Impact Factor ISSN: 1449-1907
Int J Med Sci 2018; 15(6):617-627. doi:10.7150/ijms.23924 This issue Cite
Research Paper
Anti-biofilm and Anti-Virulence Efficacy of Celastrol Against Stenotrophomonas maltophilia
1. Department of Biomedical Laboratory Science, College of Medical Sciences, Soonchunhyang University, Asan, Chungnam 31538, Republic of Korea
2. Department of Clinical Laboratory Science, Hyejeon College, Hongseoung, Chungnam 32244, Republic of Korea
Received 2017-11-19; Accepted 2018-3-2; Published 2018-3-28
Abstract
Stenotrophomonas maltophilia is a multi-drug resistant opportunistic pathogen that causes nosocomial infections in immunocompromised patients. This pathogen is difficult to treat owing to its intrinsic multidrug resistance and ability to form antimicrobial-tolerant biofilms. In the present study, we aimed to assess the potential use of celastrol as a novel anti-biofilm and/or anti-virulence agent against S. maltophilia. Results showed that celastrol at its sub-inhibitory doses decreased biofilm formation and disrupt the established biofilms produced by S. maltophilia. Celastrol-induced decrease in biofilm formation was dose-dependent based on the results of the microtiter plate biofilm assays and confocal laser scanning microscopy. In addition, our data validated the anti-virulence efficacy of celastrol, wherein it significantly interfered with the production of protease and motility of S. maltophilia. To support these phenotypic results, transcriptional analysis revealed that celastrol down-regulated the expression of biofilm- and virulence- associated genes (smeYZ, fsnR, and bfmAK) in S. maltophilia. Interestingly, celastrol significantly inhibited the expression of smeYZ gene, which encodes the resistance-nodulation-division (RND)-type efflux pump, SmeYZ. Overall, our findings suggested that celastrol might be a promising bioactive agent for treatment of biofilm- and virulence-related infections caused by the multi-drug resistant S. maltophilia.
Keywords: Celastrol, Stenotrophomonas maltophilia, Anti-biofilm, Anti-virulence, Resistance-nodulation-division (RND)-type efflux pump
Introduction
Stenotrophomonas maltophilia, a microorganism ubiquitously present in the environment, is an emerging multidrug resistant opportunistic human pathogen that often causes nosocomial infections. S. maltophilia is generally associated with septicemia and various infections, particularly respiratory infections, in immunosuppressed or immunocompromised patients, including those with cystic fibrosis (CF) [1, 2]. This aerobic gram-negative bacterium can also cause severe diseases, such as pneumonia, bacteremia, and endocarditis in patients exposed to invasive clinical devices and/or those receiving broad-spectrum antibiotics [3, 4]. One difficulty in treating S. maltophilia infections has been attributed to its intrinsic multidrug resistance [5]. Most clinical isolates of S. maltophilia exhibit multidrug resistance to β-lactams, macrolides, and aminoglycosides owing to its multidrug efflux pumps and overexpression of resistant determinants, such as L1/L2 β-lactamases and aminoglycoside-modifying enzymes [3]. In addition, S. maltophilia genome encodes several molecular machineries that contribute to its resistance to various antibiotics [6]. Therefore, S. maltophilia has become a substantial threat to human health.
Resistance-nodulation-division (RND)-type efflux pumps can confer resistance against various drugs. They significantly contribute to the multidrug resistance of S. maltophilia [5]. Besides their known roles in antibiotic extrusion, RND-type efflux pumps are increasingly recognized as components required for the bacterial physiological functions and virulence [7]. Of the diverse putative RND-type efflux systems described in S. maltophilia, SmeYZ efflux pump is constitutively expressed. It was shown that inactivation of smeYZ resulted in attenuation of drug resistance and virulence-associated characteristics, which decreased the in vivo virulence of S. maltophilia [8].
A broad spectrum of clinical syndromes has been shown to be caused by S. maltophilia infections [9]. However, little is known about its virulence factors. One of the most crucial virulence factors of S. maltophilia is its formidable ability to form biofilms, which are sessile communities of microbial cells that can adhere to one another on biotic and abiotic surfaces [1]. These adherent cells are often encased in a self-produced matrix of an extracellular polymeric substance, forming a complex three-dimensional architecture [10-12]. Biofilm-grown cells exhibit phenotypic properties that differ from those of the organisms grown planktonically, including increased resistance to multiple antibiotics, host immune responses, and multiple stressors [13-15]. One of the mechanisms that afford resistance against numerous antibiotics involve the delayed penetration of the antimicrobial drugs through the multilayer biofilm matrix. Therefore, treatment of biofilm-related infections is clinically difficult, resulting in chronic infections [16, 17].
Biofilm formation is a complex and multifactorial event. It depends on the surface characteristics, strain motility, genetic factors, and other factors [10, 12, 16]. Biofilm formation correlates with high resistance to antibiotics [18]. Recently, screening of a random transposon insertion mutant library of S. maltophilia ATCC 13637 identified the fsnR gene that encodes an orphan response regulator, FsnR. This regulatory protein plays a fundamental role in mediating the transcription of flagellar genes, cell motility, and biofilm formation [19]. Besides, two histidine kinase (HK) genes were shown to encode a two-component signal transduction system (TCS), the BfmAK system. This system has been confirmed to play a critical role in biofilm formation in S. maltophilia [20].
Conventional antibiotics used to treat biofilm-forming bacterial pathogens were not designed to act against the recalcitrant biofilm. Alternatively, they were designed to inhibit the growth of bacteria or kill them. This approach exerts selective pressure upon the pathogenic bacteria, which induces bacterial resistance against the particular drug being used [21]. With the increasing influence of drug resistance, novel therapies or antibiotic agents, including natural chemicals and plant products that can regulate biofilm formation, remove established biofilms, and inhibit the expression of virulence-related factors, are needed to treat multidrug resistant S. maltophilia infections. These agents have received an increasing attention as a new class of antimicrobial compounds with anti-biofilm activity for treatment of infectious diseases [22, 23].
Celastrol is a pentacyclic triterpenoid compound mainly found in the roots of Tripterygium wilfordii, commonly known as thunder god vine. It has been shown that celastrol has multiple pharmacological effects, including anti-inflammatory, anti-cancer, hepatoprotective, and immunomodulatory effects [24, 25]. As a natural product, celastrol might exert inhibitory effects on biofilm formation. Recent in vitro studies showed that celastrol exhibited antimicrobial and anti-biofilm effects against the gram-positive pathogen Staphylococcus aureus [16, 26].
