Impact Factor
ISSN: 1449-1907
Int J Med Sci 2022; 19(9):1417-1429. doi:10.7150/ijms.73305 This issue Cite
Research Paper
Identification of three hub genes related to the prognosis of idiopathic pulmonary fibrosis using bioinformatics analysis
1. Department of respiratory and critical care medicine, the first affiliated hospital of Anhui medical university, Hefei 230022, China.
2. Department of Infectious Diseases, Hefei second people's hospital, Hefei 230001, China.
3. Department of occupational medicine, Hefei third clinical college of Anhui Medical University, Hefei 230022, China.
4. Department of pulmonary and critical care medicine, Suzhou Dushu Lake Hospital, Suzhou, 215006, China.
5. Department of pulmonary and critical care medicine, Dushu Lake Hospital Affiliated to Soochow University, Medical Center of Soochow University, Suzhou, 215006, China.
* These authors contributed equally to this work.
Abstract
Background: Idiopathic pulmonary fibrosis (IPF) is a chronic respiratory disease characterized by peripheral distribution of bilateral pulmonary fibrosis that is more pronounced at the base. IPF has a short median survival time and a poor prognosis. Therefore, it is necessary to identify effective prognostic indicators to guide the treatment of patients with IPF.
Methods: We downloaded microarray data of bronchoalveolar lavage cells from the Gene Expression Omnibus (GEO), containing 176 IPF patients and 20 controls. The top 5,000 genes in the median absolute deviation were classified into different color modules using weighted gene co-expression network analysis (WGCNA), and the modules significantly associated with both survival time and survival status were identified as prognostic modules. We used Lasso Cox regression and multivariate Cox regression to search for hub genes related to prognosis from the differentially expressed genes (DEGs) in the prognostic modules and constructed a risk model and nomogram accordingly. Moreover, based on the risk model, we divided IPF patients into high-risk and low-risk groups to determine the biological functions and immune cell subtypes associated with the prognosis of IPF using gene set enrichment analysis and immune cell infiltration analysis.
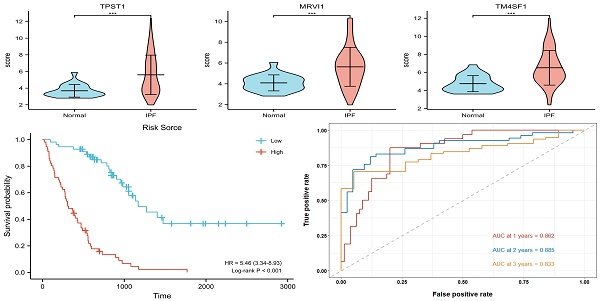
Results: A total of 153 DEGs located in the prognostic modules, three (TPST1, MRVI1, and TM4SF1) of which were eventually defined as prognostic hub genes. A risk model was constructed based on the expression levels of the three hub genes, and the accuracy of the model was evaluated using time-dependent receiver operating characteristic (ROC) curves. The areas under the curve for 1-, 2-, and 3-year survival rates were 0.862, 0.885, and 0.833, respectively. The results of enrichment analysis showed that inflammation and immune processes significantly affected the prognosis of patients with IPF. The degree of mast and natural killer (NK) cell infiltration also increases the prognostic risk of IPF.
Conclusions: We identified three hub genes as independent molecular markers to predict the prognosis of patients with IPF and constructed a prognostic model that may be helpful in promoting therapeutic gains for IPF patients.
Keywords: IPF, Prognosis, Bronchoalveolar lavage cells, Genes, Bioinformatics

 Global reach, higher impact
Global reach, higher impact