Impact Factor ISSN: 1449-1907
Int J Med Sci 2022; 19(8):1340-1356. doi:10.7150/ijms.76168 This issue Cite
Review
The emerging role of miRNAs in the pathogenesis of COVID-19: Protective effects of nutraceutical polyphenolic compounds against SARS-CoV-2 infection
1. Division of Chest Medicine, Kaohsiung Municipal Min-Sheng Hospital, Kaohsiung, Taiwan, ROC
2. Division of Chest Medicine, Department of Internal Medicine, CHENG HSIN General Hospital, Taipei, Taiwan, ROC
3. Division of Chest Medicine, Department of Internal Medicine, Kaohsiung Veterans General Hospital, Kaohsiung, Taiwan, ROC
4. Department of Clinical Pathology and Medical Research, Taipei Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation, New Taipei City, Taiwan, ROC
5. Division of Nephrology, Department of Medicine, Taipei Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation, New Taipei City, Taiwan, ROC
6. Division of Nephrology, Department of Medicine, Fu-Jen Catholic University Hospital, School of Medicine, Fu-Jen Catholic University, New Taipei City, Taiwan, ROC
7. Department of Research, Taipei Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation, New Taipei City, Taiwan, ROC
#These authors equal contribution
Received 2022-6-15; Accepted 2022-7-8; Published 2022-7-18
Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection can cause immunosuppression and cytokine storm, leading to lung damage and death. The clinical efficacy of anti-SARS-CoV-2 drugs in preventing viral entry into host cells and suppressing viral replication remains inadequate. MicroRNAs (miRNAs) are crucial to the immune response to and pathogenesis of coronaviruses, such as SARS-CoV-2. However, the specific roles of miRNAs in the life cycle of SARS-CoV-2 remain unclear. miRNAs can participate in SARS-CoV-2 infection and pathogenesis through at least four possible mechanisms: 1. host cell miRNA expression interfering with SARS-CoV-2 cell entry, 2. SARS-CoV-2-derived RNA transcripts acting as competitive endogenous RNAs (ceRNAs) that may attenuate host cell miRNA expression, 3. host cell miRNA expression modulating SARS-CoV-2 replication, and 4. SARS-CoV-2-encoded miRNAs silencing the expression of host protein-coding genes. SARS-CoV-2-related miRNAs may be used as diagnostic or prognostic biomarkers for predicting outcomes among patients with SARS-CoV-2 infection. Furthermore, accumulating evidence suggests that dietary polyphenolic compounds may protect against SARS-CoV-2 infection by modulating host cell miRNA expression. These findings have major implications for the future diagnosis and treatment of COVID-19.
Introduction
Coronavirus disease 2019 (COVID-19) has emerged as a new epidemic disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection [1]. COVID-19 has spread rapidly and placed tremendous strain on global health-care systems. SARS-CoV-2 targets angiotensin-converting enzyme 2 (ACE2) receptors in human lung and gastrointestinal tissues. The receptor-binding domain (RBD) of the SARS-CoV-2 spike (S) protein binds to ACE2 on the plasma membranes of infected cells, initiating receptor-mediated endocytosis. SARS-CoV-2 variants, such as the Delta and Omicron variants, have distinct viral S (S1 or S2) proteins and RBDs, which may make these variants more infectious by enhancing their affinity for ACE2 [2, 3]. The high infectivity of these new variants has led to the acceleration of the spread of COVID-19 worldwide. Most patients with COVID-19 exhibit some typical clinical symptoms, such as fever, fatigue, cough, expectoration, sputum production, and shortness of breath. Children with COVID-19 tend to present with similar symptoms, which are usually milder than those experienced by adult patients [4, 5]. Older adult patients with COVID-19 have a higher frequency of severe illness and COVID-19-related death than do younger patients [5]. These findings suggest that comorbidities and age may be crucial factors in determining the severity of COVID-19.
Several strategies for blocking the interaction between SARS-CoV-2 and ACE2 receptors, thus preventing the spread of infection, have been developed. Soluble RBD mimetics or antibodies against ACE2 receptors bind to ACE2 receptors and prevent the virus from binding to and entering host cells. A similar effect can be achieved by directly targeting and binding to the RBDs of coronavirus S proteins by using the extracellular domain of ACE2 as bait [6]. SARS-CoV-2 infection may induce the expression of several host cell genes, which may produce antiviral or proviral effects and even help the virus evade the immune response [7]. Therefore, several anti-SARS-CoV-2 drugs, including such as favipiravir, remdesivir, lopinavir, ritonavir, chloroquine, ribavirin, and umifenovir, have been developed to prevent viral entry into host cells and suppress various steps of viral replication [8]. However, the clinical efficacy of these drugs for COVID-19 remains limited.
Numerous treatments and vaccines for COVID-19 have already been developed; however, the detailed molecular pathogenic mechanisms must be further elucidated to facilitate the development of new, more effective therapeutics against SARS-CoV-2 variants. microRNAs (miRNAs) are small noncoding RNA molecules that regulate the expression of genes posttranscriptionally. Recent studies have confirmed that host and viral miRNAs are necessary for successful SARS-CoV-2 infection. Several miRNAs can inhibit the expression of proteins involved in the SARS-CoV-2 life cycle, such as ACE2, TMPRSS2, S proteins, and Nsp12. Therefore, miRNAs acting on the viral entry pathway may serve as potential therapeutic tools against COVID-19 [9]. Naturally occurring nutraceutical polyphenolic compounds such as resveratrol have antioxidant, antitumor, antiviral, and free radical-scavenging properties; therefore, they may be used as adjunctive therapy for COVID-19 [10]. Preclinical studies have revealed that resveratrol has promising effects for the treatment of COVID-19 [11]. Polyphenolic compounds also interact with cellular signaling pathways, regulate gene and miRNA expression, affect the activity of transcription factors [12]. The use of harmless nutraceuticals to inhibit viral entry into cells and viral replication by modulating host cell entry-related miRNAs is a feasible approach to addressing the emergence of new SARS-CoV-2 variants. In this review, we present recent findings on the role of miRNAs in SARS-CoV-2 cell entry. We also describe various polyphenols that are widely used as nutraceuticals and explore the epigenetic ability of these compounds to modulate miRNA expression and to potentially prevent SARS-CoV-2 cell entry and viral replication [12].
MicroRNA biogenesis and mechanisms
MicroRNAs (miRNAs) are small RNAs with critical functions in several physiological processes [13]. Mature miRNAs are produced from primary transcripts (pri-miRNAs), which are processed by Drosha proteins into precursor miRNAs (pre-miRNAs), each of which consists of 5p and 3p arms and a terminal loop. The pre-miRNAs are transported to the cytoplasm by exportin 5 and are divided by Dicer to release the terminal loop and 5p/3p duplex. Finally, according to the hydrogen bond theory, the 5p or 3p arm is selectively loaded onto the RNA-induced silencing complex [14-16]. However, studies have reported different arm selection preferences of miRNA (for the 5p or 3p arms) in different tissues, developmental stages, and species and during cancer progression [17-25]. Researchers have hypothesized that these arm selection preferences are regulated by target-mediated miRNA protection [26, 27]. MiRNA performs its biological functions by targeting the 3′-untranslated region (3′-UTR) of protein-coding genes to degrade mRNA or inhibit protein translation. Therefore, an single pre-miRNA structure can generate two mature miRNAs, miR-#-5p and -3p, which might have distinct functions in distinct cell types or under distinct physiological conditions [28]. miRNAs play key roles in antiviral responses and viral pathogenesis in hosts infected with herpesviruses, polyomaviruses, retroviruses, pestiviruses, and hepatitis viruses as well as coronaviruses [29]. The induction of miRNA expression by viral infection boosts the immune response by altering the gene expression profiles of host cells. The miRNAs induced by these viruses can be used to identify targets involved in the viral life cycle. Therefore, several studies have developed and evaluated the use of virus-induced miRNAs in antiviral therapies, including treatments for human immunodeficiency virus 1, herpes simplex virus, dengue fever, influenza, and hepatitis C [30].
Life Cycle of SARS-CoV-2
SARS-CoV-2 is a single-stranded positive RNA virus with a large genome (approximately 29.9 kb in length), which comprises a variable number of open reading frames encoding 16 nonstructural proteins (NSPs) as well as structural proteins (including spike [S], envelope [E], membrane [M], and nucleocapsid [N] proteins) and six accessory proteins (3a, 6, 7a, 7b, 8, and 10) [22]. MiRNAs derived from host cells may have a positive or negative effect on the viral life cycle and viral replication by directly binding to genomic or transcriptional SARS-CoV-2 RNA. These SARS-CoV-2-regulated host miRNAs can affect the host's immune system by indirectly regulating the expression of key genes in the message transmission pathway [31].
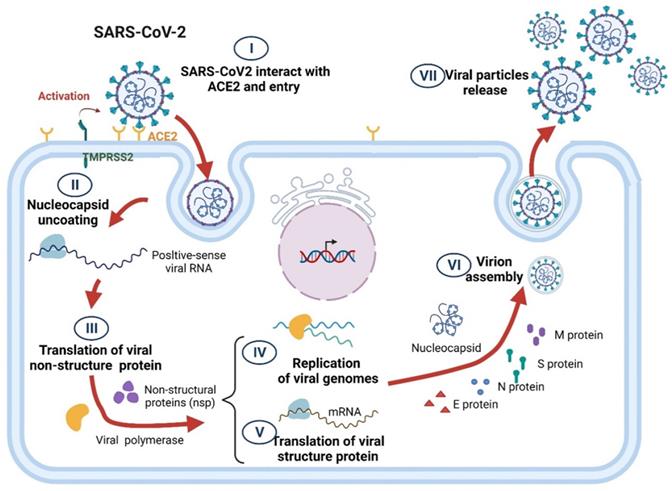
The life cycle of SARS-CoV-2 comprises several steps, including virus particle attachment, entry into host cells, genome replication, essential protein synthesis, viral component assembly, and release. In brief, transmembrane serine protease 2 (TMPRSS2) causes SARS-CoV-2 S proteins to interact with the angiotensin-converting enzyme 2 (ACE2) surface receptors of a host cell, thereby enabling the viruses to enter the host cell through endocytosis [32]. After proteolysis, the viral RNA is released, which initiates the viral genome replication and the translation of essential viral proteins using the host cell's replication and protein translation machinery. Numerous structural proteins are rapidly produced, and viral components, including the viral RNA genome and structural proteins, are packaged into SARS-CoV-2 particles and released through exocytosis to infect surrounding cells (Figure 1) [33, 34].
Schematic of SARS-CoV-2 life cycle.
MicoRNA participate in SARS-CoV-2 pathogenesis
In human cells, miRNAs can also play a role in preventing viral infection by blocking target pathways required for viral penetration, replication, and spread, including the p38 MAPK, PI3K/Akt, FAK, IFN-gamma, TGF-beta, interleukin, IGF1, and TRAIL signaling pathways [7, 35-37]. Khan et al. identified three miRNAs (hsa-miR-17-5p, hsa-miR-20b-5p, and hsa-miR-323a-5p) that exhibit antiviral effects against SARS-CoV-2 during host infection [7]. Some miRNAs, including hsa-miR-8066, hsa-miR-5197-3p, and hsa-miR-3934-3p, modulate the SARS-CoV-2 life cycle by influencing biosynthesis [38]. Sato et al. reported that hsa-miR-15b-5p interacts with RNA polymerase, which might affect SARS-CoV-2 replication [39]. A comprehensive analysis of the cell-free RNA profiles of the plasma of patients with COVID-19 and healthy controls revealed that the expression of hsa-let-7 family members, hsa-miR-23a-3p, hsa-miR-23b-3p, hsa-miR-451a, and hsa-miR-316 in patients with COVID-19 was significantly lower than that in healthy individuals. The downregulation of these miRNAs in patients with COVID-19 leads to IL-6/IL-6R hyperactivation by directly targeting the 3'UTR of IL-6/IL-6R, thereby enhancing the cytokine storm induced by SARS-CoV-2 infection [40].