In this context, the present study was conducted with the aim of assessing the antimicrobial, anti-biofilm, and anti-virulence activities of celastrol against S. maltophilia using in vitro phenotypic and genotypic analyses.
Materials and Methods
Bacterial strains, growth measurements, and celastrol
S. maltophilia strain ATCC 13637 was purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). Clinical isolate, GNU2233 (from respiratory tract), were kindly provided by Gyeongsang National University Hospital Culture Collection for Pathogens. For general purpose, all strains were grown aerobically at 37 ºC in cation-adjusted Mueller-Hinton broth (CA-MHB; Difco, Becton-Dickinson and Company, USA). For cell growth measurements, absorbance at 600 nm (A600 nm) were measured using Multiskan GO plate reader (Thermo Fisher Scientific, USA). Each experiment was performed using at least three independent cultures, as described previously [26]. Celastrol was purchased from Cayman Chemicals (Ann Arbor, MI) and dissolved in dimethyl sulfoxide (DMSO).
Crystal violet biofilm assay
To determine the effect of celastrol on biofilm formation and established biofilms, crystal-violet staining experiments were performed [1]. First, S. maltophilia strains (ATCC 13637 and GNU2233) were inoculated into CA-MHB broth and cultured overnight to measure the blocking effect of celastrol on biofilm formation. S. maltophilia ATCC 13637 and GNU2233 were standardized to match approximately 0.5 McFarland standards. They were then diluted at 1:20 (A600 nm, 0.005) with CA-MHB. Then 200 μL of the dilution was inoculated into the wells of sterile 96-well polystyrene tissue culture plate (Falcon) with or without sublethal celastrol concentrations (two-fold dilutions from 1.25 μg/mL to 10 μg/mL for ATCC 13637, and from 5 μg/mL to 40 μg/mL for GNU2233). For each concentration of celastrol, 8 wells were used. Media treated with 1% DMSO was used as positive control. After 24 h of incubation at 37 °C, the culture media was discarded and wells were washed three times with 200 μL of sterilized phosphate buffered saline (PBS) to remove non-adherent cells. Adherent biofilms were fixed by drying at 60 °C for 1 h and stained with Hucker-modified 0.1% crystal violet for 5 min. After discarding the remained stain by tapping on paper towel, stained biofilms were thoroughly rinsed three times with sterile deionized water, dried at 60 °C for 1 h, and extracted with 250 μL of 33% glacial acetic acid for 15 min, as described previously [26]. Dissolved biofilms were quantified by measuring the absorbance values at wavelength of 595 nm using the Multiskan GO plate reader. To examine the effect of celastrol on already established biofilms, 200 μL of bacteria culture diluted in media was inoculated into 96-well tissue culture plate and incubated at 37 °C for 24 h to allow biofilm formation. After the wells were washed three times with PBS, 200 μL of CA-MHB containing sub-inhibitory concentrations of celastrol was added to each well. After further incubation for 24 h, biofilms were stained and quantified as described above. Each experiment was performed at least three times in parallel.
XTT reduction assay
To measure cell viability of S. maltophilia in established biofilms following treatment with celastrol, XTT Cell proliferation kit (ATCC, Manassas, VA, USA) was used according to the manufacturer's recommendations [27]. As described above, biofilms of two S. maltophilia strains (ATCC 13637 and GNU2233) were prepared in 96-well polystyrene tissue culture plate. After 24 h of incubation, biofilms were rinsed three times with PBS and incubated with fresh CA-MHB media containing celastrol (10, 20, 40, and 80 μg/mL for ATCC 13637 or 40, 60, 80, and 100 μg/mL for GNU2233) or 1 % of DMSO for 24 h. Before reacting with cells from biofilms, XTT reagent and activation reagent were mixed at a ratio of 50:1 (v/v). Then 50 μL of activated-XTT solution was added into each well and incubated at 37 °C for 3 h in the dark. After incubation, reduction of XTT solution was determined by absorbance reading at wavelength of 475 nm using the Multiskan GO plate reader. The absorbance reading at wavelength of 600 nm was used as background (to be subtracted from the reading at 475 nm).
Confocal laser scanning microscopy (CLSM)
Confocal laser scanning microscopy was performed to visualize biomass and cells of biofilm affected by celastrol using published procedures [28] with some modifications. Briefly, overnight bacterial culture of S. maltophilia ATCC 13637 was adjusted and inoculated into tissue culture treated 24-well glass bottom imaging plates (Effendorf AG, Hamburg, Germany, Cat. no.: 0030741021). After 24 h of incubation, suspension medium was discarded and each well was washed three times with 1 mL of sterile PBS. Preformed biofilms were then treated with celastrol (40 or 80 µg/mL) or DMSO 1 % (v/v) and incubated at 37 °C for 24 h. After washing out non-adherent cells with PBS, biofilms were fixed with 3.7 % (v/v) formaldehyde for 1 h, washed with sterile PBS, and then stained with three different fluorescent dyes. The amino group was stained with 10 µg/µL of fluorescein isothiocyanate isomer Ⅰ (FITC; Sigma-Aldrich, Germany) for 1 h at room temperature (RT). To visualize carbohydrate, wells were stained with 0.1 µg/µL of Concanavalin A-Alexa Fluor 594 conjugate (ConA;C-11253, Molecular Probes, Eugene, OR, USA) at room temperature (RT) for 20 min. Extracellular nucleic acids were stained with 1 mg/L of 4, 6-diamidino-2-phenylindoldihydrochloride (DAPI; Molecular Probes, Eugene OR, USA) at RT for 40 min. After each staining step, wells were washed with 1 mL of PBS to remove unbound dye and dried for 15 min. All procedures were conducted in the dark. The excitation laser wavelengths for FITC, ConA, and DAPI were 495, 590, and 358 nm, respectively. Microscopic analysis was performed using a Zeiss LSM-710 confocal laser microscope (Carl Zeiss, Thornwood, NY, USA). Images were constructed using ZEN software (Carl Zeiss, Thornwood, NY, USA), as described previously [26].
