Impact Factor ISSN: 1449-1907
Int J Med Sci 2022; 19(8):1357-1363. doi:10.7150/ijms.73150 This issue Cite
Review
Research Progress in the Medical Application of Heavy Water, Especially in the Field of D2O-Raman Spectroscopy
Department of Breast Surgery, First Hospital of Jilin University, Changchun, Jilin 130021
Received 2022-3-22; Accepted 2022-7-7; Published 2022-7-18
Abstract
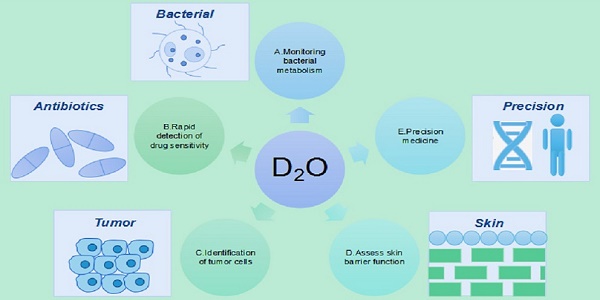
Heavy water is an ideal contrast agent for metabolic activity and can be adapted to a wide range of biological systems owing to its non-invasiveness, universal applicability, and cost-effectiveness. As a new type of probe, the heavy isotope of water has been widely used in the study of cell development, metabolism, tissue homeostasis, aging, and tumor heterogeneity. Herein, we review findings supporting the applications of and research on heavy water in monitoring of bacterial metabolism, rapid detection of drug sensitivity, identification of tumor cells, precision medicine, and evaluation of skin barrier function and promote the use of heavy water as a suitable marker for the development of detection and treatment methodologies.
Keywords: heavy water, deuterium oxide, Raman microspectroscopy, isotypes, probe, medicine.
Introduction
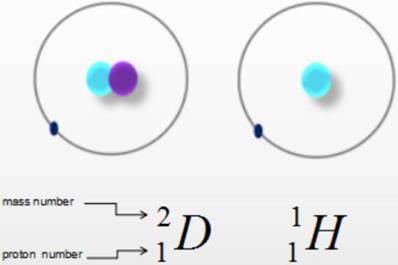
Heavy water, also known as deuterium oxide or deuterium water, is a compound of heavy hydrogen (D) and oxygen (O) (Fig. 1). A molecule of heavy water is composed of two heavy hydrogen and one oxygen atom, with the molecular and chemical formula D2O. Heavy water is similar to ordinary water in appearance but has a higher density of 1.1079 g/cm3, as well as higher freezing and boiling points of 3.82 °C and 101.42 °C, respectively. The relative molecular mass of heavy water, 20.0275 Mr, is higher than that of water (H2O, 18.0153 Mr) by about 11%, hence the name “heavy” water. The differences between deuterium and hydrogen are negligible; while they have different neutron numbers, mass numbers, and physical properties, they have the same number of protons, number of outermost electrons, and chemical properties. Hence, the chemical properties of heavy water and ordinary water are also very similar. It has been proved that a small amount of heavy water does not cause adverse reactions [1-2]. Heavy water has been widely used as a probe in many fields.
Chemical formula. Water is a compound of hydrogen (H) and oxygen (O), while heavy water is a compound of heavy hydrogen (D) and oxygen (O).
Quantitative determination of heavy water concentration
Currently, the commonly used methods to quantify heavy water concentration include the density method, mass spectrometry, and spectroscopy techniques [3]. Among these, Raman spectroscopy has attracted increasing attention as a highly sensitive molecular imaging technique for studying complex biological systems, including cells, tissues, and various biological materials [4]. Raman spectroscopy analyzes the biochemical components of a sample by measuring the inelastic scattering of light by different molecular species, producing a spectrum based on the chemical bonds present within the analyzed samples. This technique allows non-destructive, label-free spectral imaging and analysis of cells, tissues, and nanoparticles [5-7]. As a new quantitative method, heavy water-labeled single-cell Raman microspectroscopy can reduce damage to cells and human interference factors. This method does not alter the cell composition or rely on cell culture and thus allows the rapid, quantitative, and nondestructive evaluation of the effect of drugs on the real-time metabolic activity of microorganisms at the single-cell level [8-10]. Simultaneously, the results of Raman spectroscopy provide abundant information, which can reflect the biochemical and structural characteristics of the material of interest. In theory, this method can detect the entire spectrum of substances in the cell, especially the “fingerprint area” in the range of ~600-1800 cm-1, which can distinguish cell types and explore the mechanism of microbial stress [11-13]. Thus, this technique has a broad application prospect and important guiding significance.
