Impact Factor ISSN: 1449-1907
Int J Med Sci 2006; 3(3):108-111. doi:10.7150/ijms.3.108 This issue Cite
Research Paper
Association of the T+294C polymorphism in PPAR δ with low HDL cholesterol and coronary heart disease risk in women
Zentrum für Innere Medizin, Universitätsklinikum Hamburg-Eppendorf, Martinistr. 52, 20246 Hamburg, Germany
Received 2006-2-28; Accepted 2006-6-11; Published 2006-6-13
Abstract
Background: The +T294C polymorphism in PPARδ represents a functional SNP affecting transcriptional activity of the PPARδ gene. To address whether this polymorphism is associated with the risk for coronary heart disease and/or plasma lipid levels in women, we studied a group of 967 female patients with hyperlipidaemia in the presence (n=453) or absence (n=514) of coronary heart disease.
Methods: 967 female patients with or without coronary heart disease were genotyped using mutagenically separated polymerase chain reaction (MS-PCR). Statistical analysis was performed according to genotype with parameters of lipid metabolism as dependant variables.
Results: A highly significant association between the rare C allele and lower plasma HDL concentrations was found in female subjects. The effect remained significant after correcting for multiparametric testing according to Bonferoni and was seen only in subjects with a BMI below the median. Moreover, a significant association of the C-allele with coronary heart disease and BMI was obtained. Regarding the entire group, trends towards higher VLDL and LDL levels were observed.
Conclusions: Our data show for the first time that the PPARδ +T294C polymorphism is associated with lipid levels and coronary heart disease in women. However, the molecular mechanism of action remains to be elucidated.
Keywords: PPARδ, +T294C, coronary heart disease
1. Background
Peroxisome Proliferator-activated receptors (PPAR) are ligand activated transcription factors involved in the regulation of energy balance [1]. Three closely related members belong to the PPAR subgroup termed PPARα, PPARγ, and PPARδ. While PPARα is mainly expressed in liver, muscle, kidney and heart, PPARγ is most abundant in adipocytes, intestinal cells and macrophages and PPARδ is expressed in many tissues [2-4]. Each subgroup is activated by a certain variety of fatty acids and their derivatives and by specific pharmacological ligands. After forming obligate heterodimers with the retinoid X receptor, PPARs bind to specific elements in the promoter region of target genes called PPAR response elements (PRE), thereby altering metabolism by activating a network of downstream genes. Whereas PPARα promotes ß-oxidation, PPARγ has been attributed the role of a master regulator of adipocyte differentiation. In contrast, the precise function of PPARδ is less well established. Treatment of obese rhesus monkeys with the synthetic PPARδ agonist GW501516 resulted in an increase of high-density lipoprotein cholesterol (HDL) levels and a decrease of plasma triglycerides [5]. In addition, db/db mice expressing an activated form of PPARδ are resistant to obesity and hyperlipidaemia when overfed and muscle-specific overexpression of the receptor increased the number of muscle fibers with high oxidative metabolic capability [6,7]. Taken together, these results imply an important role of PPARδ in lipid metabolism and thermogenesis.
The +294T/C polymorphism in exon 4 of the PPAR δ gene was initially described by Skogsberg et al [8]. It was shown that the polymorphism influenced binding of Sp-1 resulting in higher transcriptional activity for the rare C allele than the common T allele. In a group of 543 healthy, middle-aged men the C genotype was associated with elevated levels of low-density lipoprotein (LDL) cholesterol and apolipoprotein B (Apo B). In 580 male subjects with hyperlipidaemia recruited from the West of Scotland Coronary Prevention Study (WOSCOPS) carriers of the C allele had significantly lower HDL plasma concentrations and homozygotes had a tendency towards a higher risk for coronary heart disease (CHD) [9]. In order to investigate whether these associations are also valid in women, we studied the +294T/C polymorphism in a group of 967 female patients with mixed hyperlipidaemia in the presence or absence of coronary heart disease.