Swimming motility assay
The effect of celastrol on swimming motility was assessed using 0.15% agar plate containing 1% tryptone and 0.5% NaCl as described previously [29] with slight modifications. Briefly, 1% DMSO (the control) or sublethal doses of celastrol was incorporated into motility agar plates. Two S. maltophilia strains (ATCC 13637 and GNU2233) grown overnight in CA-MHB under shaking conditions were diluted to OD600 of 1.0. Thereafter, 5 µL aliquot of bacterial culture was inoculated into the center of the motility agar surface. The inoculated plates were then incubated at 37 °C for 24 h and the diameter (mm) of circular swimming zone was measured. Each experiment was performed in triplicates with three independent cultures.
Secreted protease activity assay
The effect of celastrol on the activity of protease secreted by two S. maltophilia strains (ATCC 13637 and GNU2233) was determined using Mueller-Hinton broth casein agar containing 1% skim milk as described previously [8] with some modifications. Briefly, each overnight culture grown in CA-MHB under shaking was adjusted to OD600 of 0.005. The diluted bacterial culture was incubated with or without sublethal doses of celastrol at 37 °C for 24 h. Cell cultures were centrifuged at 12,000 x g for 15 min at 4 °C. Clear supernatant was collected from either celastrol treated or untreated sample via filtration using 0.45-µm filters (Corning). Thereafter, 40 µL of cell free culture supernatant (both treated and untreated) was inoculated into 6-mm diameter hole in the center of casein agar plate. After incubation at 37 °C for 48 h, proteolytic activities of cell free supernatants of strains were assessed by measuring the diameters of transparent zones surrounding the holes. Each experiment was performed in triplicates with three independent cultures.
RNA isolation
For transcriptional analysis of S. maltophilia, total bacterial RNA was isolated using the following procedure. Briefly, overnight cultures of S. maltophilia strains (ATCC 13637 and GNU2233) were diluted with 30 mL of CA-MHB medium to OD600 of 0.005. All cultures were then grown at 37 ºC for 6 h with shaking (250 rpm). After that, cell suspensions were cultured for another 4 hours at 37 ºC in the presence or absence of celastrol at final concentration of 5 or 20 μg/mL. The two concentrations were chosen because they showed significant anti-biofilm and anti-virulence activities. Then 2 mL of bacterial culture from each sample was centrifuged at 25,000 x g for 1 min at 4 ºC to collect cells. Bacterial total RNA was extracted and purified from pelleted cells using NucleoSpin RNA mini kit (Macherey-Nagel, Düren, Germany) according to the manufacturer's protocol. To remove residual genomic DNA, extracted total RNA was filtrated with silica columns and subjected to DNase treatment. The quality and concentration of isolated RNA samples were assessed using BioDrop µLITE (BioDrop Ltd., Cambridge, UK), as described previously [26].
qRT-PCR
The transcription levels of virulence- and biofilm- related genes (smeYZ, fsnR, bfmA, and bfmK) in S. maltophilia ATCC 13637 and GNU2233 treated with or without celastrol were determined using qRT-PCR. Gene-specific primers were used for the amplification of these genes. Transfer-messenger RNA (tmRNA) housekeeping gene [19] was used as an internal control. Primers used in this study were designed based on the genome sequence of S. maltophilia K279a [6]. Primer sequences are listed in Table 1. ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO, Japan) was used to generate cDNA using 1 μg of total RNA. qRT-PCRs were performed in sets of triplicates using THUNDERBIRD SYBR qPCR Mix (TOYOBO, Japan) on an Mx3000P instrument (Agilent Technologies). Thermal cycling parameters were set as follows: at 95 ºC for 1 min followed by 40 cycles of at 95 ºC for 15 s and at 63 ºC for smeYZ gene 1 min (at 60 ºC for tmRNA gene for 1 min and at 61 ºC for 1 min for other genes). This was followed by1 cycle each at 95 ºC for 1 min, at 60 ºC for 30 s, and at 95 ºC for 30 s. The expression values were normalized against tmRNA. All measurements were performed with three independent cultures and three replications within each qRT-PCR reaction, as described previously [26].
Nucleotide sequences of primers used for quantitative RT-PCR analysis.
| Genes | Primer sequences |
|---|---|
| tmRNA | Forward: 5'- GGG GGT GCA CTG GTT TCG -3' Reverse: 5'- TGG TGG AGG TGG GCG GAA T -3' |
| smeYZ | Forward: 5'- CAG ATC GAC CAG GTC TAC -3' Reverse: 5'- CAT TGA TAC CGG AGA ACA-3' |
| fsnR | Forward: 5'- CAT CCT TGA CCA CGA ACC -3' Reverse: 5'- TGA TGG ACC TGT CAT TGC -3' |
| bfmA | Forward: 5'- ATG ACG ACT CCC GCC CGT -3' Reverse: 5'- CGA CCT GCA GCA GCA CGT C -3' |
| bfmK | Forward: 5'- GCG GGT CGG TGT ACT GAA GG -3' Reverse: 5'- ATC GGC CAA GCG TCG TAG -3' |
Statistical analysis
Data are expressed as means ± standard deviations (SD). Significant difference between treated and untreated samples was determined using Student's t-test. Statistical significance was considered at p < 0.05.