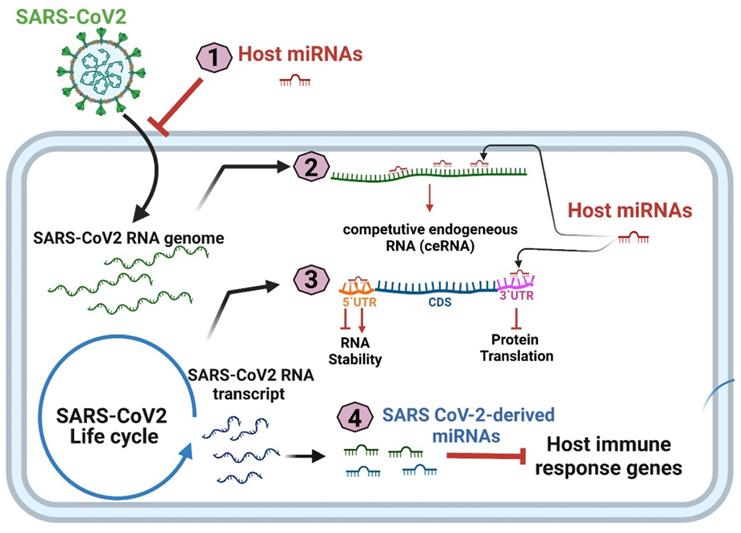
Meidert et al. reported that the differential expression of hsa-miR-3168, hsa-miR-146a-5p, and hsa-let-7e-5p in patients with COVID-19 can affect the expression of inflammation-related genes, including IL-6, IL-8, and Toll-like receptor 4 (TLR4)[41]. In addition to influencing inflammation-related pathways, host miRNAs may negatively regulate the viral life cycle by inhibiting viral entry or the translation of the viral genome; alternatively, the binding of miRNAs to the 5′ or 3′UTR may lead to greater RNA stability and increased viral replication [42]. Overall, miRNAs participate in SARS-CoV-2 infection and pathogenesis through four mechanisms: (1) host cell miRNA expression interfering with SARS-CoV-2 cell entry; (2) SARS-CoV-2-derived RNA transcripts acting as competitive endogenous RNAs (ceRNAs) that may attenuate host cell miRNA expression; (3) host cell miRNA expression modulating SARS-CoV-2 replication, and (4) SARS-CoV-2-encoded miRNAs protecting SARS-CoV-2 from degradation and silencing the expression of host protein-coding genes (Figure 2).
Four putative mechanisms of miRNA participation in SARS-CoV-2 pathogenesis.
Host cell miRNA expression interferes with SARS-CoV-2 cell entry
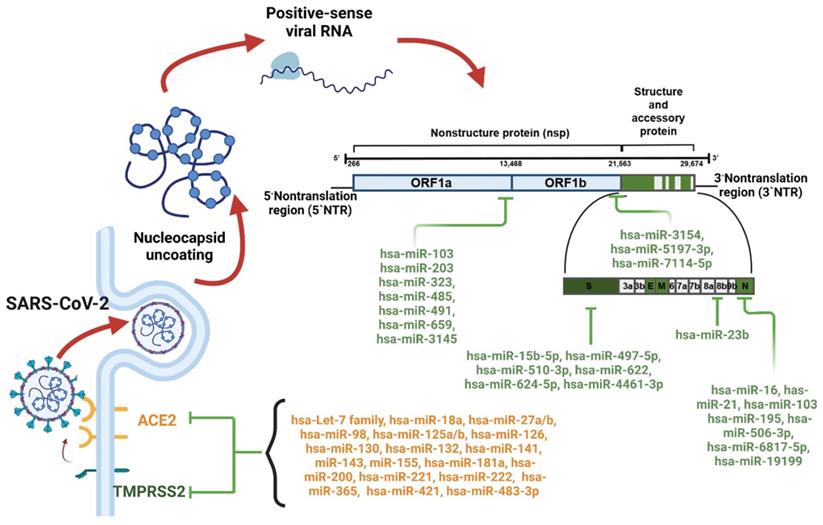
Numerous human miRNAs have binding sites across the 3′UTRs of ACE2 and TMPRSS2 and therefore exhibit antiviral effects (Figure 3 and Table 1). Hsa-miR-200c inhibits ACE2 by targeting ACE2 mRNA in primary cardiomyocytes, which suggests that hsa-miR-200c could reduce the risk of SARS-CoV-2 infection [43]. In addition, hsa-let-7e and hsa-miR-98-5p could suppress TMPRSS2 gene expression by directly targeting the 3'UTR of TMPRSS2 [44, 45]. Nersisyan et al. reported that hsa-miR-125a, hsa-miR-141, and hsa-miR-200 family members could regulate ACE2 expression by directly targeting the 3'-UTR of ACE2 [45]. A similar study revealed that the host miRNAs hsa-miR-9-5p and hsa-miR-218-5p could target the 3'-UTR of ACE2 and that hsa-Let-7d-5p, hsa-Let-7e-5p, hsa-miR-494-3p, hsa-miR-382-3p, and hsa-miR-181c-5p could target the 3'-UTR of TMPRSS2 [46]. Hsa-miR-18 and hsa-miR-125b play central roles in acute renal injury in patients with SARS-CoV-2 infections by directly binding to ACE2. Furthermore, anti-hsa-miR-18 and anti-hsa-miR-125b are effective against SARS-CoV-2 infection [47].
Human miRNAs regulating SARS-CoV-2 infection and replication.
Human miRNAs directly bind to the RNA transcripts of SARS-CoV-2.
| SARS-VoV2 | miRNAs | Reference |
|---|---|---|
| S protein | hsa-miR-510-3p, hsa-miR-624-5p, hsa-miR-497-5p, hsa-miR-622, hsa-miR-761, hsa-miR-15b-5p, hsa-miR-196a-5p, hsa-miR-565, hsa-miR-151a-5p, hsa-miR-380-5p, hsa-miR-449a, hsa-miR-4464, hsa-miR1234-3p, hsa-miR-7107-5p, hsa-885-5p | [53-55] |
| Replication related RNA | hsa-miR-1307-3p | [67] |
| 3' untranslated region | hsa-miR-3613-5p, hsa-miR-8066 | [79] |
| NSPs/ORF1a/b | hsa-miR-203b-3p, hsa-miR-103a-1-5p, hsa-miR-6818-5p, hsa-miR-624-5p, hsa-miR-378c, hsa-miR-3202, hsa-miR-5591-5p, hsa-miR-8082, hsa-miR-939-5p, hsa-miR-549a-3p and hsa-miR-6515-5p | [7, 58] |
| ORF8 | hsa-miR-12129, hsa-miR-2392, hsa-miR-23b-5p and hsa-miR-5047 | [7] |
| Nucleocapsid | hsa-miR-21-3p, hsa-miR-195-5p, hsa-miR-16-5p, hsa-miR-3065-5p, hsa-miR-424-5p, hsa-miR-421, hsa-miR-6817-5p, hsa-miR-506-3p and hsa-miR-12119 | [7, 56] |
SARS-CoV-2-derived RNA transcripts act as competitive endogenous RNAs (ceRNAs) that may attenuate host cell miRNA expression
RNA transcripts perform crucial biological functions by interacting with and titrating the expression of endogenous miRNAs [48, 49]. Therefore, both the RNA transcripts and genomic RNA of SARS-CoV-2 can act as ceRNA to regulate the activity of endogenous miRNAs. Host-pathogen interactions via host cellular components play a major role in viral infection. Viral miRNAs have been identified as key players in host-virus interactions. In addition to protein-coding mRNAs, noncoding RNAs may be targeted in infected cells, and viruses can exploit the host miRNA network through the ceRNA effect [50]. ceRNAs are transcripts that can regulate each other posttranscriptionally by competing for shared miRNAs. Because any transcripts harboring miRNA bonding sites can theoretically function as ceRNAs; they represent a widespread form of posttranscriptional regulation of gene expression in both physiology and pathology [51]. Accumulating evidence indicates that ceRNA networks link the functions of protein-coding mRNAs with those of noncoding RNAs such as miRNAs, long noncoding RNAs [52], pseudogenic RNAs, and circular RNAs [51], thereby affecting and regulating the expression of target genes. Because host miRNAs can bind to the coding DNA sequence (CDS) regions of viral RNAs, even without interfering with viral RNA function, overconsumption of host miRNAs (known as the sponge effect) may lead to a reduction in the availability of such miRNAs, thus increasing the severity of COVID-19 infection. Using target prediction tools, numerous studies have identified putative antiviral host miRNAs targeting virus genes. Haddad et al. identified 10 miRNAs (hsa-miR-4288, hsa-miR-195-5p, hsa-miR-16-5p, hsa-miR-15b-5p, he-miR-15a-5p, hsa-miR-6838-5p, hsa-miR-497-5p, hsa-miR-424-5p, hsa-miR-3133, and hsa-miR-21-3p) that can bind to the single-stranded RNA of the full-length SARS-CoV-2 genome. Among these miRNAs, hsa-miR-510-3p, hsa-miR-624-5p, and hsa-miR-497-5p exhibit high potential to target the mRNA encoding the S glycoprotein of SARS-CoV-2 [53]. Hsa-miR-15b-5p, hsa-miR-151a-5p, hsa-miR-196a-5p, hsa-miR-380-5p, hsa-miR-449a, hsa-miR-565, hsa-miR-622, and hsa-miR-761 target the S protein gene [54]. Four human miRNAs (hsa-miR-4464, hsa-miR1234-3p, hsa-miR-7107-5p, and hsa-885-5p) exhibit perfect complementarity with the RBD of the S gene of SARS-CoV-2 [55]. Nersisyan et al. performed computational prediction to identify critical miRNAs that interact with human coronaviruses and identified six miRNAs (hsa-miR-21-3p, hsa-miR-195-5p, hsa-miR-16-5p, hsa-miR-3065-5p, hsa-miR-424-5p, and hsa-miR-421) that exhibited high binding probability across all the analyzed viruses [56]. Another study, which used 67 different SARS-CoV-2 genomes to identify conserved miRNA binding sites, revealed that 10 miRNAs (hsa-miR-103a-1-5p, hsa-miR-6818-5p, hsa-miR-624-5p, hsa-miR-378c, hsa-miR-3202, hsa-miR-5591-5p, hsa-miR-8082, hsa-miR-939-5p, hsa-miR-549a-3p, and hsa-miR-6515-5p) can bind to the ORF1a/b region of the SARS-CoV-2 genome [7].
Using the bioinformatics approach, Chow et al. identified 128 human miRNAs with the potential to bind to the genomic RNA of SARS-CoV-2 [38]. Among them, the expression levels of six miRNAs differed significantly between lung cells infected with SARS-CoV-2 and healthy lung cells: the expression levels of hsa-Let-7a-3p, miR-16-2-3p, hsa-miR-135-5p, and miR-1275 were significantly downregulated in the SARS-CoV-2-infected lung cells, whereas those of hsa-miR-139-5p and hsa-miR-155-3p were significantly upregulated [38]. Using various target prediction tools, Fulzele et al. identified 873 human miRNAs targeting the SARS-CoV-2 genome [57]. Pathway enrichment analysis revealed that these miRNAs are involved in various age-related signaling pathways. The authors' findings indicate that older adult patients with SARS-CoV-2 infections tend to have greater disease severity and higher mortality rates than do younger patients because of low miRNA expression, which is directly suppressed through sponging. Another team identified 479 human miRNAs that could bind to the SARS-CoV-2 genome. Among them, 369 targeted an ORF1a/b sequence [58]. Taken together, the results of these three studies indicate that 11 human miRNAs (hsa-miR-15b-3p, hsa-miR-19b-1-5p, hsa-miR-19b-2-5p, hsa-miR-125a-3p, hsa-miR-141-3p, hsa-miR-153-5p, hsa-miR-196a-5p, hsa-miR-1202, hsa-miR-1301-3p, hsa-miR-4758-5p, and hsa-miR-5047) may target sequences of SARS-CoV-2.