2. Methods
Patients
All subjects were recruited between 1999 and 2004 from patients who attended the obesity and cardiology outpatient clinics of the University of Hamburg Hospital. Table 1 summarizes the patient characteristics. The vast majority were Caucasians and to our knowledge there were no related subjects among the samples. Informed consent was obtained from all patients and the study was approved by the local ethics commission. After measuring lipid values at their first visit to the clinic any lipid lowering medication was discontinued unless in patients presented with unstable angina or recent myocardial infarction. Only patients were included in whom a cessation of medication could be performed. All patients received dietary advice in a 30-60 minute discussion with a dietary assistant in which they were instructed to reduce their daily fat intake to a maximum of 40-55 g. Patient compliance was supervised by handing out a nutritional diary which was submitted at each of the further visits to the clinic. After 6 weeks the patients attended the clinic again and it is the lipid values and biometric data obtained at this visit which were used for statistical analysis in this study. The presence of chd was assessed by medical records and/or self-reported questionnaire and each case was validated by cardiac ultrasound and/or electrocardiographic examinations or by coronary angiography.
Patient characteristics.
| CHD- | CHD+ | p-value | |
|---|---|---|---|
| n | 514 | 453 | |
| Age (years) | 42.2 | 44.1 | |
| BMI (kg/m²) | 25.9±6 | 28.8±7.4 | 0.01 |
| Cholesterol (mg/dl) | 269.6±65.7 | 285.3±131.3 | ns |
| LDL | 172.9±59.1 | 195.2±76.3 | 0.02 |
| HDL | 56.5±18.4 | 43.7±12 | <0.001 |
| VLDL | 28.5±14.2 | 29.3±15.3 | 0.015 |
| Triglycerides (mg/dl) | 220.8±119.9 | 237.9±112 | 0.001 |
| Apo B (mg/dl) | 131.7±28.2 | 189±29.9 | 0.005 |
| Apo A1 (mg/dl) | 162.8±5.8 | 128±34.1 | 0.03 |
| Lp(a) (mg/dl) | 24.1±8.7 | 37.4±9.6 | ns |
Biochemical measurements
Plasma total cholesterol and triglycerides were determined using GPO-PAP and CHOD-PAP kits, respectively, from Boehringer Mannheim (Mannheim, Germany). HDL was measured following precipitation of apoB containing lipoproteins with phosphotungstate (Boehringer Mannheim). Apoliprotein AI (Apo AI) and Apo B were measured using the Beckmann Array 360 (Beckmann Instruments).
Genotyping
DNA was extracted from white blood cells using standard methods. A mutagenically separated polymerase chain reaction (MS-PCR) was developed to separate wildtype (wt) and mutated alleles as described [10]. The longer primer for detection of the wt allele was 5'-TTC AAG CCC TGA TGA TAA GGT CTT TGG CAT TAG ATG CTG TTT TGT TTT-3.' The shorter primer was 5'-CTT TTG GCA TTA GAT GCT GTT TTG TCC TG-3.' As reverse primer we used 5'-CTT CCT CCT GTG GCT GCT C-3.' PCR condition were: 96°C for an initial 5 minutes (min.) followed by 96°C 1 min., 65°C 1 min., 72°C 1 min. for 43 cycles and final extension at 72° for 10 min. MS-PCR created two fragment of divergent length which were detected by silver staining on polyacrylamid gel.
Statistical Analysis
Allele frequencies were determined by gene counting and compared using the ℵ2 test. The data was analysed using SPSS 12.0 software. The association between the +294T/C polymorphism and serum lipids, coronary heart disease, and body mass index (BMI) was calculated using the Student's t-test analysis with genotype as the group variable and total cholesterol, triglycerides, vldl-, ldl-, hdl cholesterol and lipoprotein (a) as dependent variables. Bonferoni's correction for multiple testing was performed by multiplying the p value with the number of tests where appropriate. To prevent generating subgroups too small for analysis, allelic variants were dichotomized in TT and TC/CC. A p-value of 0.05 or lower was considered statistically significant. Data were adjusted for age, smoking, and BMI where appropriate.