Results
Celastrol inhibited biofilm formation in S. maltophilia without affecting planktonic cell growth
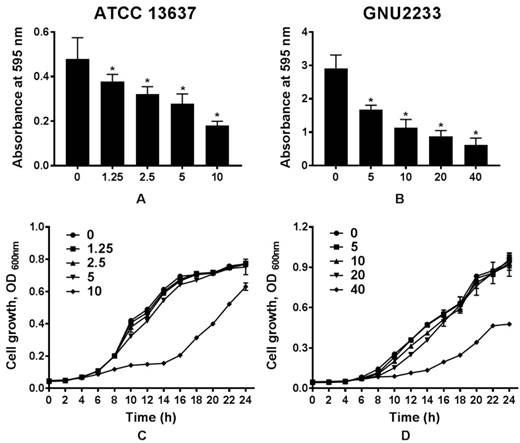
To determine whether celastrol could affect biofilm formation in S. maltophilia strains (ATCC 13637 and GNU2233), the biofilm formation abilities were compared in the presence or absence of celastrol. Flat-bottom 96-well polystyrene microtiter plates were used. Phenotypic analyses were performed using crystal violet based biomass staining assay. As shown in Figure 1, celastrol decreased biofilm formation in the two S. maltophilia strains in a dose-dependent manner, indicating that celastrol exhibited concentration-dependent anti-biofilm effects. The inhibitory effects of celastrol against biofilm formation in ATCC 13637 and GNU2233 were observed at concentrations of 5 and 20 μg/ml (biofilm reduced by 54.76 and 72.51%, respectively, compared to the control; Figure 1A and 1B). Planktonic cell growth of S. maltophilia was also evaluated to examine the anti-biofilm and anti-virulence activities of celastrol in the absence of its bactericidal activity. It was found that celastrol (at the concentrations tested) failed to exert any antimicrobial effects against the planktonic S. maltophilia cells (Figure 1C-1D). In this context, the cell growth of S. maltophilia in the presence of different concentrations of celastrol over different time periods indicated that celastrol exhibited a bacteriostatic effect. Furthermore, the antimicrobial activity of celastrol was evaluated by determining its minimum inhibitory concentration (MIC). The MICs of celastrol against S. maltophilia ATCC 13637 and GNU2233 were 20 and 80 μg/ml, respectively. It is noteworthy that celastrol (at the concentrations tested) exerted significant (p < 0.05) inhibitory effects against biofilm formation in the S. maltophilia without reducing cell growth. These results indicated that the observed attenuation of biofilm formation exerted by celastrol was attributed to its anti-biofilm effects, not to its antimicrobial effects.
Effect of celastrol on biofilm formation of S. maltophilia. Biofilms produced by S. maltophilia strains ((A) ATCC 13637 and (B) GNU2233) were quantified by measuring the absorbance values at OD595nm in the presence or absence of celastrol at increasing doses. Planktonic cell growth of S. maltophilia strains ((C) ATCC 13637 and (D) GNU2233) in the presence of celastrol was determined by measuring absorbance at wavelength of 600 nm. Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to that of DMSO treated control.
Dispersion of established S. maltophilia biofilms by celastrol
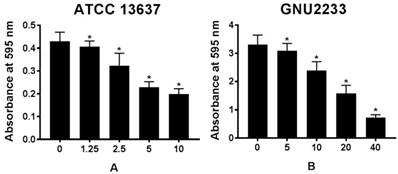
The biofilm dispersion activity of celastrol was investigated in S. maltophilia strains (ATCC 13637 and GNU2233) by measuring the biomass of biofilms. As shown in Figure 2, compared to the untreated controls (DMSO alone), celastrol treatment dose-dependently disassembled the established biofilms of S. maltophilia tested, indicating that celastrol exhibited concentration-dependent biofilm dispersion effects. In a more detailed study, celastrol exerted a significant (p < 0.05) dispersion activity against the established biofilms produced by S. maltophilia at all concentrations tested without inhibiting the growth of planktonic S. maltophilia cells. Therefore, celastrol inhibited and dispersed the biofilms produced by S. maltophilia. In addition, it significantly reduced the biomass of biofilms in a dose-dependent manner.
Effect of celastrol on established biofilms formed by S. maltophilia strains ((A) ATCC 13637 and (B) GNU2233). To determine the ability of celastrol to disrupt mature biofilms, S. maltophilia strains were cultured at 37 °C for 24 h to form established biofilms in microtitre plates. Celastrol was then added at different concentrations. These cultures were incubated further for 24 h. Total biofilm formation of S. maltophilia strains were examined by measuring absorbance at OD595nm. Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to that of DMSO treated control.
Effects of celastrol on the metabolism of established biofilms of S. maltophilia based on the XTT reduction assay
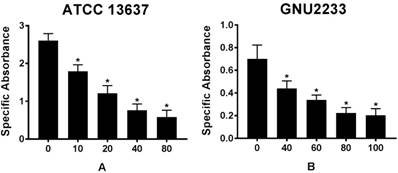
To further emphasize that celastrol could reduce the viability of biofilm cells, a colorimetric XTT reduction analysis was performed after treatment of the established biofilms with celastrol. Results of the XTT reduction assay revealed that the metabolic activity of S. maltophilia strains (ATCC 13637 and GNU2233) decreased with increasing the concentration of celastrol (p < 0.05, Figure 3). After 24-hour exposure of S. maltophilia biofilms to celastrol at concentrations of 80 and 100 μg/ml, the specific absorbance values were significantly (p < 0.05) reduced, compared to that of the DMSO-treated control group (Figure 3). In particular, celastrol at 80 μg/ml significantly decreased the metabolic activity of S. maltophilia ATCC 13637 biofilm cells by more than 80%, compared to the DMSO-treated control (Figure 3A). At a concentration of 100 μg/ml, celastrol diminished the viability of GNU2233 biofilm cells by more than 90% (Figure 3B).
Effect of celastrol on metabolic activities of established biofilms formed by S. maltophilia strains ((A) ATCC 13637 and (B) GNU2233). The viability of cells within the biofilms treated with or without celastrol was evaluated by XTT viability assay. Specific absorbance indicates A490nm (Test) - A490nm (Blank) - A655nm (Test). Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to that of DMSO treated control.