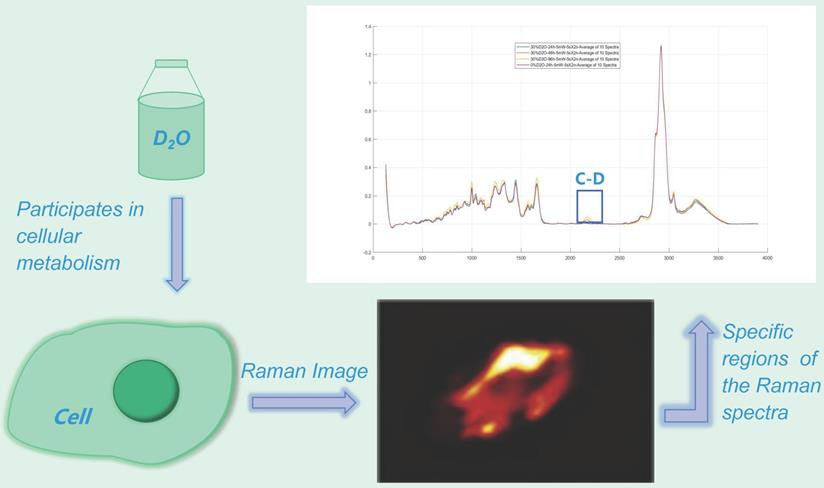
The presence of heavy water results in heavy water peaks (C-D peaks) in specific regions (~2040-2300 cm-1) of the Raman spectra of metabolically active microorganisms (Fig. 2) [14-15], which correspond to symmetric and asymmetric C-D stretching vibrations caused primarily by lipids and proteins [12,16-17]. Furthermore, the minimum inhibitory concentration based on metabolic activity (MIC-MA) can be used to quantitatively evaluate the effect of drugs on real-time metabolic activity of microorganisms based on the ΔC-D ratio (the difference between the current time and the baseline C-D) [8-9]. This technology has been widely used in microbial species identification and has also been widely used to evaluate the effect of drugs on bacterial metabolic activity.
Heavy water Raman spectroscopy imaging. The heavy water peak (C-D peak) can be measured in the specific region of the Raman spectrum (~ 2040-2300 cm-1) by using a certain concentration of heavy-water cultured microorganisms.
Biosafety of heavy water
Compared with other isotope markers, heavy water exerts little effect on microorganisms; furthermore, it is cost effective. The growth of bacteria and single-cell fungi in heavy water within a certain concentration range was not significantly inhibited (P > 0.05) compared with that in normal water [8]. Prior studies have suggested that cells and single-celled fungi can tolerate a certain concentration of heavy water, and thus, heavy water toxicity is not a limiting factor in this technique. In mice, a concentration of heavy water below 20% had no effect on physiological processes or cell division; did not modulate physiology, growth, appetite, or reproduction; and was found to have no teratogenic effect in multi-generation studies [18-21]. Studies have found that drinking 60-70 ml of heavy water daily would not cause adverse reactions [23-24]. Therefore, deuterium has been widely used as a stable isotope to evaluate the human body composition and metabolic rate [19,24-25].
The determination of heavy water concentration was used for rapid evaluation of the effect of drugs on the metabolic activity of cells and single-cell fungi, with high sensitivity [26]. The labeling of heavy water as a probe has been widely used environmental studies. Li et al. used the heavy water labeling method to understand the metabolic flux of microbial communities in complex soil systems. Through heavy water modification of soil microorganisms, bacteria that release phosphate were identified according to the ratio of C-D stretching vibration to Raman spectroscopy [27]. Eichorst et al. also found the same method of heavy water labeling, which was more conducive to the objective evaluation of differences in metabolic activity of soil bacteria and proved that deuterium content was suitable for the detection of metabolic activity indicators [28]. A recent study by Taubert et al., using heavy water as a probe in the study of groundwater microbial communities, proved that this probe could identify active microorganisms in groundwater and their functional characteristics [29]. The special value of heavy water is reflected in the application of atomic energy technology. Heavy water reduces neutron velocity and controls nuclear fission by acting as a retarder in nuclear reactors.