3. Results
The frequency of the C-allele was 0.24 and the genotypes were distributed according to the Hardy Weinberg Equilibrium. Overall the C allele frequency was higher in patients with coronary heart disease than in those without, but this difference was not statistically significant. The plasma lipid values according to +294T/C genotype are shown in table 2. There was a trend towards higher LDL in the group of C allele carriers but with this was not statistically significant. However patients carrying the C allele presented a significantly lower VLDL. In the entire group no significant interaction between genotype and coronary heart disease could be detected. Eliminating patients with diabetes mellitus or triglycerides higher than 1.000 mg/dl which likely represent cases of secondary hyperlipidaemia did not alter our findings. A possible gene-to-gene interaction between PPARδ +294T/C and Apolipoprotein E (ApoE) was analysed for each ApoE genotype separately but did not present statistically significant results (data not shown).
Plasma lipid values according to genotype.
| TT | CT/CC | p-value | |
|---|---|---|---|
| n | 599 | 368 | ns |
| Age (years) | 43.4±14.8 | 42.8±15.9 | ns |
| BMI (kg/m²) | 24.9±6 | 24.1±6.3 | ns |
| Cholesterol (mg/dl) | 266±63.3 | 267±80.3 | ns |
| LDL | 172±73.2 | 182±55.5 | ns |
| HDL | 53±18.7 | 53±17.8 | ns |
| VLDL | 27±15.4 | 24±12.5 | 0.033 |
| Triglycerides (mg/dl) | 147±137.3 | 127±169.3 | ns |
| Apo B (mg/dl) | 129±35.6 | 131±26.3 | ns |
| Apo A1 (mg/dl) | 138±26.9 | 172±35.8 | ns |
| Lp(a) (mg/dl) | 11±9.7 | 6±12.1 | ns |
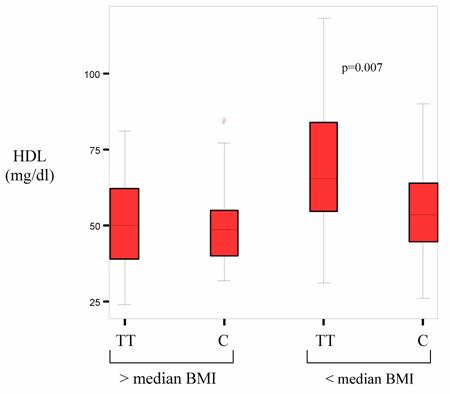
HDL plasma levels are inversely related to BMI. Thus potential associations of PPARδ with HDL-C levels could be masked by the profound effect of overweight on HDL-C in the patients with high BMI. To establish whether differences regarding the BMI affected our data the allele frequencies in patients with a BMI above the median were compared with those below the median. Patient characteristics for each group are presented in table 3. Within the group of leaner patients there was a significantly higher incidence of coronary heart disease for carriers of at least one C allele (p=0.033) (table 4). Among those with a BMI below 24.6 carriers of the rare C allele had a significantly higher BMI than T allele homocygotes (p=0.023). Whereas no effect was seen on total cholesterol and LDL, C allele carriers had a lower HDL than T homozygotes and this difference was statistically highly significant (p=0.007) (figure 1). Further division of patients created subgroups to small for statistical analysis.