Effects of celastrol on biofilm structure
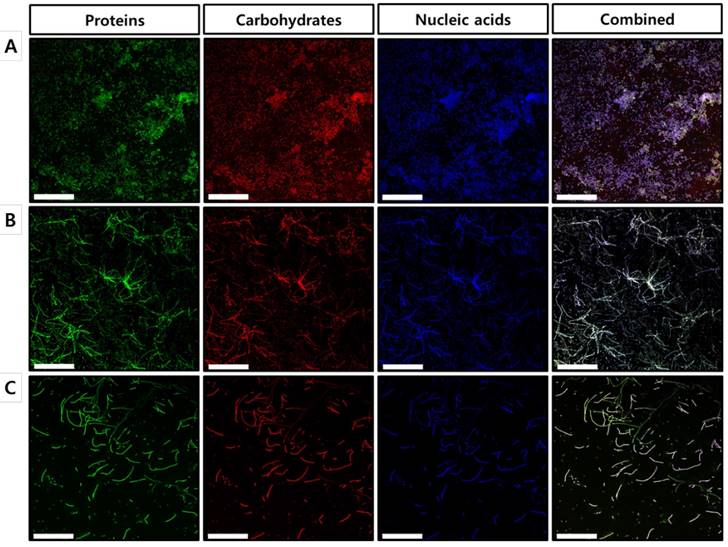
To further confirm our findings regarding the effects of celastrol on the established biofilms formed by S. maltophilia ATCC 13637, biofilm matrix components were individually stained with FITC, concanavalin A, and DAPI, followed by characterization using confocal laser scanning microscopy (CLSM). As shown in the CLSM images, biofilms treated with celastrol at concentrations of 40 and 80 μg/ml exhibited a significantly different structure, compared to that of the DMSO-treated control (Figure 4), which is consistent with the results of the 96-well biofilm assays (Figure 2 and 3). Micrographs of DMSO-treated control (Figure 4A) showed a spatial biomass distribution of the formed biofilms and a relatively thick coating of the biofilms with well-organized architecture characterized by large clumps over a glass surface. However, after treatment with celastrol (Figure 4B and 4C), the established biofilms were disrupted and scattered as microcolonies, wherein the biofilm cells remained-in the form of elongated chains, indicating that cells within the formed biofilms were probably challenged, killed, and detached from the established biofilms. It has been reported that phenotypic heterogeneity can occur in response to stressors, including antibiotic treatment. This is one of the mechanisms used by the bacteria to survive the changes in the environment [30]. In addition, the other study showed that phenotypic heterogeneity was observed when S. maltophilia K279a was challenged with ampicillin, resulting in the formation of long cell chains [31]. The results of this study are consistent with our findings in S. maltophilia ATCC 13637 after treatment with celastrol. Because microbial biofilms were formed at the bottom and sides of the 96-well plates, CLSM was used to visualize the biofilms formed on the glass bottom plates. Microscopic examination showed that celastrol at 40 and 80 μg/ml markedly eradicated the biofilms formed on the glass bottom surface. It also penetrated the biofilm, eventually killing the bacteria.
Microscopic evaluation of the effect of celastrol on established biofilms formed on glass surface using a confocal laser scanning microscope. CLSM micrographs of S. maltophilia ATCC 13637 biofilms treated with (A) DMSO 1 % (v/v) (control) and celastrol at concentration of (B) 40 μg/mL or (B) 80 μg/mL. Green (FITC) and red (Concanavalin A) colors indicate proteins and carbohydrates of EPS, respectively. Blue fluorescent staining (DAPI) represents extracellular DNA. CLSM images of panel (A) of DMSO-treated control S. maltophilia biofilms have spatial biomass distribution of mature biofilms and thick coating of biofilm with compact architecture characterized by large clumps. On the other hand, in panels (B) and (C) CLSM images of S. maltophilia biofilm treated with celastrol at 40 and 80 μg/mL, the established biofilms are almost disrupted and biofilm cells are remained as long cell chains. Magnification: 400x for each dose. Scale bar: 50 μm.
Celastrol reduced the swimming motility of S. maltophilia
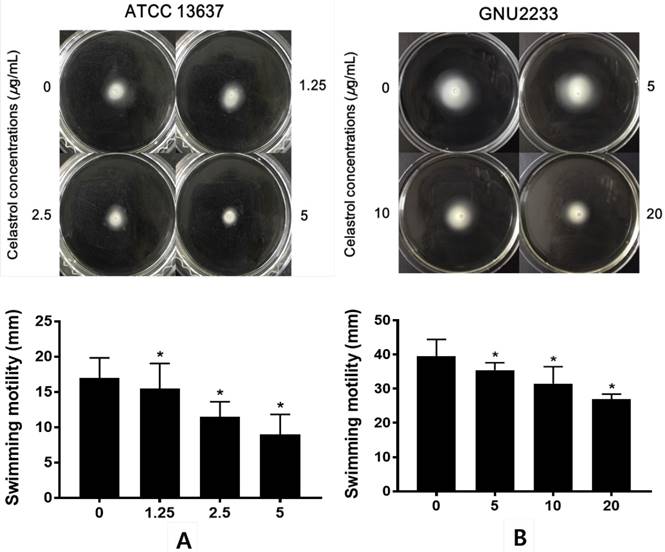
The effects of celastrol on the swimming motility of S. maltophilia strains (ATCC 13637 and GNU2233) were examined in a phenotypic study using semisolid agar. As shown in Figure 5, after 24-hour incubation, S. maltophilia showed a significant (p < 0.05) decrease in the diameter of the swimming zones in the agar plate containing celastrol. Celastrol decreased the diameter in a dose-dependent manner. Since the concentrations of celastrol tested were sub-inhibitory concentrations, the reduction of swimming motility could not be attributed to growth disturbance. In particular, the swimming motility zone of ATCC 13637 strain treated with celastrol at 5 μg/ml was 9 mm in diameter, which was smaller than that of the DMSO-treated control by 47.06%. In addition, celastrol at 20 μg/ml reduced the swimming motility of GNU2233 strain by more than 31.65%. Therefore, celastrol at sub-inhibitory concentrations reduced the swimming motility of both S. maltophilia strains.
Effect of celastrol on motility of S. maltophilia. Motility assays were performed using 0.15% agar plate containing 1% tryptone and 0.5% NaCl. (A) ATCC 13637 and (B) GNU2233 strains were inoculated in the center of motility agar surface including DMSO 1% (v/v) or celastrol at sub-inhibitory concentrations. The inoculated plates were incubated at 37 °C and the diameter (mm) of circular swimming zone was measured at 24 h. Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to that of DMSO treated control.
Celastrol inhibited protease secretion by S. maltophilia
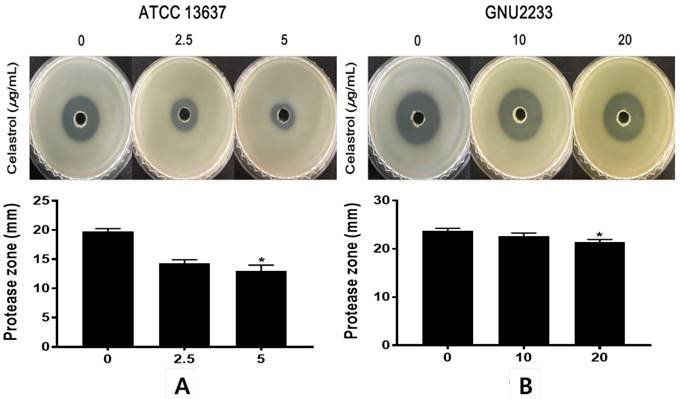
To investigate the effects of celastrol on the secretion of extracellular protease, another virulence-associated factor of S. maltophilia, a phenotypic assay was performed using a casein agar plate. After 48-hour incubation in casein agar plate, the hydrolysis zone formed around the inoculated supernatant was measured. As shown in Figure 6, celastrol at sub-inhibitory concentrations dose-dependently inhibited (p < 0.05) the proteolytic activity of S. maltophilia strains (ATCC 13637 and GNU2233). Notably, celastrol at 5 μg/ml significantly (p < 0.05) reduced the diameters of the halo zones of the ATCC 13637 strain by more than 49.1%, compared to the DMSO-treated control. In addition, S. maltophilia GNU2233 treated with celastrol at 20 μg/ml showed a decrease in the proteolytic activity by approximately 13.62%. These results showed that celastrol at sub-inhibitory concentrations inhibited protease secretion by S. maltophilia strains.
