Impact Factor
ISSN: 1449-1907
Int J Med Sci 2021; 18(15):3412-3424. doi:10.7150/ijms.61309 This issue Cite
Research Paper
Integrated analysis of the molecular mechanisms in idiopathic pulmonary fibrosis
1. Department of Respiratory and Critical Care Medicine, The First Affiliated Hospital of Anhui Medical University, Hefei 230022, China
2. Department of General Medicine, Hefei Second People's Hospital, Hefei 230001, China
3. Department of Thoracic Surgery, the first affiliated hospital of Anhui medical university, Hefei 230022, China
4. Hefei Third Clinical College of Anhui Medical University, Hefei 230022, China.
5. Hefei Prevention and Treatment Center for Occupational Diseases, Hefei 230022, China.
*Ke Zhu, Aiqun Xu and Wanli Xia contributed equally to this work.
Abstract
Rationale: Idiopathic pulmonary fibrosis (IPF) is one of the most aggressive forms of idiopathic interstitial pneumonia. Some miRNAs may be associated with IPF and may affect the occurrence and development of IPF in various pathways. Many miRNAs and genes that may be involved in the development of IPF have been discovered using chip and high throughput technologies.
Methods: We analyzed one miRNA and four mRNA databases. We identified hub genes and pathways related to IPF using GO, KEGG enrichment analysis, gene set variation analysis (GSVA), PPI network construction, and hub gene analysis. A comprehensive analysis of differentially expressed miRNAs (DEMs), predicted miRNA target genes, and differentially expressed genes (DEGs) led to the creation of a miRNA-mRNA regulatory network in IPF.
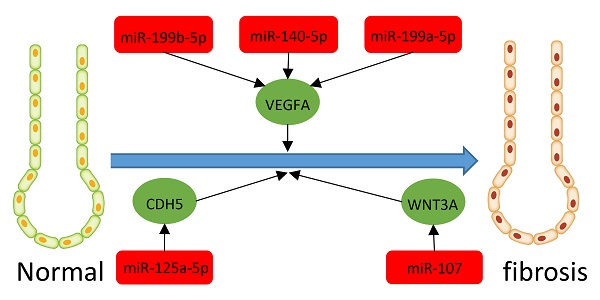
Results: We found 203 DEGs and 165 DEMs that were associated with IPF. The findings of enrichment analyses showed that these DEGs were mainly involved in antimicrobial humoral response, antimicrobial humoral immune response mediated by antimicrobial peptide, extracellular matrix organization, cell killing, and organ or tissue specific immune response. The VEGFA, CDH5, and WNT3A genes overlapped between hub genes and the miRNA-mRNA regulatory network. The miRNAs including miR-199b-5p, miR-140-5p, miR-199a-5p, miR-125A-5p, and miR-107 that we predicted would regulate the VEGFA, CDH5, and WNT3A genes, which were also associated with IPF or other fibrosis-related diseases. GSVA indicated that metabolic processes of UTP and IMP, immune response, regulation of Th2 cell cytokine production, and positive regulation of NK cell-mediated immunity are associated with the pathogenesis and treatment of IPF. These pathways also interact with VEGFA, CDH5, and WNT3A.
Conclusion: These findings provide a new research direction for the diagnosis and treatment of IPF.
Keywords: IPF, Lung tissue, Genes, microRNAs, Bioinformatics

 Global reach, higher impact
Global reach, higher impact