Clinical characteristics of patients above and below median BMI (24.6)
| Below median BMI | Above median BMI | |
|---|---|---|
| n total | 561 | 406 |
| n with chd | 247 | 206 |
| n without chd | 314 | 200 |
| Age (years) | 41.3±16.3 | 45.5±14.4 |
| BMI (kg/m²) | 22.6±1.5 | 33.8±4.2 |
| Cholesterol (mg/dl) | 268±13.9 | 279.2±44.8 |
| LDL | 184±21.1 | 180.3±52.6 |
| HDL | 60±12.5 | 38.8±19.2 |
| VLDL | 22.5±4.1 | 38.1±14.9 |
| Triglycerides (mg/dl) | 116.4±180.9 | 384±214.4 |
| Apo B (mg/dl) | 100.1±22.6 | 239.2±31.3 |
| Apo A1 (mg/dl) | 159±26.8 | 129.3±44.5 |
| Lp(a) (mg/dl) | 6.1±9.2 | 64.2±14.3 |
Patients below median BMI (24.6 kg/m²) according to genotype
| TT | CT/CC | CT | CC | p-value* | |
|---|---|---|---|---|---|
| n | 342 | 219 | 211 | 8 | |
| chd present | 81 | 71 | 70 | 1 | 0.033 |
| BMI (kg/m²) | 22.1±1.5 | 22.9±1.3 | 22.9±1.1 | 23.1±1.1 | 0.023 |
| Cholesterol (mg/dl) | 270.5±51.2 | 266.2±89.4 | 266.4±93.2 | 262±64.5 | ns |
| LDL | 183±46.8 | 189±54.8 | 194±53.9 | 174.6±62.5 | ns |
| HDL | 65.5±20.6 | 53.5±16 | 54±16 | 58.2±17.8 | 0.004 |
| VLDL | 22±13.2 | 23±8.8 | 23.4±8.8 | 19.2±4.8 | ns |
| Triglycerides (mg/dl) | 112±115.4 | 118±181.6 | 118.9±84.5 | 96.4±23.8 | ns |
| Apo B (mg/dl) | 100±22.6 | 103.3±21.1 | 103±11.1 | 111.5±9.8 | ns |
| Apo A1 (mg/dl) | 159±26.8 | 161±24.3 | 161.1±13.3 | 159±22.1 | ns |
| Lp(a) (mg/dl) | 6±4.2 | 12±14.1 | 12.1±8.8 | 10.7±3.4 | ns |
*referring to TT vs CT/TT; Student's t-test.
HDL-levels according to genotype above and below median BMI. Boxes represent T allele homocygotes (TT) or C allele carriers (C).
4. Discussion and Conclusions
To address the question whether the +T294C polymorphism in PPARδ is associated with the risk for coronary heart disease and/or plasma lipid levels in women, we investigated this genetic variant in a group of female patients with hyperlipidaemia in the presence or absence of coronary heart disease. As previously observed in men by Skogsberg et al. [9], we found a highly significant association between the rare C allele and lower plasma HDL concentrations in female subjects. The effect remained significant after correcting for multiparametric testing according to Bonferoni and was seen only in subjects with a BMI below the median. In addition, the non-corrected data revealed a significant association of the C-allele with coronary heart disease. Regarding the entire group, trends towards higher VLDL and LDL levels were observed. Results in support of our findings were also obtained by Chen et al. [11], who studied 4 PPARδ haplotypes and found that effects on plasma lipids and coronary lesions were primarily due to the influence of the +T294C SNP. The C allele frequency of 0.24 in our high risk patient group was similar as the frequency of 0.21 reported by Chen et al. in their study for chd patients derived from the lipoprotein coronary atherosclerosis study. Conversely, Skogsberg et al. found a somewhat lower C allele frequency in their study with healthy young men from Sweden [9].
Treatment of obese rhesus monkeys with GW 501516, a selective PPARδ agonist, led to a significant improvement of metabolic traits characterized by a rise of HDL and lowering of LDL, triglycerides, and insulin [5]. In addition, transgenic expression of an activated PPARδ in mice conferred resistance to obesity and hyperlipidaemia [6]. Considering that the C allele is linked to a higher basal transcriptional activity, it is currently unclear why this allele is associated with lower HDL and a trend towards higher LDL levels in humans. Of note both animal studies used obese models to investigate the effect of an activated PPARδ. In an obese state the activities of numerous interacting genes are altered which may influence the effect of PPARδ on metabolism. In our data and in the study by Skogsberg et al., the effect of the C allele on lipid levels increased if over-weight individuals were excluded from the study.