Effect of celastrol on protease production of S. maltophilia. Proteolytic activity was determined using casein agar. (A) ATCC 13637 and (B) GNU2233 cells were incubated with or without sub-inhibitory concentrations of celastrol at 37 °C for 24 h. Secreted protease assays were performed using culture supernatants. Diameters of transparent zones surrounding the holes were measured at 48 h. Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to DMSO treated controls.
Celastrol induced transcriptional changes in virulence- and biofilm-related genes of S. maltophilia
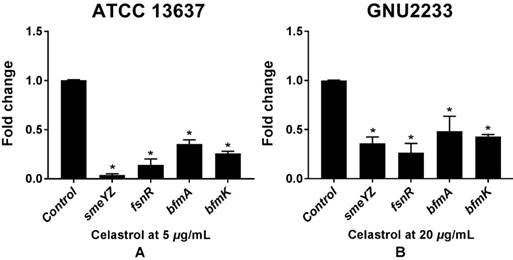
To investigate the molecular mechanisms of the anti-biofilm and anti-virulence activities of celastrol, qRT-PCR was performed to measure the expression of the genes (Table 1) related to biofilm formation and virulence in both celastrol-treated and DMSO-treated S. maltophilia cells (ATCC 13637 and GNU2233). Celastrol affected the expression of many genes, compared to the internal housekeeping control gene (Figure 7). In consistence with the results of inhibition of biofilm formation (Figure 1) and attenuation of virulence-related characteristics (Figure 5 and 6), celastrol significantly reduced the gene expression of smeYZ, encoding SmeYZ efflux pump (involved in swimming, protease secretion, and biofilm formation), by more than 30- and 3-fold in the ATCC 13637 and GNU2233 strains, respectively (Figure 7A and 7B, respectively; p < 0.05). In addition, celastrol significantly decreased the gene expression of fsnR, encoding the orphan response regulator FsnR (involved in motility and biofilm formation), by more than 10- and 3-fold in the ATCC 13637 and GNU2233 strains, respectively (Figure 7A and 7B, respectively; p < 0.05). Furthermore, the expression of two HK genes, encoding the BfmAK system involved in biofilm formation and development, was significantly down-regulated by celastrol by more than 3-fold in both S. maltophilia strains (Figure 7Aa and 7B; p < 0.05). Overall, the repression of these genes by celastrol confirmed the observed inhibitory effects of celastrol against the virulence factors and biofilm formation in S. maltophilia.
Transcriptional profiles of S. maltophilia cells treated with or without celastrol. ATCC 13637 and GNU 2233 were cultivated in CA-MHB medium at 37 °C for 6 h with shaking (250 rpm). Thereafter, they were further incubated at 37 °C for 4 h with or without celastrol at (A) 5 μg/mL (ATCC 13637) or (B) 20 μg/mL (GNU2233) without shaking. Transcriptional profiles were examined by qRT-PCR analysis. Relative gene expressions indicate transcriptional levels after treatment with celastrol versus DMSO treated control (value of 1.0). The tmRNA was used to normalize the expression levels of genes tested. All measurements were conducted with three individual cultures. Additionally, three replications were used for each qRT-PCR reaction. Mean values of triplicate independent studies are shown. Error bars represent standard deviations (SDs). Asterisks (*) indicate statistically significant change (p < 0.05) compared to that of DMSO treated control.
Discussion
The ever increasing emergence of multidrug resistant opportunistic pathogens, such as S. maltophilia, in clinical settings has become a worldwide public health problem owing to the lack of effective treatment measures, its intrinsic multidrug resistance, antibiotic-modifying enzymes, and antibiotic-resistant biofilm formation. Furthermore, superbugs such as S. maltophilia can cause chronic infections through formation of biofilms, which are difficult to encounter with conventional antibiotics [3]. Thus, discovery and development of alternative anti-pathogenic agents is urgently needed to reduce the emergence of antibiotic resistance and overcome life-threatening infections caused by antibiotic-resistant organisms. This alternative approach, unlike the conventional antibiotics that aim to inhibit cell growth, can combat the pathogenesis of biofilm-related infections caused by multidrug resistant pathogens without increasing the risk of drug resistance emergence [32]. The present study provided phenotypic and genotypic evidence that celastrol could be a potential anti-pathogenic agent for treatment of S. maltophilia infections since it showed significant anti-biofilm and anti-virulence activities against S. maltophilia without affecting their planktonic cell growth.
Recent pharmacological and clinical studies have shown that friedelane-type triterpenoids exert various pharmacological effects, including anti-oxidant, anti-tumor, anti-inflammatory, anti-HIV, and anti-bacterial bioactivities [33]. Herein, at sub-inhibitory concentrations, celastrol was not lethal to S. maltophilia. It dose-dependently decreased biofilm formation, motility, and protease secretion by S. maltophilia. Additionally, celastrol disrupted the previously established biofilms and interfered with the expression of biofilm- and virulence-associated genes.
The mechanisms underlying biofilm formation in S. maltophilia involve a complex network modulated by environmental factors, including temperature, phosphate, pH, antibiotics, metal ions, and other factors [3]. Moreover, S. maltophilia genome contains diverse array of multidrug efflux pumps and many other molecular machineries that can increase the resistance of S. maltophilia to various antibiotics [6]. In this regard, S. maltophilia infections have become a critical threat to human health.
To gain further insights into whether celastrol could inhibit the molecular mechanisms involved in biofilm formation and repress the expression of virulence-associated genes in S. maltophilia, the transcription levels of various genes were examined using qRT-PCR analysis. As expected, biofilm and virulence regulators (SmeYZ, FsnR, and BfmAK system) were significantly down-regulated by celastrol (Figure 7), resulting in reduced biofilm formation (Figure 1) and attenuated virulence-related characteristics (Figure 5 and 6), which validated the phenotypic effects of celastrol.