Medical applications of heavy water in the field of D2O-Raman spectroscopy
Monitoring metabolism in individual bacteria
Using heavy water with stable isotope labeling, deuterium intake was found to be a reliable indicator of general bacterial metabolic activity [30-31]. Because neither H2O nor D2O have Raman peaks in the C-D region, the background from water did not interfere with this approach [32]. Therefore, studies have been established to examine the effects of carbon sources and bacteria on deuterium uptake by quantitatively measuring the assimilation of heavy water into a single bacterium [33-34]. The deuterium assimilation rate was higher in the presence of simple substrates, such as sugar, compared to that with complex carbon substrates, and the difference was significant in bacterial isolates. The quantitative determination of deuterium content in heavy water was further used to distinguish between various types of bacteria and their metabolic activities; thus, this detection method could be combined with chemometrics to construct a powerful bacterial monitoring method. The absorption of deuterium as a marker of bacterial metabolic activity determined using Raman microspectroscopy was strongly affected by the organic carbon source used by a single bacterium for growth, as well as by the cell itself [35].
Rapid detection technology of drug sensitivity
To counteract the common problem of antibiotic resistance [36-37], it is urgent that researchers develop technology to rapidly detect antibiotic drug sensitivity in clinic [38-39]. As metabolism suppressive drugs ultimately alter cellular macromolecule metabolism, a method for detecting macromolecule-specific metabolites following drug therapy would be valuable. The application of heavy water as a probe detection method and the C-D band shift as a biomarker of cellular metabolic activity to quantitatively evaluate the metabolic activity of bacteria in their environment in a culture-independent manner at the single-cell level has been proposed to determine the efficacy of antibiotics through metabolic inhibition [30]. In particular, the ability to evaluate specific metabolites and newly synthesized macromolecules provides greater insights into the underlying processes by which cells respond to drugs. Using heavy water as a probe to detect low numbers of bacteria solved the clinical problem of low sample content. By combining single-cell Raman spectroscopy and heavy water labeling, the active response of bacteria to antibiotics can be evaluated according to the deuterium-related characteristic peaks after only 30 minutes [40]. By evaluating the differences in heavy water assimilation activity between drug-resistant bacteria and sensitive bacteria under the action of antibiotics, the total time from urine collection to drug sensitivity reading can be reduced to 2.5 hours, which can guide clinicians to perform rapid diagnosis and screen effective antibiotics in time [41]. Therefore, this method can be the basis of a new antimicrobial screening platform at the single-cell level.
Identification of tumor stem cells
Fast-growing tumor cells also appear to incorporate more D from heavy water compared to other cells; this property would be detected directly with a Raman microscope. Using heavy water in combination with Raman microscopy, the boundaries of a tumor can be revealed by its inherently higher metabolic activities compared to the surrounding normal tissue. Thus, Raman spectroscopy using heavy water as a probe can help identify tumor stem cells with specific patterns of metabolic activity [42-44]. Studies have found that unsaturated lipid levels in ovarian cancer stem cells increased significantly compared to those in normal cells [45-46]. In addition, the combination of heavy water-Raman spectroscopy and fluorescence labeling can be used to monitor the metabolic activity of specific cells and lineages in situ, especially the metabolic cooperation between glial cells and neurons [47]. In addition to monitoring the synthesis of lipids and proteins [48], heavy water-Raman spectroscopy can be used to monitor protein turnover, lipid consumption, and macromolecular degradation in tumor cells [49].
Positioning of precise targeted drug use
Extracellular vesicles are biologically derived nanocarriers important for intercellular communication and transportation that have been proposed as disease biomarkers and therapeutic drug carriers [50-51]. The combination of heavy water-Raman spectroscopy imaging and bioactive molecules provides an opportunity to study the production and uptake of extracellular vesicles in various normal and dysfunctional states, as well as a direction for precise drug therapy [52-53]. Applying this approach to various newly designed drugs that target cell metabolism can identify which macromolecules are specifically targeted by the drug. This method can produce information-rich whole-cell spectral data and can be used to directly visualize and analyze extracellular vesicles at the two- and three-dimensional levels, thus providing guidance for the design of future extracellular vesicle treatment systems [54-55].