Since Skogsberg et al. and also Chen et al. [8, 11] have shown that the PPARδ T+294C SNP is associated with elevated plasma LDL cholesterol and apoB levels in men and we show in this work that similar associations are operating in women, we also looked for a potential gene-to-gene interaction between PPARδ and apoE. Allelic variation in apoE is known to be associated consistently with plasma concentrations of cholesterol, LDL cholesterol and apoB in men and women (reviewed in ref. [12]). However, our data do not provide evidence for a gene-to-gene interaction between both genes and exclude the possibility that the observed associations are due to apoE rather than PPARδ in our study.
Conflicts of interest
The authors have declared that no conflict of interest exists.
References
1. Evans RM, Barish GD, Wang YX. PPARs and the complex journey to obesity. Nat Med. 2004 ;4:1-7
2. Reddy JK, Hashimoto T. Peroxisomal ß-Oxidation and peroxisome proliferator activated receptor α: an adaptive metabolic system. Annu Rev Nutr. 2001 ;21:193-230
3. Spiegelman BM, Flier JS. Adipogenesis and obesity: rounding the big picture. Cell. 1996 ;87:377-389
4. Skogsberg J, Kannisto R, Roshani L, Gagne E, Hamsten A, Larsson C, Ehrenborg E. Characterization of the human peroxisome proliferator activated receptor delta gene and its expression. Int J Mol Med. 2000 ;6:73-81
5. Olivier WR, Shenk JL, Snaith MR, Russell CS, Plunket KD, Bodkin NL, Lewis MC, Winegar DA, Sznaidman ML, Lambert MH, Xu HE. A selective peroxisome proliferator-activated receptor δ agonist promotes reverse cholesterol transport. Proc Nat Acad Sci USA. 2001 ;98:5306-5311
6. Leibowitz MD, Fievet C, Hennuyer N, Peinado-Onsurbe J, Duez H, Bergera J, Cullinan CA, Sparrow CP, Baffic J, Berger GD, Santini C, Marquis RW, Tolman RL, Smith RG, Moller DE, Auwerx J. Activation of PPAR delta alters lipid metabolism in db/db mice. FEBS Lett. 2000 ;473(3):333-336
7. Luquet S, Lopez-Soriano J, Holst D, Fredenrich A, Melki J, Rassoulzandegan M, Grimaldi PA. Peroxisome proliferator-activated receptor δ controls muscle development and oxidative capability. FASEB. 2003 ;17:2299-2301
8. Skogsberg J, Kannisto K, Cassel TN, Hamsten A, Eriksson P, Ehrenborg E. Evidence That Peroxisome Proliferator-Activated Receptor Delta Influences Cholesterol Metabolism in Men. Arterioscler Thromb Vasc Biol. 2003 ;23:637-643
9. Skogsberg J, McMahon AD, Karpe F, Hamsten A, Packard CJ, Ehrenborg E. Peroxisome proliferator activated receptor delta genotype in relation to cardiovascular risk factors and risk of coronary heart disease in hypercholesterolaemic men. J Int Med. 2003 ;254:597-604
10. Rust S, Funke H, Assmann G. Mutagenically separated PCR (MS- PCR): a highly specific one step procedure for easy mutation detection. Nucl A Res. 1991 ;21:3623-3629
11. Chen S, Tsybouleva N, Ballantyne CM, Gotto AM, Marian AJ. Effects of PPARα, γ and δ haplotypes on plasma levels of lipids and progression of coronary atherosclerosis and response to statin therapy in the lipoprotein coronary atherosclerosis study. Pharmacogenet. 2004 ;14:61-71
12. Eichner JE, Dunn ST, Perveen G, Thompson DM, Stewart KE, Stroehla BC. Apolipoprotein E polymorphism and cardiovascular disease. Am J Epidemiol. 2002 ;155:487-495
Author contact
Corresponding address: Jens Aberle, e-mail: aberleuni-hamburg.de, Tel.: 0049 40 42803 4449 Fax: 0049 40 42803 8290

 Global reach, higher impact
Global reach, higher impact