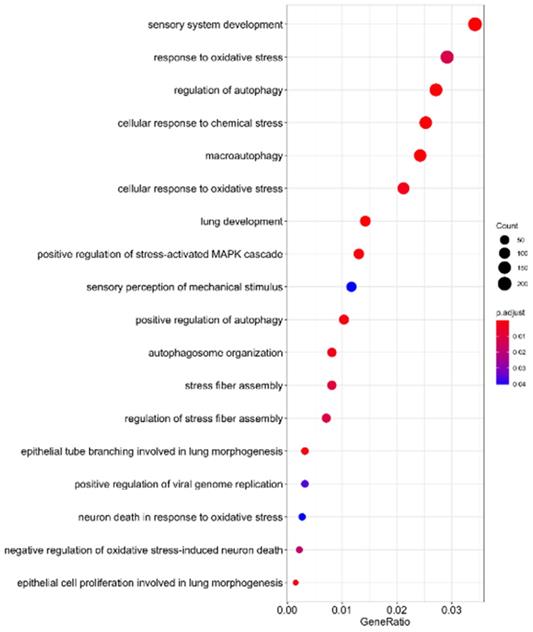
As indicated in Table 1, 41 microRNAs have been determined to target sequences of SARS-CoV-2, suggesting that these human miRNAs might be sponged during SARS-CoV-2 replication. To elucidate the functions of these sponged host miRNAs, we identified their putative target genes and conducted pathway enrichment analysis. As indicated in Figure 4, these miRNAs are involved in sensory system development, oxidative responses, autophagy, lung development, stress-activated MAPK signaling, and oxidative stress-induced neuron death. A previous study reported that patients with SARS-CoV-2 infection have excessive reactive oxygen species (ROS) levels, which facilitates the cascade of biological events that drive pathological host responses [59]. High ROS levels induce tissue damage, thrombosis, and red blood cell dysfunction, which contribute to the severity of COVID-19. Macroautophagy and autophagy are crucial to viral replication and maturation [60]. According to RNA sequencing data of SARS-CoV-2-infected cells, MAPK signaling is associated with lung fibrosis, a lethal complication of COVID-19 [61]. These findings demonstrate that SARS-CoV-2 RNA sponging host endogenous miRNAs is crucial to COVID-19 progression.
SARS-CoV-2 RNA sponging of miRNAs involved in human signaling transduction pathways.
(3) Host cell miRNA expression modulates SARS-CoV-2 replication
In addition to affecting viral entry into host cells, many host miRNAs also target and suppress the replication, translation, and protein synthesis-related gene expression of SARS-CoV-2. hsa-miRNA can bind to the SARS-CoV-2 genome. Host miRNAs can facilitate viral replication by targeting the viral 5′ noncoding region or by controlling (repressing) the translation of viral mRNA into proteins by targeting the viral 3′ noncoding region, thus enabling the virus to evade the host immune system [62, 63]. Briefly, miRNA binding to 5′ UTR leads to RNA stability and increased viral replication. miRNA binding to 3′ UTR can lead to the inhibition of viral translation or increased RNA stability and viral translation. The SARS-CoV-2 gene ORF6 and CDSs NSP13, NSP14, and NSP15 can powerfully suppress both primary interferon production and interferon signaling. Among the 27 known viral proteins of SARS-CoV-2, ORF6 most strongly suppresses both primary interferon production and interferon signaling [64]. SARS-CoV-2 ORF6 can also act as a virulence factor by regulating nucleocytoplasmic trafficking to accelerate viral replication, resulting in rapid disease progression [65]. Certain human miRNAs (including miRNA-323 and miRNA-485) target ORF1a/b, which encodes enzymes necessary for the replication and translation of SARS-CoV-2 [66]. Many human miRNAs modulate the life cycle of SARS-CoV-2 by directly targeting the RNA encoding NSPs or structural proteins (Figure 3 and Table 1). Chen et al. identified two miRNAs (hsa-miR-1307-3p and hsa-miR-3613-5p) that could prevent viral replication by targeting the 3'-UTRs of replication-related SARS-CoV-2 RNA [67]. Hsa-miR-203b-3p can suppress viral replication by targeting the sequence of ORF1ab and ORF3a [58]. The high abundance of hsa-miR-497-5p, hsa-miR-21-3p, and hsa-miR-195-5p were predicted targeted at SARS-CoV-2 genome, that degrade the RNA of coding-region genes of SARS-CoV-2, thereby suppressing SARS-CoV-2 replication [53]. By acting as host miRNA sponges, noncoding SARS-CoV-2 RNAs may dysregulate and suppress the expression of human miRNAs, including hsa-miR-10a-5p, hsa-miR-99b-5p, hsa-miR-376a-3p, and hsa-miR-548a-5p, which are involved in immune responses [68]. Bartoszewski et al. reported that hsa-miR-34a/c-5p, hsa-miR-92a-5p, hsa-miR-138-5p, hsa-miR-449c-5p, hsa-miR-766-5p, hsa-miR-3940-5p, and hsa-miR-6741-5p can bind to multiple sites on noncoding SARS-CoV-2 RNA, which can affect the host immune response [68]. Circulating hsa-miR-150-5p plays a crucial role in inhibiting SARS-CoV-2 infection by directly interacting with the nsp10 region of SARS-CoV-2 RNA [69].
(4) SARS-CoV-2-encoded miRNAs protect SARS-CoV-2 from degradation and silencing the expression of host protein-coding genes
Studies have identified multiple DNA and RNA viruses that produce pre-miRNA sequences that may undergo further maturation induced by the human RNA-induced silencing complex [70, 71]. Viral miRNAs have been identified in many human viruses, including influenza [72], EV71 [73], hepatitis A [74], and SARS-CoV [75, 76]. The structure and function of these viral miRNAs are highly similar to those of human miRNA. It is well known that the transcription and translation mechanisms of the host cell are necessary for viral replication, translation, and protein synthesis [77, 78]. Viral miRNAs protect SARS-CoV-2 mRNA from turnover and degradation in human cells by inhibiting certain host mRNA deadenylases. Viral miRNAs also target transcriptional regulators to prevent RNA polymerase II from attaching to the promoters of host genes [58]. Morales et al. used deep RNA sequencing to profile the expression of small RNAs in the lungs of SARS-CoV-2 infected mice. They identified three SARS-CoV-derived miRNAs, which were derived from the nsp3 (svRNA-nsp3.1 and -nsp3.2) and N (svRNA-N) genomic regions of SARS-CoV [75]. Using antagomir to block SARS-CoV-derived miRNAs could significantly reduce in vivo lung pathology and proinflammatory cytokine expression, which indicates that virus-derived miRNA may be a therapeutic target in patients with SARS-CoV infections. Arisan et al. identified seven key-microRNAs (miR-8066, miR-5197, miR-3611, miR-3934-3p, miR-1307-3p, miR-3691-3p, and miR-1468-5p) with sequences similar to those of human miRNA and SARS-CoV-2 [79]. These SARS-CoV-2-derived miRNAs could completely complement the target RNA transcripts of SARS-CoV-2 to prevent viral replication and protein translation. Recently, Fu et al. identified a SARS-CoV-2-derived miRNA, miR-nsp3-3p, that was encoded at nucleotides 3874-2894 on the 3'-UTRs of nsp3 genes [80]. Furthermore, the expression levels of miR-nsp3-3p can effectively predict critical illness risk and outcomes (as related to disease progression and recovery) in patients with COVID-19. Aydemir et al. used Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis and gene regulatory networks to identify 40 SARS-CoV-2-encoded miRNAs and their regulatory targets, some of which play key roles in the NFκB, JAK/STAT, and TGFβ signaling pathways and cellular epigenetic regulation pathways [81]. One study identified 27 SARS-CoV-2-encoded miRNAs that bind to regions of the SARS-CoV-2 genome. Most of the target sites of these miRNAs are located on the ORF1ab gene, and some sites are located in the 5'-UTR of the viral genome and the S gene. Virus-encoded miRNAs that bind to genomic regions can regulate viral replication and host entry [66]. Conversely, SARS-CoV-2-encoded miRNAs can also act on the host genome. miR-147-3p, encoded by SARS-CoV-2, enhances the expression of TMPRSS2 in the gut and increases the virus's ability to spread [55], and miR-5197, miR-8066, and miR-3934-3p are involved in the N-linked and O-linked glycosylation of subunit S1 and S2 proteins, which can increase the pathogenicity of SARS-CoV-2 [79]. Although most miRNAs have conserved sequences, some mutations were detected in miR-1307-3p and miR-1468-5p of SARS-CoV-2 strains, which may be involved in their pathogenicity [82]. However, targeting miRNAs that are highly conserved in all SARS-CoV-2 strains appears to be an effective preventive approach and a promising therapeutic strategy.
Circulating miRNAs act as biomarkers for COVID-19
Several circulating miRNAs have been detected in sera, plasma, urine, tears, amniotic fluid, and gastric juice [83-85]. These circulating miRNAs are highly stable in body fluids and might originate from different cell types under different physiological conditions [84, 86]. Therefore, miRNAs might be useful noninvasive biomarkers for the diagnosis and prognosis of patients with COVID-19 (Table 2).
Circulating miRNAs identified as biomarkers for diagnosis and prognosis of patients with COVID-19
| microRNA | Clinical samples | References | |
|---|---|---|---|
| Diagnosis markers | hsa-miR-10b, hsa-miR-21, hsa-miR-155, hsa-miR-208a, hsa-miR-499, hsa-miR-29a-3p, hsa-miR-146a, hsa-miR-155-5p, hsa-miR-2392, hsa-miR-155, hsa-miR-19a-3p, hsa-miR-19b-3p, hsa-miR-92a-3p, hsa-miR-150-5p, hsa-miR-375, hsa-miR-122-5p, hsa-miR-494-3p, hsa-miR-3197, hsa-miR4690-5p, hsa-miR-1915-3p and hsa-miR-3652 | COVID-19 v.s healthy control | [69, 89, 91-93] |
| hsa-miR-106b-5p, hsa-miR-221-3p, hsa-miR25-3p, hsa-miR-30a-5p | COVID-19 v.s Community-acquired pneumonias | [88] | |
| hsa-miR-155, hsa-miR-208a, hsa-miR-499 | COVID-19 v.s Influenza-ARDS | [90] | |
| Prognosis markers | miR-nsp3-3p, hsa-miR-320a/b/c, hsa-miR-200c, hsa-miR-155 | Mild/moderate patients v.s severe patients | [80, 89, 96, 97] |
| hsa-miR-29a-3p, hsa-miR-146a-3p | Acute phase v.s post-acute pahase | [91] | |
| hsa-miR-146a-5p | Drug response | [98] | |
| hsa-miR-148a-3p, hsa-miR-451a, hsa-miR-486-5p, hsa-miR-2392 | ICU v.s ward patients | [92, 99] | |
| hsa-miR-192-5p, hsa-miR323a-3p | ICU survivors v.s non-survivors | [99] | |
| hsa-miR-320b, hsa-miR483-5p | Survivors v.s non-survivors | [100] |
Diagnostic biomarkers
Early diagnosis of COVID-19 and immediate isolation of patients with COVID-19 are effective approaches to preventing the spread of SARS-CoV-2. In addition, early detection of disease deterioration, timely intervention, respiratory support, and nutritional support can effectively reduce the risk of patient mortality. Therefore, identifying potential risk factors for and predicting the course of COVID-19 are crucial for health-care professionals. In general, the nasopharyngeal swab polymerase chain reaction test is the most common tool for COVID-19 diagnosis. However, because active SARS-CoV-2 viruses may be present in the specimens collected from the nasopharyngeal cavity, the collector and examiner might be infected during the testing process. The detection of circulating miRNAs in blood samples would be a safer alternative for COVID-19 diagnosis.