Assessment of skin barrier function
Heavy water can be used as an excellent and cost-effective probe to evaluate skin barrier function. The penetration dynamics of water can be regulated by the integrity of the skin barrier [56-57]. Therefore, the application of D2O is a promising method to assess the state of the skin barrier, considering the isotope substitution and diffusion behavior of water [58-59]. Owing to the different Raman spectral characteristics of the O-D bond of heavy water and the O-H bond of the skin, the influence of external osmotic water can be minimized [60-61]. The combination of heavy water and Raman microscopy can sensitively identify small changes in the molecular composition of skin and can detect skin at different depths. Heavy water, as a skin probe, can be used to detect skin water-related properties by extracting spectra from each pixel depth and distinguishing endogenous and exogenous hydrogen bonds [62-63]. By using different hydrogen-bonding water types to calculate the relative water content, the total water content of different skin depths can be calculated [64].
Other medical applications of heavy water
Deuterium in water has been reported to be rapidly balanced with mediators such as urine, saliva or serum [65-67], which can be used to measure the total amount of water in the body [68]. Lichtenbelt et al. showed that a 10-hour sampling time appears to be preferable for measuring total body water space and body composition by the deuterium-dilution technique [69]. Studies have shown that D2O can be used as a tracer to measure tissue perfusion and blood flow [70-71]. Recently, Lin Chen et al. demonstrated that D2O can be used as a new contrast agent to guide intravascular neurointervention and that deuterium-based MRI is a secure and practical method that can to accurately identify the perfusion area and to predict the affected area, which can guide real-time endovascular intervention [72]. Studies have demonstrated the potential of the deuterium compound-based MRS method in assessing tissue metabolokinetics [73]. DMRS-based deuterium metabolic imaging has also been shown to be useful in detecting tumor cells [74]. Recently, Laurie et al. proposed quantitative exchange-label turnover MRS, which can improve the sensitivity of the metabolic map and directly monitor cell metabolism in vivo. This approach is expected to be a new approach for exploring metabolic disorders in a wide range of human diseases [75]. Based on the hypothesis that the conversion of amino acids in the presence of D2O leads to the production of deuterium-labeled amino acids [76-77], some studies have proved that the rate of protein synthesis can be estimated under the action of D2O [77-79]. Herath et al. demonstrated that deuterium-based high-resolution mass spectrometry provides a useful method for the quantification of low levels of deuterium enrichment that is not limited to specific molecular classes; this method is expected to be useful for the study of metabolic flux of deuterium-labeled tracers [80].
Development prospects
The exploration of the applications of heavy water provides novel insights and aids the development of new diagnostic and therapeutic strategies. Heavy water can be applied to the study of a variety of developmental processes, including cell development, metabolism, tissue homeostasis, drug resistance, and aging. As a nondestructive, noninvasive, and context-free imaging method, the combination of heavy water and Raman microscopy can be used to visualize the kinetics of protein synthesis, lipid production, and DNA metabolism in various model organisms at a low cost and without tissue bias. The latest developments in the applications of the heavy water labeling method indicate the possibility of real-time tracking of single cells, thus providing further understanding of the transport of single cells and making the in vitro study of single cells possible. This method will facilitate the comprehensive real-time molecular characterization and imaging of single cells in vitro to promote the understanding of single-cell biology. The newly developed classification strategy based on the C-D stretching vibration range avoids interference from other Raman bands owing to its higher sensitivity and thus saves time by negating the need to analyze large data sets. Heavy water provides a basis for the rapid clinical diagnosis and selection of appropriate antibiotics to treat bacterial infection. In particular, it provides a valuable method to facilitate the treatment and diagnosis of critical bacterial infections and can also be used to assay drug susceptibility. The recognition ability of heavy water to cancer stem cells with specific metabolic activities provides a new direction and visual angle for the diagnosis and treatment of clinical tumors. The combination of heavy water with Raman spectroscopy could be used to reveal the molecular components with biochemical significance through multivariable analysis, such as in the rapid identification of metabolically active drug-resistant cells or cancer stem cells with metabolic pattern changes after chemotherapy or in the exploration of the roles of such cells in chemotherapy failure, and could thus provide a direction for the development of personalized cancer therapy. Through the determination of heavy water concentration, data processing, and analysis framework, we can further promote the comparative study of extracellular vesicle uptake under different conditions or by different cell types, which may provide further guidance in the study of the role of different molecules, targeting and uptake of extracellular vesicles, and design of extracellular vesicle-based therapeutics. Heavy water provides data on different penetration depths and molecular skin effects to varying degrees, supporting the idea of multiple roles of heavy water in the skin as a convenient and inexpensive target.
