Impact Factor
ISSN: 1449-1907
Int J Med Sci 2021; 18(7):1648-1656. doi:10.7150/ijms.53575 This issue Cite
Research Paper
Systematic elucidation of the mechanism of Jingyin granule in the treatment of Novel Coronavirus (COVID-19) Pneumonia via Network Pharmacology
1. Institute of Infection Disease, Department of Liver Diseases, Central Laboratory, ShuGuang Hospital affiliated to Shanghai University of Chinese Traditional Medicine, Shanghai, China.
2. Department of Liver Surgery, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai, China.
Abstract
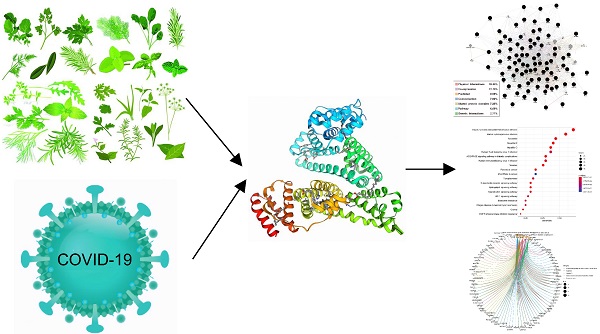
Background: Jingyin granule is one of the widely used traditional Chinese medicine mixture composed of multiple herbs in the treatment of respiratory system diseases. The mechanism of its therapeutic effects has still been obscure. The aim of this study is to use the network pharmacology approach for identification of the main active ingredients of Jingyin granule against COVID-19 targets and to explore their therapeutic mechanism.
Material and Method: In this study, the ingredients of Jingyin granule were evaluated by the usage of Traditional Chinese Medicine Systems Pharmacology Database and Traditional Chinese Medicine Integrated Database, and the interactions between potential gene targets and ingredients were identified using the SwissTargetPrediction database. Meanwhile the possible efficient targets COVID-19 acts on were identified via Online Mendelian Inheritance in Man database, DisGeNET database and GeneCards database. In addition, functions, components and pathways were identified by Gene Ontology enrichment analysis and Kyoto Encyclopedia of Genes and Genomes pathway analysis. Protein interaction, ingredients-targets network was established.
Results: Our findings showed that numerous ingredients of Jingyin granule could act on COVID-19 with 88 target genes. GO enrichment analysis, KEGG pathway analysis, and protein-protein interaction network revealed that these targets were interrelated with regulation of immune function, directly targeting disease genes.
Conclusions: Jingyin granule could be utilized to exert systematic pharmacological effects. Jingyin granule could directly target the major genes, and also regulate the immune system, acting as oblique disease treatment.
Keywords: Jingyin granule, ingredient, druggability, network pharmacology

 Global reach, higher impact
Global reach, higher impact