Hsa-miR-10b expression is downregulated in the peripheral blood of patients with COVID-19. Furthermore, hsa-miR-10b expression is significantly and negatively correlated with serum IL-2 and IL-8 in the blood of patients with COVID-19, which suggests that hsa-miR-10b may contribute to cytokine storm [87]. In a study comparing the differential miRNA serum profiles patients with COVID-19 and community-acquired pneumonia (CAP), the researched identified four miRNAs (hsa-miR-25-3p, hsa-miR-30a-5p, hsa-miR-106b-5p, and hsa-miR-221-3p) for inclusion in a panel that could be used to significantly discriminate between patients with COVID-19 and CAP [88]. Another study demonstrated that the expression of hsa-miR-21, hsa-miR-155, hsa-miR-208a, and hsa-miR-499 in the serum or plasma of patients with COVID-19 was significantly higher than that in the serum of plasma of healthy individuals [89]. Receiver operating characteristic (ROC) analysis revealed that hsa-miR-155, hsa-miR-208a, and hsa-miR-499 serum levels could be used to distinguish between COVID-19 and influenza-induced acute respiratory distress syndrome [90]. Donyavi et al. conducted an additional ROC analysis and determined that hsa-miR-29a-3p, hsa-miR-146a, and hsa-miR-155-5p expression levels could serve as biomarkers for COVID-19 diagnosis with high specificity and sensitivity [91]. McDonald et al. observed that hsa-miR-2392 expression was significantly upregulated in the serum and urine of patients with COVID-19 [92]. By comprehensively analyzing miRNA profiles, Akula et al. reported that hsa-miR-150-5p, hsa-miR-375, hsa-miR-122-5p, and hsa-miR-494-3p expression levels were significantly upregulated in the plasma of patients with COVID-19, whereas those of hsa-miR-3197, hsa-miR-4690-5p, hsa-miR-1915-3p, and hsa-miR-3652 were significantly downregulated [69]. Fayyad-Kazan et al. identified eight miRNAs (hsa-miR-15a-5p, hsa-miR-17-5p, hsa-miR-19a-3p, hsa-miR-19b-3p, hsa-miR-23a-3p, hsa-miR-92a-3p, hsa-miR-142-5p, and hsa-miR-320a) differentially expressed in patients with SARS-CoV-2 and healthy controls. Among them, hsa-miR-19a-3p, hsa-miR-19b-3p, and hsa-miR-92a-3p were combined into a diagnostic panel, which may serve as a diagnostic biomarker with an AUC of 0.917 (p =0.0001) [93].
Prognostic biomarkers
The use of prognostic biomarkers to monitor clinical progress and classify patients can enable health-care providers to provide more precise personalized treatment [94]. The reason why some patients with COVID-19 are more likely to experience severe disease symptoms remains unknown. Usually, patients with severe COVID-19 have a high viral load, high oxidative stress, and high levels of inflammatory cytokines [95]. COVID-19 can be classified as mild, moderate, severe, or critical. SARS-CoV-2 infection leads to rapid innate immunity activation; therefore, some levels of hematological parameters (white blood cells, lymphopenia, C-reactive protein (CRP), lactate dehydrogenase, and creatine kinase), proinflammatory cytokines (IL-1B, IL-6, IL-8, and G-CSF), and chemokines (MCP1, IP10, and MIP1a) are significantly elevated in the blood of patients with COVID-19 [9,10]. Elevated levels of these markers are more common among patients with severe COVID-19 than among those with mild COVID-19 and warrant inclusion in risk stratification models.
Using next generation sequencing to profile the small RNA in serum of patients with COVID-19 and heathy controls, researchers identified SARS-CoV-2-derived miR-nsp3-3p as a potential biomarker for the prediction of a patient's risk of severe disease [80]. Similar studies showed that hsa-miR-320 family genes, hsa-miR-200c, and hsa-miR-155 are differentially expressed in patients with COVID-19 and are significant correlated with certain clinicopathological characteristics, including CRP, IL-6, and D-dimer levels [89, 96, 97]. Hsa-miR-2392 is key to multiple downstream signaling pathways, including those related to inflammation, glycolysis, and hypoxia. High hsa-miR-2392 expression is strongly associated with poor outcomes among patients with COVID-19. Furthermore, anti-hsa-miR-2392 was reported to reduce SARS-CoV-2 viability in both in vivo and in vitro models [92].
Studies have reported that the expression of certain circulating miRNAs in the blood can be used as a biomarker for predicting a patient's response to drug treatment. Sabbatinelli et al. observed low hsa-miR-146a-5p expression in the serum of patients with COVID-19 who did not respond to tocilizumab treatment and determined that low circulating hsa-miR-146a expression in serum was strongly associated with adverse outcomes among patients with COVID-19 [98]. Another group reported that both hsa-miR-29a-3p and hsa-miR-146a-3p expression could be used to precisely distinguish between acute and post-acute COVID-19 [91]. A panel of three circulating miRNAs (hsa-miR-148a-3p, hsa-miR-451a and hsa-miR-486-5p) were identified as prognostic biomarkers that could be used to distinguish between patients with COVID-19 admitted to the ICU and those admitted to a clinical ward [99]. In addition, the expression of hsa-miR-192-5p and hsa-miR-323a-3p exhibit discretionary potential to predict patient survival [99]. Two circulating serum miRNAs, hsa-miR-320b, and hsa-miR-483-5p, were also identified as biomarkers for predicting survival outcomes among patients with COVID-19 [100].
Natural compounds can block SARS-CoV-2 infection by regulating miRNA expression
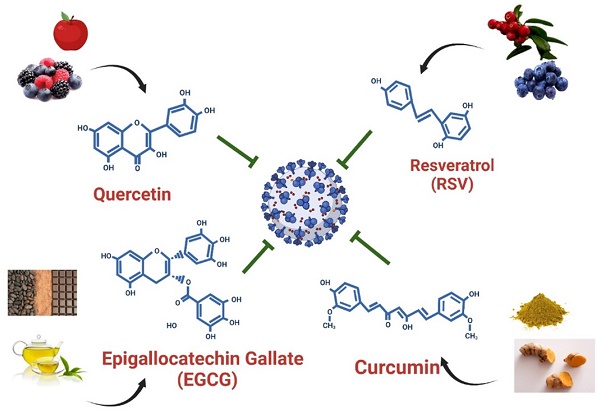
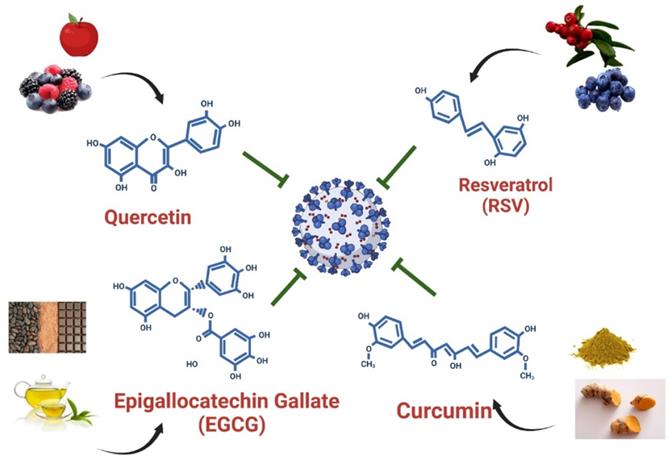
Polyphenols are the most abundant dietary antioxidants and are commonly found in fruits, vegetables, chocolate and wine [12, 101, 102]. Epidemiological studies have revealed that dietary polyphenol intake can alleviate the symptoms of some chronic diseases, including type 2 diabetes, cardiovascular disease, and COVID-19 [103]. The progression of COVID-19 is often associated with a high degree of inflammation and overproduction of ROS, resulting in poor outcomes [12, 104]. Natural polyphenols are powerful antioxidants and protect against chronic diseases by altering cell signaling pathways and regulating the expression of antioxidant-related genes, indicating that dietary polyphenolic compounds have potential benefits for the prevention of SARS-CoV-2 infection. A limited number of studies have revealed that dietary polyphenolic compounds may act directly against coronaviruses in vitro by modulating expression of host miRNAs [105, 106]. In a recent review, authors identified 125 polyphenols-modulating host miRNAs and 644 miRNAs that can interact with SARS-CoV-2 genome by performing literature search [106]. Comparison of two groups of miRNA candidates revealed 17 host miRNAs with both capacity to interact with SARS-CoV-2 genome and which expression can be regulate by polyphenols, including hsa-let-7a-3p, has-miR-25-5p, hsa-miR-1246, hsa-miR-125a-5p, hsa-miR-1262, hsa-miR-1290, hsa-miR-148a-5p, hsa-miR-154-3p, hsa-miR-21-3p, hsa-miR-320c, hsa-miR-335-5p, has-miR-34a-5p, hsa-miR-377-5p, hsa-miR-455-3p, has-miR-499a-5p, hsa-miR-544a and hsa-miR-744-3p [106]. These polyphenols-induced miRNAs suggested that exerted anti-SARS-CoV-2 capacity by influencing processes of viral replication or entry. In this review, we identified four polyphenols—quercetin, epigallocatechin gallate (ECGC), curcumin, and resveratrol—that have been frequently used as therapeutic nutritional supplements or to prevent SARS-CoV-2 infection (Figure 5). We further searched literatures to demonstrate that four polyphenolic compounds may protect against SARS-CoV-2 infection by modulating host cell miRNA expression.
Dietary compounds may protect against SARS-CoV-2 infection. Foods containing quercetin, EGCG, curcumin, and resveratrol and the structures of these polyphenols are illustrated.
Quercetin is the main representative of the flavonoid subclass of flavanols and is mainly present in fruits such as apples, berries, and grapes (Figure 5). Quercetin is most often present in food as quercetin-3-glucoside (isoquercetin) and is hydrolyzed in the small intestine and rapidly absorbed [107, 108]. It exerts antiviral effects by interacting with viral proteins to suppress viral infection, thereby blocking viral replication and entry into host cells [109]. Furthermore, quercetin has been determined to be safe and effective in reducing the serum concentrations of ALP, q-CRP, and LDH, and recently, patients with COVID-19 have been treated with daily quercetin supplements in addition to antiviral drugs [110]. One study demonstrated that quercetin can effectively inhibit SARS-CoV-2 Mpro activity in HuH-7 cells in vitro, with IC50 values ranging from 0.125 to 12.9 μM (Table 3) [111]. Mangiavacchi et al. reported that two quercetin compounds, quercetin (1) and quercetin (2d), could block SARS-CoV-2 replication in infected cells at nontoxic concentrations, with IC50 values of 192 and 8 µM, respectively (Table 3) [112]. Accumulating studies have revealed that quercetin may act as a therapeutic agent to suppress human cancer cell growth by regulating miRNA expression [113-115]. Following treatment with quercetin, the expression of let-7 family miRNAs significantly increased in patients with pancreatic ductal adenocarcinoma [113]. Upregulation of let-7 family miRNAs can inhibit K-Ras gene activity, thereby affecting cancer cell proliferation and migration. In addition, quercetin-induced miR-200b-3p converts symmetric cell division to asymmetric cell division by reversing the Notch/Numb ratio; inhibiting self-renewal; and activating the potential of cells to differentiate into adipocytes, osteocytes, and chondrocytes [115]. Furthermore, quercetin induces miR-16 expression in human cancer. High miR-16 expression can inhibit claudin-2, a mediator of intestinal barrier leakage during intestinal inflammation [116]. Among these quercetin-regulated microRNAs, let-7 family miRNA and hsa-miR-200b target TMPRSS2 and ACE-2, respectively [43, 45]. Furthermore, has-miR-16 expression suppresses the formation of structural proteins by directly targeting nucleocapsids [56]. By modulating miRNA expression, quercetin may prevent SARS-CoV-2 cell entry and replication. Overall, the daily dietary intake of quercetin can prevent SARS-CoV-2 infection and slow COVID-19 progression by modulating let-7 family miRNA, hsa-miR-200 family miRNA, and hsa-miR-16 expression (Figure 6).