As an emerging class of biomarker, heavy water has considerable advantages over traditional markers. Currently, several studies have partially elucidated the value of heavy water in medical research. However, the applications of heavy water in medical research need to be further explored, and heavy water is expected to play a more important role in the development of novel clinical management modalities.
Abbreviations
D: heavy hydrogen; H2O: water; MIC-MA: minimum inhibitory concentration based on metabolic activity; O: oxygen.
Acknowledgements
We would like to thank Editage for English language editing.
Funding
This work was supported by the Science and Technology Development Project of Jilin Province (Grant numbers 20210401057YY and 20200403083SF to Song Dong).
Author contributions
X W: Conceptualization, Data curation, Writing—original draft, Writing—review and editing. NM L: Software, Visualization. YF Z: Methodology, Resources. F Y: Investigation, Validation. ZJ Z: Formal analysis. D S: Conceptualization, Funding acquisition, Project administration, Supervision. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Gisolfi CV, Summers RW, Schedl HP, Bleiler TL, Oppliger RA. Human intestinal water absorption: Direct vs. indirect measurements. Am J Physiol. 1990;258:G216-G222
2. Kushner DJ, Baker A, Dunstall TG. Pharmacological uses and perspectives of heavy water and deuterated compounds. Can J Physiol Pharmacol. 1999;77:79-88
3. Lemaster DM. Uniform and selective deuteration in two-dimensional NMR of proteins. Annu Rev Biophys Biophys Chem. 1990;19:243-266
4. Butler HJ, Ashton L, Bird B. et al. Using Raman spectroscopy to characterize biological materials. Nat Protoc. 2016;11:664-687
5. Penders J, Pence Ij, Horgan CC. et al. Single particle automated Raman trapping analysis. Nat Commun. 2018;9:4256
6. Puppels GJ, de Mul FF, Otto C. et al. Studying single living cells and chromosomes by confocal Raman microspectroscopy. Nature. 1990;347:301-303
7. Klein K, Gigler AM, Aschenbrenner T. et al. Label. Label-Free live-cell imaging with confocal Raman microscopy. Biophys J. 2012;102:360-368
8. Berry D, Mader E, Lee TK. et al. Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells. Proc Natl Acad Sci U S A. 2015;112:E194-E203
9. Tao YF, Wang Y, Huang S. et al. Metabolic-activity-based assessment of antimicrobial effects by D2O-labeled single-cell Raman microspectroscopy. Anal Chem. 2017;89:4108-4115
10. Wang Y, Song YZ, Tao Y. et al. Reverse and multiple stable isotope probing to study bacterial metabolism and interactions at the single cell level. Anal Chem. 2016;88:9443-9450
11. Potma E, de Boeij WP, van Haastert PJ, Wiersma DA. Real-time visualization of intracellular hydrodynamics in single living cells. Proc Natl Acad Sci U S A. 2001;98:1577-1582
12. Justice NB, Li Z, Wang YF. et al. (15)N- and (2)H proteomic stable isotope probing links nitrogen flow to archaeal heterotrophic activity. Environ Microbiol. 2014;16:3224-3237
13. Wegener G, Bausch M, Holler T. et al. Assessing sub-seafloor microbial activity by combined stable isotope probing with deuterated water and 13C-bicarbonate. Environ Microbiol. 2012;14:1517-1527
14. Kumar B N V, Guo S, Bocklitz T, Rösch P, Popp J. Demonstration of carbon catabolite repression in naphthalene degrading soil bacteria via Raman spectroscopy based stable isotope probing. Anal Chem. 2016;88:7574-7582
15. Kubryk P, Kölschbach JS, Marozava S. et al. Exploring the potential of stable isotope (resonance) Raman microspectroscopy and surface-enhanced Raman scattering for the analysis of microorganisms at single cell level. Anal Chem. 2015;87:6622-6630
16. Movasaghi Z, Rehman S, ur Rehman DI. Fourier transform infrared (FTIR) spectroscopy of biological tissues. Appl Spectrosc Rev. 2008;43:134-179