It has been shown that the RND-type efflux pump, SmeYZ contributes to intrinsic and acquired multidrug resistance to aminoglycosides and trimethoprim-sulfamethoxazole owing to its antimicrobial extrusion functions [34]. In addition, it is involved in biofilm formation, flagella formation, oxidative stress susceptibility, swimming motility, and protease secretion [8], which supported our findings that suppression of smeYZ gene expression (Figure 7) correlated with the attenuation of biofilm formation Figure 1), swimming motility (Figure 5), and protease secretion (Figure 6).
A previous study has reported that fsnR gene encodes the orphan response regulator, FsnR. Genetic and biochemical evidence suggested that FsnR could trigger the transcription of at least two flagella-related operons and mediate cell motility and biofilm formation. Suppression of fsnR gene expression is a crucial strategy to inhibit S. maltophilia cell motility and biofilm formation [19]. Our results of the transcriptional analysis showed that celastrol inhibited the expression of fsnR gene (Figure 7). These findings suggested that down-regulation of fsnR gene expression could interfere with the biofilm formation ability of S. maltophilia, thus eradicating S. maltophilia infections, particularly those biofilm-related infections.
A recent study has suggested that two HK genes encode a two-component TCS, the BfmAK system. This system has been found to play multiple roles in the formation of S. maltophilia biofilms [20]. BfmA has been found to be a transcription factor that stimulates the transcription of bfmA-bfmK operon and smlt0800 (acoT, a gene encoding acyl coenzyme A thioesterase) by directly binding to their promoter regions; in addition, it has been linked to biofilm development [20]. Based on our results, inhibition of bfmA and bfmk gene expression by celastrol possibly contributed to the reduction of biofilm formation in S. maltophilia (Figure 1 and 7). These findings suggested that celastrol could inhibit the expression of several important biofilm- and virulence- regulating genes in S. maltophilia.
When mature biofilms are completely formed by dreadful biofilm-forming pathogens, conventional antibiotics usually fail to eradicate the infection because the multilayer architecture of the biofilm acts as a diffusion barrier. This can provide protection to biofilm-grown cells against antimicrobial agents or antibiotics. Therefore, high biofilm disrupting concentrations are required [35], and strategies focusing on destroying the formed biofilms are of particular importance. The effects of celastrol on the established biofilms of S. maltophilia strains were further tested using microtiter plate assay and CLSM. Results revealed that celastrol not only induced disruption of the complex architecture of the biofilms but also killed many sessile cells within the biofilms by loosening the microcolonies. It is plausible to infer that this is linked to increased antibiotic penetration promoted by celastrol into the established biofilms of S. maltophilia. It is noteworthy that this does make clinical sense; otherwise, biofilm-related clinical infections will proceed without prevention.
To counter the dreadful pathogenic bacteria, antibiotics, which can induce resistance emergence, have been used for a long time. Therefore, novel strategies are urgently needed to control antibiotic-resistant S. maltophilia. To decrease the bacterial virulence and overcome biofilm-associated resistance, prevention of biofilm formation, eradication of pre-existing biofilms, and halting of the ongoing virulence factor production could be used as alternative control strategies. Results of this study showed that celastrol, a pentacyclic triterpenoid, possessed anti-biofilm and anti-virulence activities. It also inhibited the expression of genes related to biofilm formation and virulence of S. maltophilia.
Overall, celastrol might be a potential anti-pathogenic agent by targeting biofilm formation and virulence factors without inhibiting bacterial growth. It could be administered in combination with antibiotics or other bactericides to induce synergistic effects and achieve better control of persistent S. maltophilia infections. In addition, the anti-biofilm and anti-virulence activities of celastrol demonstrated the value of plant-derived compounds as promising and potent bioactive agents.
Abbreviations
ATCC: American type culture collection; CA-MHB: cation-adjusted Mueller Hinton broth; CLSM: confocal laser scanning microscopy; ConA: Concanavalin A; DAPI: 4',6-diamidino-2-phenylindole dihydrochloride; DMSO: dimethyl sulfoxide; FITC: fluorescein isothiocyanate; MIC: minimum inhibitory concentration; PBS: phosphate buffered saline; qRT-PCR: quantitative reverse transcription-polymerase chain reaction; RND: resistance-nodulation-division.