In vitro studies investigating the effects of polyphenols against SARS-CoV-2 infection
| Polyphenols | Cell Model | SARS-CoV-2 strain | Concentration for used | References |
|---|---|---|---|---|
| Quercetin | Huh-7 | Human coronavirus 229E | 2.5uM~50uM | [111] |
| Vero E6 cells | SARS-CoV-2 strain 026V-03883 | Compound 1: 200uM~500uM; Compound 2d: 10uM~100uM | [112] | |
| EGCG | Huh-7 | Human coronavirus 229E | 2.5uM~50M | [111] |
| HEK293T-hACE2 and Vero E6 cells | SARS-CoV strain Frankfurt-1 | 1.25ug/ml ~25ug/ml | [118] | |
| Resveratrol | Vero E6 cells, Calu-3 cells and primary human bronchial epithel (PBECs) | SARS-CoV-2 strain NL/2020 | 15uM~150uM | [126] |
| Vero E6 cells | SARS-CoV-2 (BetaCoV/Shenzhen/SZTH-003/2020 strain) | 1.56uM~200uM | [127] | |
| Curcumin | hACE2/A549 | eGFP-luciferase-SARS-CoV-2 pseudo-typed particles | 2.5~100 ug/ml | [138] |
| Vero E6 cells, | SARS-CoV-2 D614G strain and Delta variant | 1.25~10 ug/ml | [136] | |
| Vero E6 cells and Calu-3 cells | SARS-CoV-2 isolated form hospital | 1ug/ml~125ug/ml | [137] |
Overview of quercetin-, EGCG-, curcumin-, and resveratrol-mediated inhibition of SARS-CoV-2 entry into host cells and disruption of SARS-CoV-2 life cycle progression through upregulation of miRNA expression.
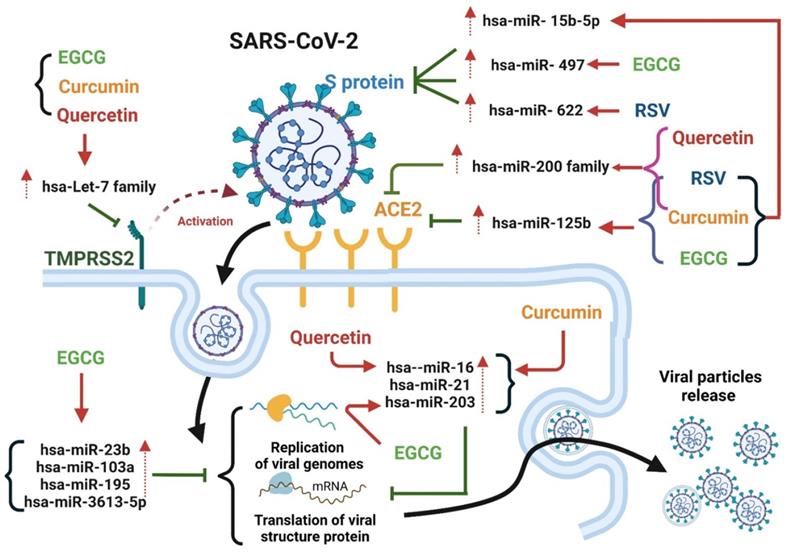
Epigallocatechin-3-gallate (EGCG) is a flavanol consisting of a catechin conjugated with gallic acid. EGCG is abundant in green tea and cocoa-based products [117]. In vivo and in vitro studies have reported the inhibitory effects of EGCG on various viruses, including SARS-CoV-2. Henss et al. reported that EGCG exhibit anti-SARS-CoV-2 activity by inhibiting the interaction between S proteins and ACE2 [118]. Furthermore, molecular docking revealed that EGCG has a high binding affinity with the SARS-CoV-2 S protein [119]. EGCG effectively inhibited SARS-CoV-2 Mpro activity and HCoV-229E replication in Huh-7 cells at a dose of 2.5 μM [111]. Henss et al. reported that EGCG effectively blocked the entry of SARS-CoV-2 into Vero E6 cells infected with the SARS-CoV strain Frankfurt-1, with an IC50 of 1.72 µg ml-1 (Table 3) [118]. Overall, EGCG might be suitable for use as a lead structure to develop more effective anti-COVID-19 drugs. Following treatment with quercetin and EGCG, hsa-let-7 family miRNA expression was specifically induced in lung cancer [113]. In another study, when EGCG was used in the IL-1β-stimulated human osteoarthritis chondrocyte cell line, it modulated the expression of various miRNAs, including let-7 family miRNAs, hsa-miR-125b-5p, and hsa-miR-497 [120]. Li et al. reported that EGCG mitigated Aβ-induced neurotoxicity by inducing miR-34a-5p and miR-125b-5p expression [121]. In addition, EGCG acted as an anti-cervical cancer agent by suppressing cervical carcinoma cell growth by upregulating hsa-miR-125b and hsa-miR-203 expression, indicating that EGCG has therapeutic potential for use in the prevention of cervical cancer [122]. Moreover. EGCG upregulated the expression of hsa-miRNA-15b in both murine and human T cells [123]. Shin et al. reported that EGCG protected against the effects of dihydrotestosterone-induced apoptosis and reduced intracellular ROS levels by altering the miRNA expression profile of human dermal papilla cells [124], wherein the expression of hsa-miR-3613-3p was significantly upregulated following treatment with EGCG. hsa-miR-3613 was predicted to bind to the 3′-UTR region of the SARS-CoV-2 genome, suggesting that EGCG prevents viral replication by targeting the sequences of replication-related SARS-CoV-2 RNAs [67]. hsa-let-7 family of miRNAs can suppress TMPRSS2 expression by binding to the 3′-UTR region of TMPRSS2. In addition, hsa-miR-125b-5p can suppress ACE2 expression by targeting 3′-UTR of ACE2, thus decreasing the likelihood of SARS-CoV-2 cell entry. In addition, hsa-miR-497-5p targets the RNA sequence of the SARS-CoV-2 S protein and may therefore inhibit the production of the S protein during the SARS-CoV-2 life cycle, resulting in the reduced yield and assembly of viral proteins. These findings suggest that EGEG interacts with SARS-CoV-2 and host cells by regulating let-7-family miRNAs, hsa-miR-15b, and hsa-miR-125b to inhibit viral entry into cells and regulating hsa-miR-497-5p expression, thus restricting production of the S protein (Figure 6).
Resveratrol (3,40,5-trihydroxystilbene) belongs to the stilbene class and thus exists in both cis and trans forms. It is abundant in grapes, grape juice, and red wine [125]. Similar to EGCG, resveratrol exerts inhibitory effects on various viruses including SARS-CoV-2. At a dose of 66 µM, resveratrol exerted a dose-dependent antiviral effect on SARS-CoV-2 in Vero E6 cells and reduced virus particle production by 50% [126]. In another study, resveratrol significantly suppressed SARS-CoV-2 replication in Vero E6 cells infected with SARS-CoV-2 (BetaCoV/Shenzhen/SZTH-003/2020 strain; Table 3), with a half-maximal effective concentration of 4.48 μM [127]. Accumulating evidence indicates that resveratrol inhibits major pathways involved in the pathogenesis of SARS-CoV-2 by modulating the expression of ACE2 and suppressing the release of proinflammatory cytokines and the production of ROS [128]. In one study, resveratrol reduced the titer of SARS-CoV-2 with minimal cytotoxic effects [129]. Therefore, resveratrol may be used alone or in combination with FDA-approved drugs to treat COVID-19 [130]. Resveratrol/CM-glucan formulations may be suitable for the simultaneous volatilization of aerosols in the treatment of patients with lower respiratory SARS-CoV-2 infections [131]. Overall, resveratrol may reduce the spread of SARS-CoV-2 in the lower respiratory tract and inhibit viral replication in the early stages of infection [10]. Fu et al. reported that resveratrol inhibits pancreatic cancer growth and metastasis by upregulating the expression of hsa-miR‑200 family members [132]. Han et al. reported that resveratrol suppressed growth inhibition by inducing hsa-miR-622 expression in 16HBE-T cells [133]. Furthermore, resveratrol exhibited neuroprotective properties in pneumococcal meningitis by modulating global miRNA expression, such as by upregulating the expression of hsa-miR-15b-5p, hsa-miR-25-3p, and hsa-miR-125b-5p. Among these resveratrol-induced miRNAs, hsa-miR-200 family miRNA and hsa-miR-125b expression can suppress ACE2 expression. Furthermore, hsa-miR-15b and hsa-miR-622 bind to the SARS-CoV-2 S protein. Taken together, these findings suggest that resveratrol can regulate miRNA expression, thus preventing SARS-CoV-2 infection by inhibiting viral entry and viral replication (Figure 6).
Curcumin is mainly present in turmeric plants and curry powders and is not widespread in food [107]. One clinical trial revealed that that treating COVID-19 with nanocurcumin significantly reduced the serum concentrations of IL-6 and IL-1β [134, 135]. One in vitro study reported that curcumin (10 µg/mL) could inhibit 99% and 99.8% of the viral activity of the DG614 strain and Delta variant of SARS-CoV-2, respectively [136]. In another study investigating the effect of turmeric root extract on SARS-CoV-2, curcumin achieved the complete neutralization of SARS-CoV-2 at a subtoxic concentration of 15.6 µg/mL (Table 3) [137]. Goc et al. reported that at concentrations of 10 to 25 μg/mL, curcumin inhibited 20% to 30% of ACE2 activity in cell-based assays (Table 3) [138]. Another study reported that curcumin exhibited a high binding affinity to the protease of SARS-CoV-2, indicating that curcumin may be suitable for use in the development of drugs that can prevent the host cell entry and replication of coronaviruses [139]. Curcumin treatment can induce the expression of hsa-miR-9, hsa-miR-200 family miRNAs, hsa-miR-203, hsa-miR-16, and hsa-let-7a in cancer cells [140, 141]. In an orthotopic xenograft model of human pancreatic cancer, curcumin treatment inhibited tumor growth by inducing hsa-let-7 expression [142]. These results indicate that curcumin can protect against SARS-CoV-2 entry by inducing the expression of let-7 family miRNAs, which suppress TMPRSS2 expression (Figure 6). In addition, curcumin treatment upregulated the expression of hsa-miR-199 and hsa-miR-200 in vivo, suggesting that curcumin treatment is beneficial for liver fibrosis [143]. In leukemia cells, curcumin treatment significantly increased the expression of hsa-miR-16, which induced cell death by apoptosis and suppressed cell proliferation [144]. Hsa-miR-16 upregulation was also observed in human breast cancer cells treated with curcumin [145]. In another study, curcumin directly induced the expression of the tumor-suppressive microRNA hsa-miR-203 in bladder cancer by hypomethylating the hsa-miR-203 promoter [146]. Curcumin suppressed the expression of genes encoding amyloid precursor proteins and amyloid-β and upregulated the expression of hsa-miR-15b-5p in swAPP695-HEK293 cells [147]. Taken together, the curcumin-induced expression of hsa-miR-15b, hsa-miR-200 family miRNA, and hsa-miR-125b suppressed the expression of ACE2 and the S protein gene, indicating that curcumin can protect against SARS-CoV-2 infection (Figure 6).
The aforementioned natural compounds exhibit potential for use as anti-SARS-CoV-2 infection agents because they reduce infection-induced inflammation and ROS generation and prevent interactions between SARS-CoV-2 and host cells. Combined use with antiviral drugs can synergistically improve the efficiency of these agents in protecting against SARS-CoV-2 with minimal side effects. Using molecular docking approaches, some studies have demonstrated that flavonoids target ACE-2 and S proteins. Among these flavonoids, quercetin, EGCG, curcumin, and resveratrol exhibit high binding affinities to ACE-2 receptors and thus inhibit the entry of SARS-CoV-2 into host cells [148-152]. In addition to directly binding to ACE-2/S proteins to prevent SARS-CoV-2 entry, miRNAs can protect against SARS-CoV-2 by downregulating the expression of ACE2, TMPRSS2, and the S protein gene. Herein, we summarize recent findings indicating that these natural compounds protect against SARS-CoV-2 entry host cells by modulating miRNA expression (Figure 6).