17. Matthäus C, Krafft C, Dietzek B. et al. Noninvasive imaging of intracellular lipid metabolism in macrophages by Raman microscopy in combination with stable isotopic labeling. Anal Chem. 2012;84:8549-8556
18. Jones PJ, Leatherdale ST. Stable isotopes in clinical research: Safety reaffirmed. Clin Sci (Lond). 1991;80:277-280
19. Hodel A, Gebbers JO, Cottier H, Laissue JA. Effects of prolonged moderate body deuteration on proliferative activity in major cell renewal systems in mice. Life Sci. 1982;30:1987-1996
20. Peng SK, Ho KJ, Taylor CB. Biologic effects of prolonged exposure to deuterium oxide. A behavioral, metabolic, and morphologic study. Arch Pathol. 1972;94:81-89
21. Katz JJ, Crespi HL, Hasterlik RJ, Thomson JF, Finkel AJ. Some observations on biological effects of deuterium, with special reference to effects on neoplastic processes. J Natl Cancer Inst. 1957;18:641-659
22. Neese RA, Misell LM, Turner S. et al. Measurement in vivo of proliferation rates of slow turnover cells by 2 H 2 O labeling of the deoxyribose moiety of DNA. Proc Natl Acad Sci USA. 2002;99:15345-15350
23. Guillermier C, Fazeli PK, Kim S. et al. Imaging mass spectrometry demonstrates age-related decline in human adipose plasticity. Jci Insight. 2017;2:e90349
24. Valencia ME, Alemán-Mateo H, Salazar G, Hernández Triana MH. Body composition by hydrometry (deuterium oxide dilution) and bioelectrical impedance in subjects aged >60 y from rural regions of Cuba, Chile and Mexico. Int J Obes. 2003;27:848-855
25. Schoeller DA. Recent advances from application of doubly labeled water to measurement of human energy expenditure. J Nutr. 1999;129:1765-1768
26. Hekmatara M, Heidari Baladehi MH, Ji YT, Xu J. D2O-probed Raman microspectroscopy distinguishes the metabolic dynamics of macromolecules in organellar anticancer drug response. Anal Chem. 2021;93:2125-2134
27. Li HZ, Bi QF, Yang K. et al. D2O-isotope-labeling approach to probing phosphate-solubilizing bacteria in complex soil communities by single-cell Raman spectroscopy. Anal Chem. 2019;91:2239-2246
28. Eichorst SA, Strasser F, Woyke T. et al. Advancements in the application of NanoSIMS and Raman microspectroscopy to investigate the activity of microbial cells in soils. Fems Microbiol Ecol. 2015 91
29. Taubert M, Stöckel S, Geesink P. et al. Tracking active groundwater microbes with D2O labelling to understand their ecosystem function. Environ Microbiol. 2018;20:369-384
30. Xu JB, Zhu D, Ibrahim AD. et al. Raman deuterium isotope probing reveals microbial metabolism at the single-cell level. Anal Chem. 2017;89:13305-13312
31. Song YZ, Cui L, López JÁS. et al. Raman-deuterium isotope probing for in-situ identification of antimicrobial resistant bacteria in Thames River. Sci Rep. 2017;7:16648
32. Brooker MH, Hancock G, Rice BC, Shapter J. Raman frequency and intensity studies of liquid H2O, H218O and D2O. J Raman Spectrosc. 1989;20:683-694
33. Kopf SH, McGlynn SE, Green-Saxena A. et al. Heavy water and (15) N labelling with NanoSIMS analysis reveals growth rate-dependent metabolic heterogeneity in chemostats. Environ Microbiol. 2015;17:2542-2556
34. Wang Y, Huang WE, Cui L, Wagner M. Single cell stable isotope probing in microbiology using Raman microspectroscopy. Curr Opin Biotechnol. 2016;41:34-42
35. Matanfack GA, Taubert M, Guo SX. et al. Influence of carbon sources on quantification of deuterium incorporation in heterotrophic bacteria: A Raman-Stable isotope labeling approach. Anal Chem. 2020;92:11429-11437
36. Andersson DI, Hughes D. Microbiological effects of sublethal levels of antibiotics. Nat Rev Microbiol. 2014;12:465-478
37. Zhu YG, Gillings M, Simonet P. et al. Microbial mass movements. Science. 2017;357:1099-1100
38. Kerremans JJ, Verboom P, Stijnen T. et al. Rapid identification and antimicrobial susceptibility testing reduce antibiotic use and accelerate pathogen-directed antibiotic use. J Antimicrob Chemother. 2008;61:428-435