Acknowledgements
This research was supported by the Soonchunhyang University Research Fund and by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2017R1D1A1B03032960).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Pompilio S, Pomponio V. et al. Phenotypic and genotypic characterization of Stenotrophomonas maltophilia isolates from patients with cystic fibrosis: genome diversity, biofilm formation, and virulence. BMC Microbiol. 2011;11:159
2. Vidigal PG, Mosel F, Koehling HL. et al. Delineation of Stenotrophomonas maltophilia isolates from cystic fibrosis patients by fatty acid methyl ester profiles and matrix-assisted laser desorption/ionization time-of-flight mass spectra using hierarchical cluster analysis and principal component analysis. J Med Microbiol. 2014;63:1615-1620
3. Brooke JS. Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clin Microbiol Rev. 2012;25(1):2-41
4. Looney WJ, Narita M, Mühlemann K. Stenotrophomonas maltophilia: an emerging opportunist human pathogen. Lancet Infect Dis. 2009;9(5):312-323
5. Sanchez MB, Hernandez A, Martinez JL. Stenotrophomonas maltophilia drug resistance. Future Microbiol. 2009;4(6):655-660
6. Crossman LC, Gould VC, Dow JM. et al. The complete genome, comparative and functional analysis of Stenotrophomonas maltophilia reveals an organism heavily shielded by drug resistance determinants. Genome Biol. 2008;9(4):R74
7. Poole K. Bacterial multidrug efflux pumps serve other functions. Microbe Mag. 2008;3(4):179-185
8. Lin YT, Huang YW, Chen SJ. et al. The SmeYZ efflux pump of Stenotrophomonas maltophilia contributes to drug resistance, virulence-related characteristics, and virulence in mice. Antimicrob Agents and Chemother. 2015;59(7):4067-4073
9. Adamek M, Overhage J, Bathe S. et al. Genotyping of environmental and clinical Stenotrophomonas maltophilia isolates and their pathogenic potential. PLoS One. 2011;6(11):e27615
10. Huang TP, Somers EB, Wong AC. Differential biofilm formation and motility associated with lipopolysaccharide/exopolysaccharide-coupled biosynthetic genes in Stenotrophomonas maltophilia. J Bacteriol. 2006;188(8):3116-3120
11. Passerini de Rossi B, Calenda M, Vay C. et al. Biofilm formation by Stenotrophomonas maltophilia isolates from device-associated nosocomial infections. Rev Argent Microbiol. 2007;39(4):204-212
12. Pompilio A, Piccolomini R, Picciani C. et al. Factors associated with adherence to and biofilm formation on polystyrene by Stenotrophomonas maltophilia: the role of cell surface hydrophobicity and motility. FEMS Microbiol Lett. 2008;287(1):41-47
13. Di Bonaventura G, Spedicato I, D'Antonio D. et al. Biofilm formation by Stenotrophomonas maltophilia: modulation by quinolones, trimethoprim-sulfamethoxazole, and ceftazidime. Antimicrob Agents Chemother. 2004;48(1):151-160
14. Passerini de Rossi B, Feldman L, Pineda MS. et al. Comparative in vitro efficacies of ethanol-, EDTA- and levofloxacin-based catheter lock solutions on eradication of Stenotrophomonas maltophilia biofilms. J Med Microbiol. 2012;61:1248-1253
15. Pompilio A, Catavitello C, Picciani C. et al. Subinhibitory concentrations of moxifloxacin decrease adhesion and biofilm formation of Stenotrophomonas maltophilia from cystic fibrosis. J Med Microbiol. 2010;59:76-81
16. Donlan RM, Costerton JW. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev. 2002;15(2):167-193
17. Wu K, Yau YC, Matukas L. et al. Biofilm compared to conventional antimicrobial susceptibility of Stenotrophomonas maltophilia isolates from cystic fibrosis patients. Antimicrob Agents Chemother. 2013;57(3):1546-1548
18. Balcázar JL, Subirats J, Borregao CM. The role of biofilms as environmental reservoirs of antibiotic resistance. Front Microbiol. 2015;6:1216
19. Kang XM, Wang FF, Zhang H. et al. Genome-wide identification of genes necessary for biofilm formation of nosocomial pathogen Stenotrophomonas maltophilia reveals orphan response regulator FsnR is a critical modulator. Appl Environ Microbiol. 2015;81(4):1200-1209
20. Zheng L, Wang FF, Ren BZ. et al. Systematic Mutational Analysis of Histidine Kinase Genes in the Nosocomial Pathogen Stenotrophomonas maltophilia Identifies BfmAK System Control of Biofilm Development. Appl Environ Microbiol. 2016;82(8):2444-2456
21. Hentzer M, Givskov M. Pharmacological inhibition of quorum sensing for the treatment of chronic bacterial infections. J Clin Invest. 2003;112(9):1300-1307
22. Molhoek EM, van Dijk A, Veldhuizen EJ. et al. A cathelicidin-2-derived peptide effectively impairs Staphylococcus epidermidis biofilms. Int J Antimicrob Agents. 2011;37(5):476-479
23. de Lima Pimenta A, Chiaradia-Delatorre LD, Mascarello A. et al. Synthetic organic compounds with potential for bacterial biofilm inhibition, a path for the identification of compounds interfering with quorum sensing. Int J Antimicrob Agents. 2013;42(6):519-523
24. Shrivastava S, Jeengar MK, Reddy VS. et al. Anticancer effect of celastrol on human triple negative breast cancer: possible involvement of oxidative stress, mitochondrial dysfunction, apoptosis and PI3K/Akt pathways. Exp Mol Pathol. 2015;98(3):313-327
25. Youn GS, Kwon DJ, Ju SM. et al. Celastrol ameliorates HIV-1 Tat-induced inflammatory responses via NF-kappaB and AP-1 inhibition and heme oxygenase-1 induction in astrocytes. Toxicol Appl Pharmacol. 2014;280(1):42-52
26. Woo SG, Lee SY, Lee SM. et al. Activity of novel inhibitors of Staphylococcus aureus biofilms. Folia Microbiol. 2017;62(2):157-167
27. Field D, O' Connor R, Cotter PD. et al. In vitro activities of nisin and nisin derivatives alone and in combination with antibiotics against Staphylococcus biofilms. Front Microbiol. 2016;7:508
28. Nosyk O, ter Haseborg E, Metzger U. et al. A standardized pre-treatment method of biofilm flocs for fluorescence microscopic characterization. J Microbiol Methods. 2008;75(3):449-456
29. Murray TS, Kazmierczak BI. FlhF is required for swimming and swarming in Pseudomonas aeruginosa. J Bacteriol. 2006;188(19):6995-7004
30. Ackermann M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat Rev Microbiol. 2015;13(8):497-508
31. Abda EM, Krysciak D, Krohn-Molt I. et al. Phenotypic heterogeneity affects Stenotrophomonas maltophilia K279a colony morphotypes and β-lactamase expression. Front Microbiol. 2015;6:1373
32. Sperandio V. Novel approaches to bacterial infection therapy by interfering with bacteria-to-bacteria signaling. Expert Rev Anti Infect Ther. 2007;5(2):271-276
33. Schmidt BM, Ribnicky DM, Lipsky PE. et al. Revisiting the ancient concept of botanical therapeutics. Nat Chem Biol. 2007;3(7):360-366
34. Wu CJ, Huang YW, Lin YT. et al. Inactivation of SmeSyRy Two-Component Regulatory System Inversely Regulates the Expression of SmeYZ and SmeDEF Efflux Pumps in Stenotrophomonas maltophilia. PLoS One. 2016;11(8):e0160943
35. Tu Quoc PH, Genevaux P, Pajunen M. et al. Isolation and characterization of biofilm formation-defective mutants of Staphylococcus aureus. Inf Immun. 2007;75(3):1079-1088
Author contact
Corresponding author: E-mail: omnibinac.kr Tel: +82-41-530-3039, Fax: +82-41-530-3085, Address: Department of Biomedical Laboratory Science, College of Medical Sciences, Soonchunhyang University, 22 Soonchunhyang-ro, Shinchang-myeon, Asan-si, Chungcheongnam-do 31538, Republic of Korea

 Global reach, higher impact
Global reach, higher impact