Conclusions
Antiviral drugs and vaccines are key cost-effective tools for controlling the COVID-19 pandemic. However, the emergence of SARS-CoV-2 variants could threaten the global impact of mass vaccination and the effectiveness of antiviral drug development. Therefore, adjunctive therapy with nutritional polyphenolic compounds would be a beneficial alternative. As miRNAs are involved in SARS-CoV-2-induced immune evasion and cytokine storm, this contributes to the progression of COVID-19. Polyphenols are known to modulate miRNAs in various disease entities, including RNA virus infection. In this article, we preliminarily explore the prospective roles of miRNAs in the pathogenesis of SARS-CoV-2, including host cell miRNA-human gene interactions, human miRNA-SARS-CoV-2 transcript interactions, SARS-CoV-2 derived RNA transcripts act as decoy human miRNAs, and SARS-CoV-2 miRNA-human gene interactions. Polyphenolic compounds can prevent SARS-CoV-2 infection and inhibit SARS-CoV-2 replication by modulating host miRNAs. Suppressing infection and replication of SARS-Cov-2 can reduce the amount of cellular ceRNA, thereby maintaining the normal physiological function of the host miRNAs. Certain circulating miRNAs might be useful biomarkers of COVID-19 or be used in alternative approaches to COVID-19 treatment. In this paper, we describe four polyphenols, namely quercetin, EGCG, curcumin, and resveratrol, that are present in foods and may provide prophylactic or therapeutic benefits for SARS-CoV-2 infection by modulating host miRNAs. In the future, additional biological experiments and clinical trials must be conducted to further clarify these findings and to further elucidate the role of these miRNAs in the pathogenesis and treatment of COVID-19.
Acknowledgements
This study was supported by Taipei Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation (TCRD-TPE-109-66, TCRD-TPE-MOST-109-17, TCRD-TPE-110-RT-1 and TCMF-CM3-111-01) and Kaohsiung Municipal Min-Sheng Hospital (KMSH-11105).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Habas K, Nganwuchu C, Shahzad F, Gopalan R, Haque M, Rahman S. et al. Resolution of coronavirus disease 2019 (COVID-19). Expert Rev Anti Infect Ther. 2020;18:1201-11
2. Cele S, Gazy I, Jackson L, Hwa SH, Tegally H, Lustig G. et al. Escape of SARS-CoV-2 501Y.V2 from neutralization by convalescent plasma. Nature. 2021;593:142-6
3. Garcia-Beltran WF, St Denis KJ, Hoelzemer A, Lam EC, Nitido AD, Sheehan ML. et al. mRNA-based COVID-19 vaccine boosters induce neutralizing immunity against SARS-CoV-2 Omicron variant. Cell. 2022;185:457-466
4. Cui X, Zhao Z, Zhang T, Guo W, Guo W, Zheng J. et al. A systematic review and meta-analysis of children with coronavirus disease 2019 (COVID-19). J Med Virol. 2021;93:1057-69
5. O'Driscoll M, Ribeiro Dos Santos G, Wang L, Cummings DAT, Azman AS, Paireau J. et al. Age-specific mortality and immunity patterns of SARS-CoV-2. Nature. 2021;590:140-5
6. Kruse RL. Therapeutic strategies in an outbreak scenario to treat the novel coronavirus originating in Wuhan, China. F1000Res. 2020;9:72
7. Khan MA, Sany MRU, Islam MS, Islam A. Epigenetic Regulator miRNA Pattern Differences Among SARS-CoV, SARS-CoV-2, and SARS-CoV-2 World-Wide Isolates Delineated the Mystery Behind the Epic Pathogenicity and Distinct Clinical Characteristics of Pandemic COVID-19. Front Genet. 2020;11:765
8. Majumder J, Minko T. Recent Developments on Therapeutic and Diagnostic Approaches for COVID-19. AAPS J. 2021;23:14
9. Zhang S, Amahong K, Sun X, Lian X, Liu J, Sun H. et al. The miRNA: a small but powerful RNA for COVID-19. Brief Bioinform. 2021;22:1137-49
10. Liao MT, Wu CC, Wu SV, Lee MC, Hu WC, Tsai KW. et al. Resveratrol as an Adjunctive Therapy for Excessive Oxidative Stress in Aging COVID-19 Patients. Antioxidants (Basel). 2021;10:1440
11. Ahmadian R, Biganeh H, Panahi Y, Guest PC, Jamialahmadi T, Sahebkar A. Resveratrol as a Probable Multiheaded Treatment Approach for COVID-19. Adv Exp Med Biol. 2021;1328:441-6
12. Cione E, La Torre C, Cannataro R, Caroleo MC, Plastina P, Gallelli L. Quercetin, Epigallocatechin Gallate, Curcumin, and Resveratrol: From Dietary Sources to Human MicroRNA Modulation. Molecules. 2019;25:63
13. Yekta S, Shih IH, Bartel DP. MicroRNA-directed cleavage of HOXB8 mRNA. Science (New York, NY. 2004;304:594-6
14. Newman MA, Hammond SM. Emerging paradigms of regulated microRNA processing. Genes & development. 2010;24:1086-92
15. Trabucchi M, Briata P, Filipowicz W, Rosenfeld MG, Ramos A, Gherzi R. How to control miRNA maturation? RNA Biol. 2009;6:536-40
16. Slezak-Prochazka I, Durmus S, Kroesen BJ, van den Berg A. MicroRNAs, macrocontrol: regulation of miRNA processing. RNA (New York, NY. 2010;16:1087-95
17. Leung CM, Li SC, Chen TW, Ho MR, Hu LY, Liu WS. et al. Comprehensive microRNA profiling of prostate cancer cells after ionizing radiation treatment. Oncology reports. 2014;31:1067-78
18. Cheng WC, Chung IF, Huang TS, Chang ST, Sun HJ, Tsai CF. et al. YM500: a small RNA sequencing (smRNA-seq) database for microRNA research. Nucleic acids research. 2013;41:D285-94
19. Li SC, Liao YL, Ho MR, Tsai KW, Lai CH, Lin WC. miRNA arm selection and isomiR distribution in gastric cancer. BMC genomics. 2012;13(Suppl 1):S13
20. Chang HT, Li SC, Ho MR, Pan HW, Ger LP, Hu LY. et al. Comprehensive analysis of microRNAs in breast cancer. BMC genomics. 2012;13(Suppl 7):S18
21. Li SC, Liao YL, Chan WC, Ho MR, Tsai KW, Hu LY. et al. Interrogation of rabbit miRNAs and their isomiRs. Genomics. 2011;98:453-9
22. Griffiths-Jones S, Hui JH, Marco A, Ronshaugen M. MicroRNA evolution by arm switching. EMBO Rep. 2011;12:172-7
23. Cloonan N, Wani S, Xu Q, Gu J, Lea K, Heater S. et al. MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome biology. 2011;12:R126
24. Marco A, Hui JH, Ronshaugen M, Griffiths-Jones S. Functional shifts in insect microRNA evolution. Genome biology and evolution. 2010;2:686-96
25. Guo L, Li H, Liang T, Lu J, Yang Q, Ge Q. et al. Consistent isomiR expression patterns and 3' addition events in miRNA gene clusters and families implicate functional and evolutionary relationships. Molecular biology reports. 2012;39:6699-706
26. Tsai KW, Leung CM, Lo YH, Chen TW, Chan WC, Yu SY. et al. Arm Selection Preference of MicroRNA-193a Varies in Breast Cancer. Scientific reports. 2016;6:28176
27. Chatterjee S, Fasler M, Bussing I, Grosshans H. Target-mediated protection of endogenous microRNAs in C. elegans. Developmental cell. 2011;20:388-96
28. Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11:597-610
29. Bernier A, Sagan SM. The Diverse Roles of microRNAs at the Host(-)Virus Interface. Viruses. 2018 10
30. Mallick B, Ghosh Z, Chakrabarti J. MicroRNome analysis unravels the molecular basis of SARS infection in bronchoalveolar stem cells. PLoS One. 2009;4:e7837
31. Gottwein E, Cullen BR. Viral and cellular microRNAs as determinants of viral pathogenesis and immunity. Cell Host Microbe. 2008;3:375-87
32. Shang J, Wan Y, Luo C, Ye G, Geng Q, Auerbach A. et al. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci U S A. 2020;117:11727-34
33. V'Kovski P, Kratzel A, Steiner S, Stalder H, Thiel V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat Rev Microbiol. 2021;19:155-70
34. Poduri R, Joshi G, Jagadeesh G. Drugs targeting various stages of the SARS-CoV-2 life cycle: Exploring promising drugs for the treatment of Covid-19. Cell Signal. 2020;74:109721
35. Hirasawa K, Kim A, Han HS, Han J, Jun HS, Yoon JW. Effect of p38 mitogen-activated protein kinase on the replication of encephalomyocarditis virus. J Virol. 2003;77:5649-56
36. Elbahesh H, Cline T, Baranovich T, Govorkova EA, Schultz-Cherry S, Russell CJ. Novel roles of focal adhesion kinase in cytoplasmic entry and replication of influenza A viruses. J Virol. 2014;88:6714-28
37. Diehl N, Schaal H. Make yourself at home: viral hijacking of the PI3K/Akt signaling pathway. Viruses. 2013;5:3192-212
38. Chow JT, Salmena L. Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium. Genes (Basel). 2020;11:1002
39. Sato A, Ogino Y, Tanuma SI, Uchiumi F. Human microRNA hsa-miR-15b-5p targets the RNA template component of the RNA-dependent RNA polymerase structure in severe acute respiratory syndrome coronavirus 2. Nucleosides Nucleotides Nucleic Acids. 2021;40:790-7
40. Wang Y, Li J, Zhang L, Sun HX, Zhang Z, Xu J. et al. Plasma cell-free RNA characteristics in COVID-19 patients. Genome Res. 2022;32:228-41
41. Meidert AS, Hermann S, Brandes F, Kirchner B, Buschmann D, Billaud JN. et al. Extracellular Vesicle Associated miRNAs Regulate Signaling Pathways Involved in COVID-19 Pneumonia and the Progression to Severe Acute Respiratory Corona Virus-2 Syndrome. Front Immunol. 2021;12:784028
42. Trobaugh DW, Klimstra WB. MicroRNA Regulation of RNA Virus Replication and Pathogenesis. Trends Mol Med. 2017;23:80-93
43. Lu D, Chatterjee S, Xiao K, Riedel I, Wang Y, Foo R. et al. MicroRNAs targeting the SARS-CoV-2 entry receptor ACE2 in cardiomyocytes. J Mol Cell Cardiol. 2020;148:46-9
44. Matarese A, Gambardella J, Sardu C, Santulli G. miR-98 Regulates TMPRSS2 Expression in Human Endothelial Cells: Key Implications for COVID-19. Biomedicines. 2020;8:462
45. Nersisyan S, Shkurnikov M, Turchinovich A, Knyazev E, Tonevitsky A. Integrative analysis of miRNA and mRNA sequencing data reveals potential regulatory mechanisms of ACE2 and TMPRSS2. PLoS One. 2020;15:e0235987
46. Pierce JB, Simion V, Icli B, Perez-Cremades D, Cheng HS, Feinberg MW. Computational Analysis of Targeting SARS-CoV-2, Viral Entry Proteins ACE2 and TMPRSS2, and Interferon Genes by Host MicroRNAs. Genes (Basel). 2020;11:1354
47. Widiasta A, Sribudiani Y, Nugrahapraja H, Hilmanto D, Sekarwana N, Rachmadi D. Potential role of ACE2-related microRNAs in COVID-19-associated nephropathy. Noncoding RNA Res. 2020;5:153-66
48. Tsai KW, Chong KH, Li CH, Tu YT, Chen YR, Lee MC. et al. LOC550643, a Long Non-coding RNA, Acts as Novel Oncogene in Regulating Breast Cancer Growth and Metastasis. Front Cell Dev Biol. 2021;9:695632
49. Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17:272-83
50. Arora S, Singh P, Dohare R, Jha R, Ali Syed M. Unravelling host-pathogen interactions: ceRNA network in SARS-CoV-2 infection (COVID-19). Gene. 2020;762:145057