39. van Belkum A, Bachmann TT, Lüdke G. et al. Developmental roadmap for antimicrobial susceptibility testing systems. Nat Rev Microbiol. 2019;17:51-62
40. Hong WL, Karanja CW, Abutaleb NS. et al. Antibiotic susceptibility determination within one cell cycle at single-bacterium level by stimulated Raman metabolic imaging. Anal Chem. 2018;90:3737-3743
41. Yang K, Li HZ, Zhu X. et al. Rapid antibiotic susceptibility testing of pathogenic bacteria using heavy-water-labeled single-cell Raman spectroscopy in clinical samples. Anal Chem. 2019;91:6296-6303
42. Sun XX, Yu Q. Intra-tumor heterogeneity of cancer cells and its implications for cancer treatment. Acta Pharmacol Sin. 2015;36:1219-1227
43. Zhang Y, Jin LD, Xu JJ. et al. Dynamic characterization of drug resistance and heterogeneity of the gastric cancer cell BGC823 using single-cell Raman spectroscopy. Analyst. 2017;143:164-174
44. Peixoto J, Lima J. Metabolic traits of cancer stem cells. Dis Model Mech. 2018;11:dmm033464
45. Li JJ, Condello S, Thomes-Pepin J. et al. Lipid desaturation is a metabolic marker and therapeutic target of ovarian cancer stem cells. Cell Stem Cell. 2017;20:303-314.e5
46. Zhang XN, Gillespie AL, Sessions AL. Large D/H variations in bacterial lipids reflect central metabolic pathways. Proc Natl Acad Sci USA. 2009;106:12580-12586
47. Bélanger M, Allaman I, Magistretti PJ. Brain energy metabolism: Focus on astrocyte-neuron metabolic cooperation. Cell Metab. 2011;14:724-738
48. Busch R, Kim YK, Neese RA. et al. Measurement of protein turnover rates by heavy water labeling of nonessential amino acids. Biochim Biophys Acta. 2006;1760:730-744
49. Shi LY, Zheng CG, Shen YH. et al. Optical imaging of metabolic dynamics in animals. Nat Commun. 2018;9:2995
50. Horgan CC, Nagelkerke A, Whittaker TE. et al. Molecular imaging of extracellular vesicles in vitro via Raman metabolic labelling. J Mater Chem B. 2020;8:4447-4459
51. Armstrong JPK, Stevens MM. Strategic design of extracellular vesicle drug delivery systems. Adv Drug Deliv Rev. 2018;130:12-16
52. Fu D, Holtom G, Freudiger C, Zhang X, Xie XS. Hyperspectral imaging with stimulated Raman scattering by chirped femtosecond lasers. J Phys Chem B. 2013;117:4634-4640
53. Fu D, Zhou J, Zhu WS. et al. Imaging the intracellular distribution of tyrosine kinase inhibitors in living cells with quantitative hyperspectral stimulated Raman scattering. Nat Chem. 2014;6:614-622
54. van Niel G, D'Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19:213-228
55. De Toro J, Herschlik L, Waldner C, Mongini C. Emerging roles of exosomes in normal and pathological conditions: New insights for diagnosis and therapeutic applications. Front Immunol. 2015;6:203
56. Blattner CM, Coman G, Blickenstaff NR, Maibach HI. Percutaneous absorption of water in skin: A review. Rev Environ Health. 2014;29:175-180
57. Barba C, Martí M, Semenzato A. et al. Effect of lipid modification on stratum corneum permeability. J Therm Anal Calorim. 2015;120:297-305
58. Ashtikar M, Matthäus C, Schmitt M. et al. Non-invasive depth profile imaging of the stratum corneum using confocal Raman microscopy: First insights into the method. Eur J Pharm Sci. 2013;50:601-608
59. Wang HQ, Zhang QH, Mao G. et al. Novel confocal Raman microscopy method to investigate hydration mechanisms in human skin. Skin Res Technol. 2019;25:653-661
60. Eklouh-Molinier C, Happillon T, Bouland N. et al. Investigating the relationship between changes in collagen fiber orientation during skin aging and collagen/water interactions by polarized-FTIR microimaging. Analyst. 2015;140:6260-6268
61. Zhang QH, Andrew Chan KLA, Zhang GJ. et al. Raman microspectroscopic and dynamic vapor sorption characterization of hydration in collagen and dermal tissue. Biopolymers. 2011;95:607-615
