo F, Wang N, Yu C, Fan Y, Chen D. Corrigendum to Genetic association between long non-coding RNA MIAT polymorphism and ischemic stroke susceptibility in the Chinese population. J Stroke Cerebrovasc Dis. 2024;33:108009
35. Ho PS, Ko YC, Yang YH, Shieh TY, Tsai CC. The incidence of oropharyngeal cancer in Taiwan: an endemic betel quid chewing area. J Oral Pathol Med. 2002;31:213-9
36. Ghafouri-Fard S, Azimi T, Taheri M. Myocardial Infarction Associated Transcript (MIAT): Review of its impact in the tumorigenesis. Biomed Pharmacother. 2021;133:111040
Author contact
Corresponding authors: Ying-Erh Chou, PhD. or Shun-Fa Yang, PhD. School of Medicine, Chung Shan Medical University, 110, Section 1, Chien-Kuo N. Road, Taichung, Taiwan, ROC. Fax: 886-4-24723229. E-mail: intointo814com (Ying-Erh Chou); E-mail: ysfedu.tw (Shun-Fa Yang).

 Global reach, higher impact
Global reach, higher impact