51. Qi X, Zhang DH, Wu N, Xiao JH, Wang X, Ma W. ceRNA in cancer: possible functions and clinical implications. J Med Genet. 2015;52:710-8
52. Wu X, Sui Z, Zhang H, Wang Y, Yu Z. Integrated Analysis of lncRNA-Mediated ceRNA Network in Lung Adenocarcinoma. Front Oncol. 2020;10:554759
53. Haddad H, Walid A-Z. miRNA target prediction might explain the reduced transmission of SARS-CoV-2 in Jordan, Middle East. Noncoding RNA Res. 2020;5:135-43
54. Sardar R, Satish D, Birla S, Gupta D. Integrative analyses of SARS-CoV-2 genomes from different geographical locations reveal unique features potentially consequential to host-virus interaction, pathogenesis and clues for novel therapies. Heliyon. 2020;6:e04658
55. Hosseini Rad Sm A, McLellan AD. Implications of SARS-CoV-2 Mutations for Genomic RNA Structure and Host microRNA Targeting. Int J Mol Sci. 2020;21:4807
56. Nersisyan S, Engibaryan N, Gorbonos A, Kirdey K, Makhonin A, Tonevitsky A. Potential role of cellular miRNAs in coronavirus-host interplay. PeerJ. 2020;8:e9994
57. Fulzele S, Sahay B, Yusufu I, Lee TJ, Sharma A, Kolhe R. et al. COVID-19 Virulence in Aged Patients Might Be Impacted by the Host Cellular MicroRNAs Abundance/Profile. Aging Dis. 2020;11:509-22
58. Sacar Demirci MD, Adan A. Computational analysis of microRNA-mediated interactions in SARS-CoV-2 infection. PeerJ. 2020;8:e9369
59. Schonrich G, Raftery MJ, Samstag Y. Devilishly radical NETwork in COVID-19: Oxidative stress, neutrophil extracellular traps (NETs), and T cell suppression. Adv Biol Regul. 2020;77:100741
60. Miller K, McGrath ME, Hu Z, Ariannejad S, Weston S, Frieman M. et al. Coronavirus interactions with the cellular autophagy machinery. Autophagy. 2020;16:2131-9
61. Vagapova ER, Lebedev TD, Prassolov VS. Viral fibrotic scoring and drug screen based on MAPK activity uncovers EGFR as a key regulator of COVID-19 fibrosis. Scientific reports. 2021;11:11234
62. Skalsky RL, Cullen BR. Viruses, microRNAs, and host interactions. Annu Rev Microbiol. 2010;64:123-41
63. Ghosh Z, Mallick B, Chakrabarti J. Cellular versus viral microRNAs in host-virus interaction. Nucleic acids research. 2009;37:1035-48
64. Yuen CK, Lam JY, Wong WM, Mak LF, Wang X, Chu H. et al. SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 function as potent interferon antagonists. Emerg Microbes Infect. 2020;9:1418-28
65. Miyamoto Y, Itoh Y, Suzuki T, Tanaka T, Sakai Y, Koido M. et al. SARS-CoV-2 ORF6 disrupts nucleocytoplasmic trafficking to advance viral replication. Commun Biol. 2022;5:483
66. Arghiani N, Nissan T, Matin MM. Role of microRNAs in COVID-19 with implications for therapeutics. Biomed Pharmacother. 2021;144:112247
67. Chen L, Zhong L. Genomics functional analysis and drug screening of SARS-CoV-2. Genes Dis. 2020;7:542-50
68. Bartoszewski R, Dabrowski M, Jakiela B, Matalon S, Harrod KS, Sanak M. et al. SARS-CoV-2 may regulate cellular responses through depletion of specific host miRNAs. Am J Physiol Lung Cell Mol Physiol. 2020;319:L444-L55
69. Akula SM, Bolin P, Cook PP. Cellular miR-150-5p may have a crucial role to play in the biology of SARS-CoV-2 infection by regulating nsp10 gene. RNA Biol. 2022;19:1-11
70. Mishra R, Kumar A, Ingle H, Kumar H. The Interplay Between Viral-Derived miRNAs and Host Immunity During Infection. Front Immunol. 2019;10:3079
71. Ahmad I, Valverde A, Siddiqui H, Schaller S, Naqvi AR. Viral MicroRNAs: Interfering the Interferon Signaling. Curr Pharm Des. 2020;26:446-54
72. Perez JT, Varble A, Sachidanandam R, Zlatev I, Manoharan M, Garcia-Sastre A. et al. Influenza A virus-generated small RNAs regulate the switch from transcription to replication. Proc Natl Acad Sci U S A. 2010;107:11525-30
73. Weng KF, Hung CT, Hsieh PT, Li ML, Chen GW, Kung YA. et al. A cytoplasmic RNA virus generates functional viral small RNAs and regulates viral IRES activity in mammalian cells. Nucleic acids research. 2014;42:12789-805
74. Shi J, Sun J, Wang B, Wu M, Zhang J, Duan Z. et al. Novel microRNA-like viral small regulatory RNAs arising during human hepatitis A virus infection. FASEB J. 2014;28:4381-93
75. Morales L, Oliveros JC, Fernandez-Delgado R, tenOever BR, Enjuanes L, Sola I. SARS-CoV-Encoded Small RNAs Contribute to Infection-Associated Lung Pathology. Cell Host Microbe. 2017;21:344-55
76. Battaglia R, Alonzo R, Pennisi C, Caponnetto A, Ferrara C, Stella M. et al. MicroRNA-Mediated Regulation of the Virus Cycle and Pathogenesis in the SARS-CoV-2 Disease. Int J Mol Sci. 2021 22
77. Walsh D, Mathews MB, Mohr I. Tinkering with translation: protein synthesis in virus-infected cells. Cold Spring Harb Perspect Biol. 2013;5:a012351
78. Walsh D, Mohr I. Viral subversion of the host protein synthesis machinery. Nat Rev Microbiol. 2011;9:860-75
79. Arisan ED, Dart A, Grant GH, Arisan S, Cuhadaroglu S, Lange S. et al. The Prediction of miRNAs in SARS-CoV-2 Genomes: hsa-miR Databases Identify 7 Key miRs Linked to Host Responses and Virus Pathogenicity-Related KEGG Pathways Significant for Comorbidities. Viruses. 2020;12:614
80. Fu Z, Wang J, Wang Z, Sun Y, Wu J, Zhang Y. et al. A virus-derived microRNA-like small RNA serves as a serum biomarker to prioritize the COVID-19 patients at high risk of developing severe disease. Cell Discov. 2021;7:48
81. Aydemir MN, Aydemir HB, Korkmaz EM, Budak M, Cekin N, Pinarbasi E. Computationally predicted SARS-COV-2 encoded microRNAs target NFKB, JAK/STAT and TGFB signaling pathways. Gene Rep. 2021;22:101012
82. Maranini B, Ciancio G, Ferracin M, Cultrera R, Negrini M, Sabbioni S. et al. microRNAs and Inflammatory Immune Response in SARS-CoV-2 Infection: A Narrative Review. Life (Basel). 2022;12:288
83. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K. et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell research. 2008;18:997-1006
84. Gilad S, Meiri E, Yogev Y, Benjamin S, Lebanony D, Yerushalmi N. et al. Serum microRNAs are promising novel biomarkers. PLoS One. 2008;3:e3148
85. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105:10513-8
86. Chim SS, Shing TK, Hung EC, Leung TY, Lau TK, Chiu RW. et al. Detection and characterization of placental microRNAs in maternal plasma. Clinical chemistry. 2008;54:482-90
87. Bagheri-Hosseinabadi Z, Ostad Ebrahimi H, Bahrehmand F, Taghipour G, Abbasifard M. The relationship between serum levels of interleukin-2 and IL-8 with circulating microRNA-10b in patients with COVID-19. Iran J Immunol. 2021;18:65-73
88. Martinez-Fleta P, Vera-Tome P, Jimenez-Fernandez M, Requena S, Roy-Vallejo E, Sanz-Garcia A. et al. A Differential Signature of Circulating miRNAs and Cytokines Between COVID-19 and Community-Acquired Pneumonia Uncovers Novel Physiopathological Mechanisms of COVID-19. Front Immunol. 2021;12:815651
89. Haroun RA, Osman WH, Amin RE, Hassan AK, Abo-Shanab WS, Eessa AM. Circulating plasma miR-155 is a potential biomarker for the detection of SARS-CoV-2 infection. Pathology. 2022;54:104-10
90. Garg A, Seeliger B, Derda AA, Xiao K, Gietz A, Scherf K. et al. Circulating cardiovascular microRNAs in critically ill COVID-19 patients. Eur J Heart Fail. 2021;23:468-75
91. Donyavi T, Bokharaei-Salim F, Baghi HB, Khanaliha K, Alaei Janat-Makan M, Karimi B. et al. Acute and post-acute phase of COVID-19: Analyzing expression patterns of miRNA-29a-3p, 146a-3p, 155-5p, and let-7b-3p in PBMC. Int Immunopharmacol. 2021;97:107641
92. McDonald JT, Enguita FJ, Taylor D, Griffin RJ, Priebe W, Emmett MR. et al. Role of miR-2392 in driving SARS-CoV-2 infection. Cell Rep. 2021;37:109839
93. Fayyad-Kazan M, Makki R, Skafi N, El Homsi M, Hamade A, El Majzoub R. et al. Circulating miRNAs: Potential diagnostic role for coronavirus disease 2019 (COVID-19). Infect Genet Evol. 2021;94:105020
94. Gallo Marin B, Aghagoli G, Lavine K, Yang L, Siff EJ, Chiang SS. et al. Predictors of COVID-19 severity: A literature review. Rev Med Virol. 2021;31:1-10
95. Feng E, Balint E, Poznanski SM, Ashkar AA, Loeb M. Aging and Interferons: Impacts on Inflammation and Viral Disease Outcomes. Cells. 2021;10:708
96. Duecker RP, Adam EH, Wirtz S, Gronau L, Khodamoradi Y, Eberhardt FJ. et al. The MiR-320 Family Is Strongly Downregulated in Patients with COVID-19 Induced Severe Respiratory Failure. Int J Mol Sci. 2021;22:10351
97. Pimenta R, Viana NI, Dos Santos GA, Candido P, Guimaraes VR, Romao P. et al. MiR-200c-3p expression may be associated with worsening of the clinical course of patients with COVID-19. Mol Biol Res Commun. 2021;10:141-7
98. Sabbatinelli J, Giuliani A, Matacchione G, Latini S, Laprovitera N, Pomponio G. et al. Decreased serum levels of the inflammaging marker miR-146a are associated with clinical non-response to tocilizumab in COVID-19 patients. Mech Ageing Dev. 2021;193:111413
99. de Gonzalo-Calvo D, Benitez ID, Pinilla L, Carratala A, Moncusi-Moix A, Gort-Paniello C. et al. Circulating microRNA profiles predict the severity of COVID-19 in hospitalized patients. Transl Res. 2021;236:147-59
100. Giuliani A, Matacchione G, Ramini D, Di Rosa M, Bonfigli AR, Sabbatinelli J. et al. Circulating miR-320b and miR-483-5p levels are associated with COVID-19 in-hospital mortality. Mech Ageing Dev. 2022;202:111636
101. Fazio A, Iacopetta D, La Torre C, Ceramella J, Muia N, Catalano A. et al. Finding solutions for agricultural wastes: antioxidant and antitumor properties of pomegranate Akko peel extracts and beta-glucan recovery. Food Funct. 2018;9:6618-31
102. Roman GC, Jackson RE, Gadhia R, Roman AN, Reis J. Mediterranean diet: The role of long-chain omega-3 fatty acids in fish; polyphenols in fruits, vegetables, cereals, coffee, tea, cacao and wine; probiotics and vitamins in prevention of stroke, age-related cognitive decline, and Alzheimer disease. Rev Neurol (Paris). 2019;175:724-41
103. Mhatre S, Srivastava T, Naik S, Patravale V. Antiviral activity of green tea and black tea polyphenols in prophylaxis and treatment of COVID-19: A review. Phytomedicine. 2021;85:153286
104. Iddir M, Brito A, Dingeo G, Fernandez Del Campo SS, Samouda H, La Frano MR. et al. Strengthening the Immune System and Reducing Inflammation and Oxidative Stress through Diet and Nutrition: Considerations during the COVID-19 Crisis. Nutrients. 2020;12:1562
