Impact Factor ISSN: 1449-1907
Int J Med Sci 2022; 19(9):1482-1501. doi:10.7150/ijms.73404 This issue Cite
Research Paper
Comprehensive Analysis of Transcriptional Expression of hsa-mir-21 Predicted Target Genes and Immune Characteristics in Kidney Renal Clear Cell Carcinoma
1. Department of Urological Surgery, Second Affiliated Hospital of Shantou University Medical College, Shantou, Guangdong, China.
2. Department of Hematology, Second Affiliated Hospital of Shantou University Medical College, Shantou, Guangdong, China.
3. Department of Clinical Laboratory Medicine, Second Affiliated Hospital of Shantou University Medical College, Shantou, Guangdong, China.
*Co-first authors with equal contributions to this work.
Abstract
Background: To uncover advanced prognosis biomarkers in patient with kidney renal clear cell carcinoma (KIRC), our study was the first to make a comprehensive analysis of hsa-mir-21 predicted target genes and explore the immune characteristics in KIRC.
Methods: In this study, the comprehensive analysis of hsa-mir-21 predicted target genes and immune characteristics in KIRC were analyzed via TIMER2.0, UALCAN, Metascape, Kaplan-Meier plotter, Human Protein Atlas, CancerSEA, JASPAR, GEPIA, R package: GSVA package (version 1.34.0) & immune infiltration algorithm (ssGSEA) and R package: RMS package (version 6.2-0) & SURVIVAL package (version 3.2-10).
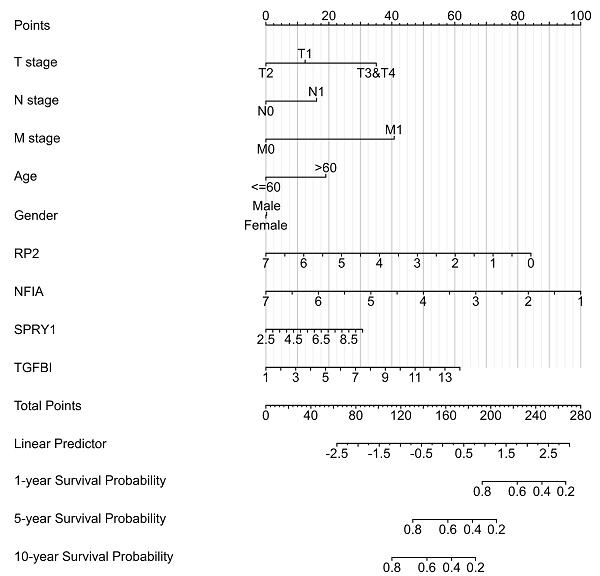
Results: Up-transcriptional expressions of RP2, NFIA, SPRY1 were significantly associated with favorable prognosis in KIRC, whereas that of TGFBI was markedly significantly to unfavorable prognosis. Additionally, RP2, NFIA, SPRY1 and TGFBI were significantly relevant to the immune infiltration in KIRC. Finally, ZNF263 was a common predicted transcription factor of RP2, NFIA, SPRY1 and TGFBI, which can as an independent indicator for prognosis in KIRC patients.
Conclusions: Hsa-mir-21 predicted target genes (RP2, NFIA, SPRY1 and TGFBI) and the common transcription factor ZNF263 could be the advanced prognosis biomarkers in KIRC patients.
Keywords: Hsa-mir-21, Predicted Target Gene, Immune Characteristics, Transcription Factor, Prognosis, Prediction Model, Kidney Renal Clear Cell Carcinoma

 Global reach, higher impact
Global reach, higher impact