62. Liu YL, Lunter DJ. Tracking heavy-water-incorporated confocal Raman spectroscopy for evaluating the effects of pegylated emulsifiers on skin barrier. J Biophotonics. 2020;13:e202000286
63. Pattenaude SR, Streacker LM, Ben-Amotz D. Temperature and polarization dependent Raman spectra of liquid H2O and D2O. J Raman Spectrosc. 2018;49:1860-1866
64. Morawietz T, Marsalek O, Pattenaude SR. et al. The interplay of structure and dynamics in the Raman spectrum of liquid water over the full frequency and temperature range. J Phys Chem Lett. 2018;9:851-857
65. Wong WW, Cochran WJ, Klish WJ. et al. In vivo isotope-fractionation factors and the measurement of deuterium- and oxygen-18-dilution spaces from plasma, urine, saliva, respiratory water vapor, and carbon dioxide. Am J Clin Nutr. 1988;47:1-6
66. Mendez J, Prokop E, Picon-Reategui E. et al. Total body water by D2O dilution using saliva samples and gas chromatography. J appl physiol. 1970;28:354-357 &
67. Schloerb PR, Friis-hansen BJ, Edelman IS, Solomon AK, Moore FD. The measurement of total body water in the human subject by deuterium oxide dilution; with a consideration of the dynamics of deuterium distribution. J clin invest. 1950;29:1296-1310
68. Moore FD. Determination of total body water and solids with isotopes. Science. 1946;104:157-160
69. van Marken Lichtenbelt WDV, Westerterp KR, Wouters L. Deuterium dilution as a method for determining total body water: Effect of test protocol and sampling time. Br j nutr. 1994;72:491-497
70. Kim SG, Ackerman JJH. Multicompartment analysis of blood flow and tissue perfusion employing D2O as a freely diffusible tracer: A novel deuterium NMR technique demonstrated via application with murine RIF-1 tumors. Magn reson med. 1988;8:410-426
71. Wang FN, Peng SL, Lu CT, Peng HH, Yeh TC. Water signal attenuation by D2O infusion as a novel contrast mechanism for 1H perfusion MRI. Nmr biomed. 2013;26:692-698
72. Chen L, Liu J, Chu C. et al. Deuterium oxide as a contrast medium for real-time MRI-guided endovascular neurointervention. Theranostics. 2021;11:6240-6250
73. Lu M, Zhu XH, Zhang Y, Mateescu G, Chen W. Quantitative assessment of brain glucose metabolic rates using in vivo deuterium magnetic resonance spectroscopy. J Cereb Blood Flow Metab. 2017;37:3518-3530
74. De Feyter HM, Behar KL, Corbin ZA. et al. Deuterium metabolic imaging (DMI) for MRI-based 3D mapping of metabolism in vivo. Sci adv. 2018;4:eaat7314
75. Rich LJ, Bagga P, Wilson NE. et al. 1H magnetic resonance spectroscopy of 2H-to-1H exchange quantifies the dynamics of cellular metabolism in vivo. Nat biomed eng. 2020;4:335-342
76. Kasumov T, Ilchenko S, Li L. et al. Measuring protein synthesis using metabolic ²H labeling, high-resolution mass spectrometry, and an algorithm. Anal biochem. 2011;412:47-55
77. Kasumov T, Willard B, Li L. et al. 2 H 2 O-Based High-Density Lipoprotein Turnover Method for the Assessment of Dynamic High-Density Lipoprotein Function in Mice. ATVB. 2013;33:1994-2003
78. Price JC, Holmes WE, Li KW. et al. Measurement of human plasma proteome dynamics with (2)H(2)O and liquid chromatography tandem mass spectrometry. Anal biochem. 2012;420:73-83
79. Rachdaoui N, Austin L, Kramer E. et al. Measuring proteome dynamics in vivo: As easy as adding water? Mol cell proteomics. 2009;8:2653-2663
80. Herath KB, Zhong WD, Yang J. et al. Determination of low levels of H-2 labeling using high-resolution mass spectrometry: Application in studies of lipid flux and beyond. Rapid commun mass sp. 2013;28:239-244
Author contact
Corresponding author: Dr Dong Song, Department of Breast Surgery, First Hospital of Jilin University, 71 Xinmin Avenue, Changchun, Jilin 130021, P.R. China. songdongedu.cn

 Global reach, higher impact
Global reach, higher impact