105. Annunziata G, Sanduzzi Zamparelli M, Santoro C, Ciampaglia R, Stornaiuolo M, Tenore GC. et al. May Polyphenols Have a Role Against Coronavirus Infection? An Overview of in vitro Evidence. Front Med (Lausanne). 2020;7:240
106. Milenkovic D, Ruskovska T, Rodriguez-Mateos A, Heiss C. Polyphenols Could Prevent SARS-CoV-2 Infection by Modulating the Expression of miRNAs in the Host Cells. Aging Dis. 2021;12:1169-82
107. Esatbeyoglu T, Huebbe P, Ernst IM, Chin D, Wagner AE, Rimbach G. Curcumin-from molecule to biological function. Angew Chem Int Ed Engl. 2012;51:5308-32
108. Ranka S, Gee JM, Biro L, Brett G, Saha S, Kroon P. et al. Development of a food frequency questionnaire for the assessment of quercetin and naringenin intake. Eur J Clin Nutr. 2008;62:1131-8
109. Di Petrillo A, Orru G, Fais A, Fantini MC. Quercetin and its derivates as antiviral potentials: A comprehensive review. Phytother Res. 2022;36:266-78
110. Shohan M, Nashibi R, Mahmoudian-Sani MR, Abolnezhadian F, Ghafourian M, Alavi SM. et al. The therapeutic efficacy of quercetin in combination with antiviral drugs in hospitalized COVID-19 patients: A randomized controlled trial. Eur J Pharmacol. 2022;914:174615
111. Zhu Y, Scholle F, Kisthardt SC, Xie DY. Flavonols and dihydroflavonols inhibit the main protease activity of SARS-CoV-2 and the replication of human coronavirus 229E. Virology. 2022;571:21-33
112. Mangiavacchi F, Botwina P, Menichetti E, Bagnoli L, Rosati O, Marini F. et al. Seleno-Functionalization of Quercetin Improves the Non-Covalent Inhibition of M(pro) and Its Antiviral Activity in Cells against SARS-CoV-2. Int J Mol Sci. 2021;22:7048
113. Appari M, Babu KR, Kaczorowski A, Gross W, Herr I. Sulforaphane, quercetin and catechins complement each other in elimination of advanced pancreatic cancer by miR-let-7 induction and K-ras inhibition. Int J Oncol. 2014;45:1391-400
114. Nwaeburu CC, Bauer N, Zhao Z, Abukiwan A, Gladkich J, Benner A. et al. Up-regulation of microRNA let-7c by quercetin inhibits pancreatic cancer progression by activation of Numbl. Oncotarget. 2016;7:58367-80
115. Nwaeburu CC, Abukiwan A, Zhao Z, Herr I. Quercetin-induced miR-200b-3p regulates the mode of self-renewing divisions in pancreatic cancer. Mol Cancer. 2017;16:23
116. Sonoki H, Sato T, Endo S, Matsunaga T, Yamaguchi M, Yamazaki Y. et al. Quercetin Decreases Claudin-2 Expression Mediated by Up-Regulation of microRNA miR-16 in Lung Adenocarcinoma A549 Cells. Nutrients. 2015;7:4578-92
117. Rodriguez-Carrasco Y, Gaspari A, Graziani G, Santini A, Ritieni A. Fast analysis of polyphenols and alkaloids in cocoa-based products by ultra-high performance liquid chromatography and Orbitrap high resolution mass spectrometry (UHPLC-Q-Orbitrap-MS/MS). Food Res Int. 2018;111:229-36
118. Henss L, Auste A, Schurmann C, Schmidt C, von Rhein C, Muhlebach MD. et al. The green tea catechin epigallocatechin gallate inhibits SARS-CoV-2 infection. J Gen Virol. 2021;102:001574
119. Gowrishankar S, Muthumanickam S, Kamaladevi A, Karthika C, Jothi R, Boomi P. et al. Promising phytochemicals of traditional Indian herbal steam inhalation therapy to combat COVID-19 - An in silico study. Food Chem Toxicol. 2021;148:111966
120. Rasheed Z, Rasheed N, Al-Shaya O. Epigallocatechin-3-O-gallate modulates global microRNA expression in interleukin-1beta-stimulated human osteoarthritis chondrocytes: potential role of EGCG on negative co-regulation of microRNA-140-3p and ADAMTS5. Eur J Nutr. 2018;57:917-28
121. Li P, Xu Y, Wang B, Huang J, Li Q. miR-34a-5p and miR-125b-5p attenuate Abeta-induced neurotoxicity through targeting BACE1. J Neurol Sci. 2020;413:116793
122. Zhu Y, Huang Y, Liu M, Yan Q, Zhao W, Yang P. et al. Epigallocatechin gallate inhibits cell growth and regulates miRNA expression in cervical carcinoma cell lines infected with different high-risk human papillomavirus subtypes. Exp Ther Med. 2019;17:1742-8
123. Zhang S, Al-Maghout T, Bissinger R, Zeng N, Pelzl L, Salker MS. et al. Epigallocatechin-3-gallate (EGCG) up-regulates miR-15b expression thus attenuating store operated calcium entry (SOCE) into murine CD4(+) T cells and human leukaemic T cell lymphoblasts. Oncotarget. 2017;8:89500-14
124. Shin S, Kim K, Lee MJ, Lee J, Choi S, Kim KS. et al. Epigallocatechin Gallate-Mediated Alteration of the MicroRNA Expression Profile in 5alpha-Dihydrotestosterone-Treated Human Dermal Papilla Cells. Ann Dermatol. 2016;28:327-34
125. Singh AP, Singh R, Verma SS, Rai V, Kaschula CH, Maiti P. et al. Health benefits of resveratrol: Evidence from clinical studies. Med Res Rev. 2019;39:1851-91
126. Ter Ellen BM, Dinesh Kumar N, Bouma EM, Troost B, van de Pol DPI, van der Ende-Metselaar HH. et al. Resveratrol and Pterostilbene Inhibit SARS-CoV-2 Replication in Air-Liquid Interface Cultured Human Primary Bronchial Epithelial Cells. Viruses. 2021;13:1335
127. Yang M, Wei J, Huang T, Lei L, Shen C, Lai J. et al. Resveratrol inhibits the replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in cultured Vero cells. Phytother Res. 2021;35:1127-9
128. Ramdani LH, Bachari K. Potential therapeutic effects of Resveratrol against SARS-CoV-2. Acta Virol. 2020;64:276-80
129. Pasquereau S, Nehme Z, Haidar Ahmad S, Daouad F, Van Assche J, Wallet C. et al. Resveratrol Inhibits HCoV-229E and SARS-CoV-2 Coronavirus Replication In Vitro. Viruses. 2021;13:354
130. Yang L, Wang Z. Natural Products, Alone or in Combination with FDA-Approved Drugs, to Treat COVID-19 and Lung Cancer. Biomedicines. 2021;9:689
131. Perrella F, Coppola F, Petrone A, Platella C, Montesarchio D, Stringaro A. et al. Interference of Polydatin/Resveratrol in the ACE2:Spike Recognition during COVID-19 Infection. A Focus on Their Potential Mechanism of Action through Computational and Biochemical Assays. Biomolecules. 2021;11:1048
132. Fu J, Shrivastava A, Shrivastava SK, Srivastava RK, Shankar S. Triacetyl resveratrol upregulates miRNA200 and suppresses the Shh pathway in pancreatic cancer: A potential therapeutic agent. Int J Oncol. 2019;54:1306-16
133. Han Z, Yang Q, Liu B, Wu J, Li Y, Yang C. et al. MicroRNA-622 functions as a tumor suppressor by targeting K-Ras and enhancing the anticarcinogenic effect of resveratrol. Carcinogenesis. 2012;33:131-9
134. Valizadeh H, Abdolmohammadi-Vahid S, Danshina S, Ziya Gencer M, Ammari A, Sadeghi A. et al. Nano-curcumin therapy, a promising method in modulating inflammatory cytokines in COVID-19 patients. Int Immunopharmacol. 2020;89:107088
135. Hassaniazad M, Inchehsablagh BR, Kamali H, Tousi A, Eftekhar E, Jaafari MR. et al. The clinical effect of Nano micelles containing curcumin as a therapeutic supplement in patients with COVID-19 and the immune responses balance changes following treatment: A structured summary of a study protocol for a randomised controlled trial. Trials. 2020;21:876
136. Marin-Palma D, Tabares-Guevara JH, Zapata-Cardona MI, Florez-Alvarez L, Yepes LM, Rugeles MT. et al. Curcumin Inhibits In Vitro SARS-CoV-2 Infection In Vero E6 Cells through Multiple Antiviral Mechanisms. Molecules. 2021;26:6900
137. Bormann M, Alt M, Schipper L, van de Sand L, Le-Trilling VTK, Rink L. et al. Turmeric Root and Its Bioactive Ingredient Curcumin Effectively Neutralize SARS-CoV-2 In Vitro. Viruses. 2021;13:1914
138. Goc A, Sumera W, Rath M, Niedzwiecki A. Phenolic compounds disrupt spike-mediated receptor-binding and entry of SARS-CoV-2 pseudo-virions. PLoS One. 2021;16:e0253489
139. Oso BJ, Adeoye AO, Olaoye IF. Pharmacoinformatics and hypothetical studies on allicin, curcumin, and gingerol as potential candidates against COVID-19-associated proteases. J Biomol Struct Dyn. 2022;40:389-400
140. Subramaniam D, Ponnurangam S, Ramamoorthy P, Standing D, Battafarano RJ, Anant S. et al. Curcumin induces cell death in esophageal cancer cells through modulating Notch signaling. PLoS One. 2012;7:e30590
141. Mirzaei H, Masoudifar A, Sahebkar A, Zare N, Sadri Nahand J, Rashidi B. et al. MicroRNA: A novel target of curcumin in cancer therapy. J Cell Physiol. 2018;233:3004-15
142. Bao B, Ali S, Banerjee S, Wang Z, Logna F, Azmi AS. et al. Curcumin analogue CDF inhibits pancreatic tumor growth by switching on suppressor microRNAs and attenuating EZH2 expression. Cancer Res. 2012;72:335-45
143. Hassan ZK, Al-Olayan EM. Curcumin reorganizes miRNA expression in a mouse model of liver fibrosis. Asian Pac J Cancer Prev. 2012;13:5405-8
144. Gao SM, Yang JJ, Chen CQ, Chen JJ, Ye LP, Wang LY. et al. Pure curcumin decreases the expression of WT1 by upregulation of miR-15a and miR-16-1 in leukemic cells. J Exp Clin Cancer Res. 2012;31:27
145. Yang J, Cao Y, Sun J, Zhang Y. Curcumin reduces the expression of Bcl-2 by upregulating miR-15a and miR-16 in MCF-7 cells. Med Oncol. 2010;27:1114-8
146. Saini S, Arora S, Majid S, Shahryari V, Chen Y, Deng G. et al. Curcumin modulates microRNA-203-mediated regulation of the Src-Akt axis in bladder cancer. Cancer Prev Res (Phila). 2011;4:1698-709
147. Liu HY, Fu X, Li YF, Li XL, Ma ZY, Zhang Y. et al. miR-15b-5p targeting amyloid precursor protein is involved in the anti-amyloid eflect of curcumin in swAPP695-HEK293 cells. Neural Regen Res. 2019;14:1603-9
148. Gu YY, Zhang M, Cen H, Wu YF, Lu Z, Lu F. et al. Quercetin as a potential treatment for COVID-19-induced acute kidney injury: Based on network pharmacology and molecular docking study. PLoS One. 2021;16:e0245209
149. Alzaabi MM, Hamdy R, Ashmawy NS, Hamoda AM, Alkhayat F, Khademi NN. et al. Flavonoids are promising safe therapy against COVID-19. Phytochem Rev. 2022;21:291-312
150. Mhatre S, Naik S, Patravale V. A molecular docking study of EGCG and theaflavin digallate with the druggable targets of SARS-CoV-2. Comput Biol Med. 2021;129:104137
151. Wahedi HM, Ahmad S, Abbasi SW. Stilbene-based natural compounds as promising drug candidates against COVID-19. J Biomol Struct Dyn. 2021;39:3225-34
152. Jena AB, Kanungo N, Nayak V, Chainy GBN, Dandapat J. Catechin and curcumin interact with S protein of SARS-CoV2 and ACE2 of human cell membrane: insights from computational studies. Scientific reports. 2021;11:2043
Author contact
Corresponding author: Kuo-Wang Tsai, Department of Research, Taipei Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation, New Taipei, Taiwan; E-Mail: kwtsai6733com

 Global reach, higher impact
Global reach, higher impact